Abstract
The prion protein (PrP) is a cell surface protein that in disease misfolds and becomes infectious causing Creutzfeldt-Jakob disease in humans, scrapie in sheep, and chronic wasting disease in deer and elk. Little is known regarding the dimerization of PrP and its role in disease. We developed a bioluminescent prion assay (BPA) to quantify PrP dimerization by bimolecular complementation of split Gaussia luciferase (GLuc) halves that are each fused to PrP. Fusion constructs between PrP and N- and C-terminal GLuc halves were expressed on the surface of RK13 cells (RK13-DC cells) and dimerized to yield a bioluminescent signal that was decreased in the presence of eight different antibodies to PrP. Dimerization of PrP was independent of divalent cations and was induced under stress. Challenge of RK13-DC cells with seven different prion strains did not lead to detectable infection but was measurable by bioluminescence. Finally, we used BPA to screen a compound library for compounds inhibiting PrP dimerization. One of the most potent compounds to inhibit PrP dimerization was JTC-801, which also inhibited prion replication in RML-infected ScN2a and SMB cells with an EC50 of 370 nM and 220 nM, respectively. We show here that BPA is a versatile tool to study prion biology and to identify anti-prion compounds.
Similar content being viewed by others
Introduction
The prion protein (PrPC) is a natural protein that is predominantly expressed on the outer cell membrane of neurons1. The structure of PrPC is well characterized and has been determined by nuclear magnetic resonance (NMR) spectroscopy and X-ray crystallography2,3. PrPC has an unstructured, flexible N-terminus followed by a globular domain with three α-helices and little β-sheet structure, and is tethered to the cell surface by a carboxy (C)-terminal GPI anchor4. During spontaneous or templated misfolding, PrPC undergoes a conformational transition where it loses all of its α-helical content and adopts mostly a β-sheet structure that is not fully defined yet but likely to consist of a four-rung ß-solenoid architecture5,6. This β-sheet-rich conformer, PrPSc, is prone to aggregation, infectious, and toxic to neurons causing neurodegeneration and death1,7. Fascinatingly, prion diseases are the only unequivocally confirmed disease group to be sporadic, genetic, and infectious in origin. Prion diseases affect humans and some other mammals, most common in humans being sporadic Creutzfeldt-Jakob disease (sCJD), in cattle bovine spongiform encephalopathy (BSE), in sheep scrapie, and in deer and elk chronic wasting disease (CWD). PrPSc can exist in multiple conformations — strains — with specific biophysical and biochemical properties that are maintained between hosts upon transmission and determine the clinical manifestation, the phenotype, of a particular prion disease8. In humans, for instance, based on the strain, PrPSc can cause CJD or Kuru, two different human prion diseases with very different incubation times and clinical presentation9.
The physiological function of PrPC is not fully understood. A plethora of divergent functions for PrPC have been proposed over the years, leaving it unclear which of them may be more relevant10,11. More recent results showing that aged Prnp knockout mice develop a chronic demyelinating polyneuropathy12 led to the finding that PrPC functions as a ligand to the G protein-coupled receptor Adgrg6 expressed in Schwann cells13. Also, identification of PrPC as a member of the ZIP family of metal ion transporters14 helped to elucidate its role in polysialylation of neural cell adhesion molecule 1 (NCAM1) during epithelial-to-mesenchymal cell transition15. PrPC also has been reported to form homodimers that exist in a monomer-dimer equilibrium, which is a characteristic of receptor proteins involved in signal transduction, and which may also be relevant during the conversion of PrPC to PrPSc16,17.
Prion diseases are despite continuing efforts in drug screening to find a treatment, unfortunately, still without cure. Only few drugs have made it into clinical trials, all of which have either failed or are ongoing18. Next to transmission experiments to animals many sophisticated tools have been developed over the years to detect and quantify prions and the effect of anti-prion drugs in vitro. These include methods as simple as proteolytic treatment of prion-infected cell lysates with proteinase K (PK) to distinguish between PK-sensitive PrPC and PK-resistant PrPSc after immunodetection with antibodies to PrP. Lately, more refined and scalable methods such as the conformation-dependent immunoassay (CDI)8, protein-misfolding cyclic amplification (PMCA)19, the standard scrapie cell assay (SSCA)20, or the real-time quaking induced conversion assay (RT-QuIC)21 have been developed.
To our knowledge no scalable assay has been developed to date to directly study the interaction of prion proteins in cells by bioluminescence. The overall aim of this study was to develop a bioluminescent cell assay that would facilitate studying the biology of the prion protein in its native state. We report here a bioluminescent prion assay (BPA) for the quantification of PrPC dimerization. Based on the concept of protein-fragment complementation22 and the fact that PrPC can dimerize, constructs between mouse PrP and split Gaussia luciferase halves were expressed in RK13 cells, which were bioluminescent and showed that GPI-anchored fusion constructs of PrPC dimerize on the cell surface under physiological conditions. Treatment of these cells with eight different antibodies to PrP, especially those binding to the first α-helix of PrPC, was able to disrupt PrPC-mediated dimerization. Dimerization of PrPC fusion constructs did not require divalent cations and was induced under stress when divalent cations were increasingly chelated. Challenge with seven different prion strains of cells expressing PrPC fusion constructs induced bioluminescence within as little as three days. A screen of a library with 1,640 compounds identified 240 compounds inhibiting dimerization of PrPC fusion constructs by 20–85%. JTC-801, a quinoline derivative, potently inhibited dimerization of PrPC fusion constructs by 80% and prion replication in RML-infected ScN2a and SMB cells with an EC50 of 370 nM and 220 nM, respectively. Our data shows that the bioluminescent prion assay is a versatile tool to study the biology of prion proteins, and that it can be used to identify compounds inhibiting PrPC dimerization that also inhibit prion replication.
Results
Design of fusion constructs between PrP and N- and C-terminal Gaussia luciferase halves
To study dimerization of the prion protein (PrP) by bioluminescence in cells, we cloned fusion constructs between the PrP (Fig. 1a) and the N- and C-terminal halves of split Gaussia luciferase (GLuc) into a bicistronic expression vector that allows expression of two proteins from a single mRNA using an internal ribosomal entry site (IRES). Insertion of amino acids 2–93 of GLuc flanked by two flexible linker sequences consisting of GGGGSGGGS between amino acids 229 and 230 of PrP yielded PrP-NGLuc (Fig. 1b), and similar insertion of amino acids 94–168 of GLuc yielded PrP-CGLuc (Fig. 1c). To obtain N- and C-terminal halves of GLuc that are also expressed on the cell surface as PrP-NGLuc and PrP-CGLuc but essentially lack all of PrP that enables dimerization, we inserted N- and C-terminal halves of GLuc with only a C-terminal flexible linker between amino acids 22 and 230 of PrP yielding NGLuc (Fig. 1d) and CGLuc (Fig. 1e). During expression proteins are directed to the endoplasmic reticulum where the leader peptide 1–22 of PrP is cleaved off. Also, the C-terminal signal peptide 231–254 of PrP is replaced by a glycosylphosphatidylinositol (GPI)-anchor attached to serine 230 of PrP, which tethers mature proteins to the outer cell membrane.
Schematic presentation of PrP-luciferase fusion constructs. (a) A signal peptide of 22 amino acids (yellow) directs expression of mouse PrP to the endoplasmic reticulum and is cleaved off of the preprotein. Also, amino acids 231–254 (blue) are cleaved off of the preprotein and replaced by a glycosylphosphatidylinositol (GPI) anchor that is attached to serine 230 and tethers mature mouse PrP (red) to the outer cell membrane. (b) PrP-NGLuc is a fusion construct that has the N-terminal half, amino acids 2–93, of Gaussia luciferase (GLuc2–93) inserted between amino acids 229 and 230 of mouse PrP, and is flanked by the flexible linker sequence GGGGSGGGGS on each side (black). Mature PrP-NGLuc has a GPI anchor attached to its last amino acid, which is serine 230 of PrP. (c) PrP-CGLuc is a fusion construct that has the C-terminal half, amino acids 94–168, of Gaussia luciferase (GLuc94–168) inserted between amino acids 229 and 230 of mouse PrP, and is flanked by the flexible linker sequence GGGGSGGGGS on each side (black). Mature PrP-NGLuc has a GPI-anchor attached to its last amino acid, which is serine 230 of PrP. (d) NGLuc is a fusion construct that has the N-terminal half, amino acids 2–93, of Gaussia luciferase (GLuc2–93) inserted between amino acids 22 and 230 of PrP, and is flanked by the flexible linker sequence GGGGSGGGGS on each side (black). Amino acids 23–229 of PrP are missing in this construct. Mature NGLuc has a GPI-anchor attached to its last amino acid, which is serine 230 of PrP. (e) CGLuc is a fusion construct that has the N-terminal half, amino acids 94–168, of Gaussia luciferase (GLuc94–168) inserted between amino acids 22 and 230 of PrP, and is flanked by the flexible linker sequence GGGGSGGGGS on each side (black). Amino acids 23–229 of PrP are missing in this construct. Mature CGLuc has a GPI-anchor attached to its last amino acid, which is serine 230 of PrP. All constructs carry the 3F4 epitope (asterics), which allows detection with the 3F4 antibody.
Cellular expression of PrP-NGLuc and PrP-CGLuc results in PrP dimerization and bioluminescence
Because the rabbit kidney epithelial cell line RK13 lacks endogenous expression of rabbit PrP (Fig. 2a) as reported previously23, we used these cells to stably coexpress PrP-NGLuc and PrP-CGLuc (RK13-DC cells), and other control constructs. Expression of these fusion constructs on the cell surface was confirmed by FACS analysis with an antibody against GLuc (Fig. 3a,b), and the SAF32 antibody against PrP (Fig. 3c–e). Biochemical analysis of cell lysates after western blotting with antibodies to PrP showed that the fusion constructs ran at a higher molecular weight than observed for PrPC in N2a cells. Also, treatment of cell lysates with peptide-N-glycosidase F (PNGaseF) revealed that PrP-NGLuc and PrP-CGLuc were properly glycosylated in RK13 cells because deglycosylation reduced the multiple bands seen without PNGaseF treatment to two major bands representing PrP-NGLuc and the slightly smaller PrP-CGLuc (Fig. 2a and Supplementary Fig. 1A). Biochemical analysis of cell lysates after western blotting with an antibody to GLuc showed that also NGLuc and CGLuc were expressed in RK13-NGLuc/CGLuc control cells and ran at a much lower molecular weight size representative for the lack of PrP in these constructs (Supplementary Fig. 1A). Treatment of RK13-DC cells with increasing amounts of phosphoinositide phospholipase C (PI-PLC) increased the amount of protein detectable in the cell culture supernatant after western blotting with an antibody against PrP suggesting that mature PrP-NGLuc and PrP-CGLuc were anchored to the outer cell membrane by GPI (Fig. 2b,c). Live cell imaging with an antibody to PrP showed that PrP-NGLuc and PrP-CGLuc were expressed on the surface of RK13-DC cells as PrP on N2a cells (Fig. 2d and Supplementary Fig. 1B). Equally, immunocytochemistry with an antibody to GLuc showed that NGLuc and CGLuc were expressed on the surface of RK13-NGLuc/CGLuc cells (Supplementary Fig. 1B). Bioluminescence measurements in RK13-DC cells resulted in a more than 10-fold higher signal in comparison to RK13 cells expressing only PrP-NGLuc or PrP-CGLuc, or NGLuc and CGLuc together (Fig. 2e), suggesting that PrP-NGLuc and PrP-CGLuc dimerize in RK13-DC cells but not NGLuc and CGLuc in RK13-NGLuc/CGLuc negative control cells. In comparison to PrP-NGLuc and PrP-CGLuc in RK13-DC cells, bioluminescence of full-length GLuc in RK13 cells was nearly 17-fold higher (Fig. 2e).
PrP-NGLuc and PrP-CGLuc expression in RK13-DC cells results in bioluminescence. (a) Western blot analysis with the Sha31 antibody against PrP shows that RK13-DC cells stably express PrP-NGLuc and PrP-CGLuc from a bicistronic expression vector (lanes 3 and 4). Non-transfected RK13 cells do not express PrP and do not show a detectable signal for PrP (lanes 1 and 2), whereas N2a cells show the characteristic signal for wild-type PrP (lanes 5 and 6). Deglycosylation of cell lysates with peptide-N-glycosidase F (PNGase F, lanes 2, 4, and 6) resulted in lower molecular weight bands suggesting that mature PrP-NGLuc and PrP-CGLuc are properly glycosylated in RK13-DC cells as PrP is in N2a cells. Detection of tubulin on the same blot served as a loading control. Additional lanes were excised for presentation purposes. (b) Western blot analysis with the Sha31 antibody against PrP shows that treatment of RK13-DC cells with increasing amounts of phosphoinositide phospholipase C (PI-PLC) increases the amount of detectable protein released into cell culture medium suggesting that mature PrP-NGLuc and PrP-CGLuc are anchored to the outer cell membrane by GPI. Additional lanes were excised for presentation purposes. (c) Densitometric quantification of the PrP signal from three different experiments as shown in (b). (d) Life cell immunofluorescence staining with the Sha31 antibody to PrP (left panels) shows that PrP-NGLuc and PrP-CGLuc are expressed in RK13-DC cells (top row), similar to wild-type PrP in N2a cells (bottom row). Bright-field microscopy (centre panels) shows the contours of the imaged cells. An overlay of fluorescence and bright-field images (right panels) shows that PrP-NGLuc, PrP-CGLuc, and PrP are located at the cell membrane. Bar = 20 µm. (e) The bioluminescence measured from RK13-DC cells expressing both PrP-NGLuc and PrP-CGLuc was more than 10-fold above background levels measured from RK13 cells stably expressing only PrP-NGLuc or PrP-CGLuc, or both NGLuc and CGLuc together each lacking the PrP moiety. A functional luciferase consisting of its N- and C-terminal halves was only formed in RK13-DC cells where dimers between the PrP moieties of PrP-NGLuc and PrP-CGLuc could form. In comparison to PrP-NGLuc and PrP-CGLuc in RK13-DC cells, bioluminescence of full-length GLuc in RK13 cells was almost 17-fold higher. In (c,e) error bars indicate SD.
Fusion constructs are stably expressed on the surface of RK13 cells. (a) FACS analysis with an antibody to Gaussia luciferase confirms that in contrast to non-transfected RK13 cells (blue) RK13-NGLuc/CGLuc cells (red) stably express NGLuc and CGLuc lacking the PrP moiety on the cell surface. (b) Equally, the same anti-luciferase antibody also confirms expression of PrP-NGLuc and PrP-CGLuc on the surface of RK13-DC cells (red) that yield a right-shifted signal in comparison to non-transfected RK13 cells (blue). (c) FACS analysis with the SAF32 antibody against PrP confirms that in contrast to RK13 cells (blue), stably transfected RK13-PrP-NGLuc cells (red) express PrP-NGLuc on their cell surface. (d) In contrast to RK13 cells (blue), stably transfected RK13-PrP-CGLuc cells (red) express PrP-CGLuc on their surface, as determined with SAF32 antibody. (e) A signal for PrP was also detected with the SAF32 antibody in RK13-DC cells (red) stably expressing both PrP-NGLuc and PrP-CGLuc but not from non-transfected RK13 cells (blue).
Dimerization of PrP is inhibited by anti-PrP antibodies
To investigate PrP dimerization in more detail, we measured the effect of eight different antibodies (Table 1) against PrP on the bioluminescence in RK13-DC cells. The antibodies SAF32 and SAF34 bind to the octarepeat region (OR), 6D11 between the OR and the hydrophobic domain (HD), SAF83, SAF61, and Sha31 to the first α-helix (α1), EP1802Y to the far C-terminus of the third α-helix (α3), and BAR236 to an unidentified conformational epitope (Fig. 4a). In contrast to unspecific mouse IgG, which served as a negative control, all eight antibodies tested against PrP reduced the bioluminescence measured from RK13-DC cells after 24 h, suggesting that steric hindrance of bound antibodies inhibited dimerization of PrP (Fig. 4b). Binding of these antibodies to PrP was not toxic to RK13-DC cells within 24 h as determined by an MTT assay (Fig. 4c). Antibodies binding to α1 more potently inhibited dimerization of PrP than those binding to more N- or C-terminal domains of PrP or BAR236 binding to a conformational epitope. Increasing antibody concentrations to PrP led to a concentration-dependent reduction in bioluminescence from RK13-DC cells and reached maximal inhibition between 2–3 µg/mL for SAF83 and Sha31 (Fig. 4d). Inhibition of bioluminescence from RK13-DC cells with antibodies to PrP was immediate and could be measured already after 1 h and remained constant over 48 h relative to untreated cells or cells treated with unspecific mouse IgG while the overall bioluminescence intensities increased over time due to cell division (Fig. 4e).
Antibodies to PrP inhibit its dimerization. (a) We measured the effect of eight different PrP antibodies on PrP dimerization in RK13-DC cells by bioluminescence assay. SAF32 and SAF34 bind within the octarepeat region (OR), 6D11 between the octarepeat and hydrophobic domain (HD, black), SAF83, SAF61, and Sha31 bind to helix 1 (α1), EP1802Y to helix 3 (α3) in the C-terminus, and BAR236 an unspecified, conformational epitope of PrP. N = glycosylated asparagine residue, S–S = disulfide bond, dark grey shading = β-sheet structure, light grey shading = α-helical structure. (b) Bioluminescence measurements in the absence of antibodies were similar to those in the presence of unspecific mouse IgG, which did not affect bioluminescence and PrP dimerization. In contrast, antibodies binding to or near helix 1 of PrP strongly reduced the bioluminescence and dimerization of RK13-DC cells, whereas those binding to more N- and C-terminal regions of PrP, such as the octarepeat region or helix 3, had a smaller effect on bioluminescence. The effect of Bar236 on bioluminescence was close to those antibodies binding to the core of PrP. (c) The metabolic activity of RK13-DC cells after antibody treatment for 24 h was similar to untreated cells or cells treated with unspecific mouse IgG suggesting that the antibody treatment was not toxic to the cells as determined by MTT assay. In contrast, treatment of RK13-DC cells for 24 h with 10% DMSO was toxic and abolished all metabolic activity. (d) Treatment of RK13-DC cells for 24 h with increasing antibody concentrations of Sha31 or SAF83 showed a dose dependent effect on the reduction of bioluminescence and PrP dimerization. (e) Treatment of RK13-DC cells with the Sha31 or SAF83 antibody had an immediate effect on the reduction of bioluminescence, which was already observable within 1 h after treatment by comparison to untreated cells or cells treated with unspecific mouse IgG. The reduction of bioluminescence by the PrP-specific antibodies relative to the controls remained constant over time for up to 48 h, while the absolute bioluminescence increased due to cell division. Error bars indicate SD (****p < 0.0001, ***p < 0.001, **p < 0.01, *p < 0.05; one-way ANOVA, followed by Dunnett’s multiple comparisons test).
Dimerization of PrP is independent of divalent cations
To assess whether dimerization of PrP requires the presence of divalent cations, such as Cu2+ or Zn2+, we treated RK13-DC cells with increasing concentrations of ethylenediaminetetraacetic acid (EDTA) for 1 h to chelate divalent cations and measured its effect on bioluminescence (Fig. 5). To our surprise chelation of divalent cations did not reduce but induce bioluminescence from RK13-DC cells, suggesting, first, that PrP does not require divalent cations to dimerize, and, second, that dimerization of PrP increases due to EDTA-induced cell stress. Measurement of the metabolic activity of RK13-DC cells by MTT assay showed that increasing concentrations of EDTA were toxic to the cells (Fig. 5).
Dimerization of PrP does not require divalent cations. Treatment with increasing concentrations of EDTA for 1 h did not reduce bioluminescence of RK13-DC cells (red circles), suggesting that divalent cations are not required for PrP dimerization. On the contrary, addition of EDTA to RK13-DC cells induced their bioluminescence (red circles), while their metabolic activity (blue squares) was reduced as measured by MTT assay, suggesting that EDTA-induced cell stress promotes PrP dimerization. Error bars indicate SD.
BPA detects challenge with different prion strains
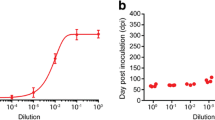
To evaluate whether challenge with prions could be detected by bioluminescence in RK13-DC cells, we challenged the cells with brain homogenate from mice that had developed prion disease after infection with the mouse-adapted scrapie RML prion strain. The bioluminescence of RML-challenged RK13-DC cells was not elevated after one day but significantly increased within three days after challenge and did not further increase after five days in comparison to untreated cells or cells treated with normal brain homogenate (NBH) from healthy mice (Fig. 6a). Survival of RK13-DC cells did not significantly differ between cells treated for 5 days with RML brain homogenate or NBH (Fig. 6b). Next to treatment with RML prions, bioluminescence in RK13-DC cells was also significantly induced after treatment with five additional mouse-adapted scrapie prion strains. These strains comprised 22 L, ME7, and 79 A (Fig. 6c) that similar to RML prions are passaged in C57BL/6 mice expressing Prnp-A, and the two prion strains 87 V and 22 A (Fig. 6d) that are passaged in VM mice expressing Prnp-B. Surprisingly, challenge of RK13-DC cells with hamster brain homogenate harbouring the hamster-adapted scrapie strain 263 K also lead to an increase in bioluminescence that was not evident after one day but progressively increased from three to five days (Fig. 6e). Similar to challenging RK13-DC cells with RML prions, challenge with 263 K prions did not affect the survival of RK13-DC cells more than NBH (Fig. 6f). To better differentiate between exogenously added PrPSc in the inoculum and endogenously converted PrP-NGLuc and PrP-CGLuc in RK13-DC cells, we had introduced the 3F4 epitope, which is absent in PrPSc in the RML brain homogenate, into the PrP moiety of PrP-NGLuc and PrP-CGLuc that allows their specific detection with the 3F4 antibody. When we tested RK13-DC cell lysates after challenge with RML prions by limited digestion with Proteinase K (PK) and western blotting with antibodies against PrP for the presence of PK-resistant PrPSc, we only detected residual PrPSc from the RML brain homogenate used to infect the cells but not PK-resistant forms of PrP-NGLuc or PrP-CGLuc in RK13-DC cells (Supplementary Fig. 2). Together with the increase in bioluminescence after prion challenge of RK13-DC cells these results suggest that PrP-NGLuc and PrP-CGLuc bind to PrPSc aggregates but that they are not converted into PK-resistant species.
Incubation of RK13-DC cells with prions induces bioluminescence. (a) We treated RK13-DC cells for 5 days with normal brain homogenate (NBH) from healthy mice or with brain homogenate from mice that had been sacrificed after they had developed disease due to an infection with RML prions. After one day of treatment the bioluminescence measured from untreated, NBH-treated, and RML-treated RK13-DC cells was comparable, whereas after three days the bioluminescence from RML-treated RK13-DC cells was significantly increased in comparison to the two negative controls. The bioluminescence levels between the three treatment groups did not change much between day three and five. (b) Survival of RK13-DC cells treated for 5 days with RML brain homogenate or NBH did not significantly differ between both groups. (c) Treatment of RK13-DC cells for five days with brain homogenates from C57BL/6 mice that had developed disease after infection with mouse-adapted scrapie strains 22 L, ME7, and 79 A caused an increase in bioluminescence in comparison to untreated cells or cells treated with NBH. (d) Equally, treatment of RK13-DC cells for five days with brain homogenates from VM mice that had developed disease after infection with the mouse-adapted scrapie strains 87 V and 22 A induced bioluminescence in comparison to untreated cells or cells treated with NBH. (e) In contrast to RK13-DC cells treated for 5 days with NBH, treatment with hamster brain homogenate containing the hamster-adapted scrapie strain 263 K progressively induced bioluminescence after 3 and 5 days. (f) An MTT assay revealed that a 5-day treatment of RK13-DC cells with the hamster-adapted scrapie strain 263 K was not more toxic than treatment with NBH. Error bars represent SD (****p < 0.0001, ***p < 0.001; one-way ANOVA, followed by Dunnett’s multiple comparisons test).
Identification of anti-prion compounds
To identify compounds inhibiting PrP dimerization, we screened Selleckchem’s Pharmacologically Active Compound Library with 1,650 compounds in RK13-DC cells and identified 240 compounds that reduced bioluminescence between 20–85% (Fig. 7a). From among these, 73 compounds reduced bioluminescence in RK13-GLuc cells stably expressing full-length GLuc, suggesting that they were either toxic or affected luciferase activity by other means. From among the remaining 167 compounds we selected 13 among the most potent (Supplementary Fig. 3), which reduced the bioluminescence of RK13-DC cells by 60–80% (Fig. 7b) but not that of RK13-GLuc cells (Supplementary Fig. 4) showing that they were not toxic or interfering with luciferase activity. From among these 13 compounds the quinoline derivative N-(4-amino-2-methylquinolin-6-yl)-2-(4-ethylphenoxymethyl)benzamide (JTC-801) (Fig. 7c) was the most potent to reduce bioluminescence in RK13-DC cells with an EC50 of less than 2 µM (Fig. 7d), a concentration that was not toxic to RK13-DC cells. Equally, JTC-801 was the only compound from among the 13 tested (Supplementary Fig. 5) to reduce the amount of PrPSc in RML-infected ScN2a cells with an EC50 of 370 nM (Fig. 7e,f), and without being toxic at these concentrations (Fig. 7f) and without reducing expression of PrPC (Supplementary Fig. 6A,B). Moreover, JTC-801 also reduced the amount of PrPSc in RML-infected SMB cells with an EC50 of 220 nM without being toxic at these concentrations (Fig. 7g,h) and without reducing expression of PrPC (Supplementary Fig. 6C,D). These results show that BPA can identify compounds inhibiting PrPC dimerization, some of which also effectively inhibit the conformational conversion of PrPC that requires interaction between misfolded and native PrP molecules.
Compound screen identifies small molecules inhibiting PrP dimerization and conversion. (a) The Pharmacologically Active Compound Library (Selleckchem) was screened in RK13-DC cells for compounds reducing bioluminescence at a concentration of 10 µM after a 24-hour treatment. From among the 1,650 compounds 240 were found to reduce bioluminescence between 20–85%. From among these 73 compounds were found to reduce bioluminescence in a counter screen in RK13 cells stably expressing full-length Gaussia luciferase (RK13-GLuc cells), suggesting that they were either toxic or negatively affecting luciferase activity. From among the remaining 167 compounds, 13 among the most potent to reduce dimerization without negatively affecting bioluminescence of RK13-GLuc cells were selected for further analysis of their anti-prion activity in ScN2a cells. (b) The 13 selected compounds strongly reduced bioluminescence of RK13-DC cells by 60–80%. (c) From among these 13 compounds the most potent to reduce bioluminescence was JTC-801, a quinoline derivative. (d) The EC50 for the reduction of bioluminescence of RK13-DC cells for JTC-801 was close to 2 µM, a concentration that did not affect cell survival within 24 h. (e,f) Biochemical and densitometric analysis shows that a 5-day treatment with JTC-801 reduces the amount of PrPSc in RML-infected ScN2a cells with an EC50 of 370 nM. For presentation purposes additional lanes were excised from the image. Treatment of ScN2a cells with JTC-801 for 5 days did not affect cell for survival for concentrations ≤1.0 µM. (g,h) Similarly, a 5-day treatment with JTC-801 reduces the amount of PrPSc in RML-infected SMB cells with an EC50 of 220 nM without being toxic for concentrations up to 1 µM. In (b,d,f,h) error bars indicate SD.
Discussion
Our data show that fusion constructs between PrPC and N- and C-terminal GLuc halves readily dimerize under physiological conditions on the surface of cells implying that dimerization of PrPC is important for its function. Although numerous functions for PrPC have been proposed in the past, the need for PrPC to dimerize to fulfil any of these functions has not been sufficiently addressed10,11,12,15. It has been suggested that dimerization of PrPC is mediated by its internal hydrophobic domain, without which dimerization and subsequent stress-induced signalling is not possible24. Here antibodies (SAF83, SAF61, and Sha31) binding proximally to the hydrophobic domain of PrP sterically hindered dimerization greater than those binding to more distal N- and C-terminal regions of PrP (SAF32, SAF34, and EP1802Y), supporting the concept that the hydrophobic domain of PrPC may be important for dimer formation. Some studies report that PrP-binding antibodies such as SAF61 induce signal transduction by crosslinking PrP molecules25. We did not study signal transduction in our cell system but our data shows that antibodies binding near the hydrophobic domain of PrPC potently inhibit its dimerization. This does not necessarily mean that antibodies inhibiting the dimerization of PrPC abolish PrPC-mediated signal transduction. In lipid rafts PrPC has also been shown to interact with transmembrane proteins such as NCAM1 to mediate cell-signalling26,27, a pathway, which is probably less active when PrPC is sequestered in its dimeric form. By inhibiting dimerization, antibodies such as SAF61 could increase the number of monomeric PrPC molecules that are free to bind NCAM1 and thus induce an alternative signalling pathway26, which remains to be shown.
The highly unstructured and conserved N-terminus of PrPC has an octarepeat region with four tandem repeats of PHGGGWGQ and two additional histidines that together can bind 5–6 divalent cations such as Cu2+ and Zn2+28,29. Using BPA we tested whether divalent cations affect dimerization of PrPC under physiological conditions. The concentration of divalent cations in 10% FBS-supplemented DMEM cell culture medium is in the low millimolar range with Ca2+ (2.93 mM) and Mg2+ (0.93 mM) being the major constituents, and with Zn2+ (4.50 µM) and Cu(II)2+ (0.34 µM) present only at micromolar concentrations30. Interestingly, EDTA concentrations greatly exceeding those of divalent cations did not reduce the bioluminescence of RK13-DC cells showing that PrPC does not require Cu2+ or Zn2+ or any other divalent cations to dimerize. Surprisingly, the bioluminescence measured from RK13-DC cells was induced in the presence of increasing EDTA concentrations, which were toxic to RK13-DC cells, suggesting that PrPC dimerizes as a cellular response to certain types of stress. Dimerization of PrPC has been reported to be protective against PrPSc-induced cell stress and to require the hydrophobic domain and GPI anchor to be protective24. Because the EDTA-induced increase in bioluminescence was measured within 1 h of treatment, stress-induced PrPC dimerization cannot be attributed to de novo protein synthesis resulting in increased protein levels but must occur through increased dimerization of monomeric PrPC molecules that are already present. PrPC has been reported to preferentially localize to lipid rafts due to its GPI anchor31,32 but PrPC is also present in non-lipid raft membrane regions, and enhanced recruitment of monomeric PrPC to lipid rafts during stress could favour its dimerization and PrPC-dependent cell signalling33.
Using BPA we could detect challenge of RK13-DC cells with six mouse-adapted and one hamster-adapted sheep scrapie strain. Very few cell lines are susceptible to infection with prions and often the spectrum of strains resulting in detectable infection for each susceptible cell line is very limited34,35,36,37,38. Our observations suggest that infected brain homogenate induces dimerization of PrP, which requires three days to become detectable by bioluminescence under these experimental conditions. To complement each other and for bioluminescence to occur, the luciferase moieties of PrP-NGLuc and PrP-CGLuc have to remain in their original conformation after binding to PrPSc. This is possible because both GLuc halves in PrP-NGLuc and PrP-CGLuc are separated from the PrP moiety by a linker domain with 10 amino acids. The extent of conformational change the PrP moiety of PrP-NGLuc and PrP-CGLuc undergoes after binding to PrPSc remains to be elucidated. Failure to detect a signal for PrP with the 3F4 antibody after proteolytic digestion of prion-challenged RK13-DC cell lysates indicates that PrP-NGLuc and PrP-CGLuc are not converted to a PK-resistant form. Surprisingly, BPA was also able to detect challenge with the hamster-adapted scrapie strain 263 K, which does not cause clinical disease in mice39. Interestingly, both hamster-adapted scrapie strains 263 K and Sc237 have been reported to cause subclinical disease in mice, without obvious generation of PK-resistant PrPSc in the subclinically infected mice. Upon passaging of brain homogenates from these subclinically infected mice, both strains resulted in clinical disease in hamsters and mice40,41. Although, NBH did not induce bioluminescence, and both, normal and infected brain homogenates, did not differ in their influence on the survival of RK13-DC cells, we cannot rule out that a component other than PrPSc from infected brains induces dimerization of PrP-NGLuc and PrP-CGLuc in RK13-DC cells.
Past efforts to identify compounds effectively inhibiting the conversion of PrPC to PrPSc in vivo have proven difficult for several reasons, particularly because the mechanism of conversion and the structure of PrPSc are poorly understood18,42. With a few exceptions most drug screening approaches are ‘black box’ approaches where compounds are screened for their potency to reduce PrPSc levels in prion-infected cells without knowing their mechanism of action43. Because PrPSc needs to bind to PrPC to convert it, our paradigm was to identify compounds that strongly bind to PrPC and thus inhibit its binding and conversion to PrPSc. Because compounds inhibiting PrPC dimerization are likely to have a strong affinity for PrPC, potentially inhibiting its conversion to PrPSc, we screened a small compound library using BPA and identified several compounds that potently inhibited dimerization of PrPC fusion constructs.
Here we have identified JTC-801 as a potent small-molecule inhibitor of prion replication. JTC-801 was first reported as a small-molecule nociceptin antagonist that binds to the nociceptin/orphanin FQ receptor (NOP) also known as the kappa-type 3 opioid receptor44. Studies in acute pain animal models revealed that it is an efficacious and potent anti-nociceptive that is available by intravenous injection and oral administration and as such may represent a novel class of analgesics45. With an EC50 of 370 nM in ScN2a cells and of 220 nM in SMB cells, JTC-801 has a comparable anti-prion efficacy to other quinoline derivatives such as quinacrine with an EC50 of 300–400 nM46,47. The fact that JTC-801 is an orally available drug and in contrast to quinacrine penetrates the brain in high enough concentrations to be potentially effective against prions in vivo, and has also been tested in Phase I and II clinical trials in Japan as an analgesic makes it a promising anti-prion drug to be tested in animal models of prion disease48,49.
In summary, the BPA is a novel scalable cell assay to quantify PrP dimerization in dependence of various stimuli. The biology of PrP is poorly understood and being able to quantify PrP dimerization as a response to stress or other stimuli could help to uncover yet unknown functions of PrP. Equally, we have shown here that BPA is a useful tool to identify novel anti-prion compounds like JTC-801 that interfere with PrPC dimerization and its conversion to PrPSc. For the scarcity of human cell lines infected with human prions, many screens utilize mouse cell lines expressing mouse PrPC and infected with mouse-adapted prions to identify anti-prion compounds that later fail to inhibit human prions from replicating when tested in mice or patients expressing human PrPC infected with CJD50. Replacement of the mouse PrP sequence in PrP-NGLuc and PrP-CGLuc with species-specific PrP, i.e. human PrPC, could greatly improve future efforts to identify species-specific anti-prion compounds.
Methods
Cloning of DNA constructs
DNA for humanized Gaussia princeps luciferase (GLuc), and fusion constructs between mouse Prnp and the N-terminal half of GLuc (PrP-NGLuc), Prnp and the C-terminal half of GLuc (PrP-CGLuc), and both constructs giving rise to NGLuc and CGLuc was custom synthesized (Thermo Fisher Scientific). The Prnp sequence included the 3F4 epitope with leucine 108 and valine 111 both mutated to methionine. The start codon of PrP-NGLuc and NGLuc was preceded by the Kozak sequence GCCACC. DNA sequences encoding amino acids 2–93 or 94–168 of GLuc were inserted between Prnp nucleotides encoding amino acids 229 and 230 of PrP and were flanked by sequences encoding a flexible GGGGSGGGS linker on both sides. To obtain the negative control constructs NGLuc and CGLuc, DNA sequences encoding amino acids 2–93 or 94–168 of GLuc with only a C-terminal GGGGSGGGS linker were inserted between Prnp nucleotides encoding amino acids 22 and 230 of PrP, eliminating most of the PrP sequence except for its N- and C-terminal signal peptides. PrP-NGLuc and PrP-CGLuc together, and NGLuc and CGLuc together were cloned into a modified pIRES2-DsRed-Express vector that allows bicistronic expression via an internal ribosomal entry site (IRES) element from a single mRNA. PrP-NGLuc and NGLuc were inserted via an N-terminal NheI and a C-terminal EcoRI restriction site, and PrP-CGLuc and CGLuc via an N-terminal PmlI and a C-terminal XhoI restriction site. GLuc was cloned into the pIRES2-DsRed-Express vector via an N-terminal NheI and a C-terminal XhoI restriction site eliminating the IRES element. All constructs were confirmed for correctness by DNA sequencing (GATC Biotech).
Cell culture
RK13 cells (American Type Culture Collection), a rabbit kidney epithelial cell line, and N2a cells (American Type Culture Collection), a mouse neuroblastoma cell line, and ScN2a and SMB cells infected with the mouse-adapted scrapie strain RML (kindly provided by Dr. David Westaway and Dr. Debbie McKenzie, University of Alberta) were grown in DMEM high glucose medium (Thermo Fisher Scientific) supplemented with 10% (v/v) foetal bovine serum (FBS; Thermo Fisher Scientific) and 1% (v/v) penicillin and streptomycin (Sigma) and incubated at 37 °C and 5% CO2. Medium was exchanged every third to fourth day. For transfections of RK13 cells, Lipofectamine 2000 (Thermo Fisher Scientific) was used following the manufacturer’s instructions. Transfected RK13 cells were analysed by fluorescence activated cell sorting (FACS) with the SAF32 antibody against PrP (Bertin Pharma) or the anti-GLuc antibody (New England Biolabs) against Gaussia luciferase for expression of PrP-NGLuc and PrP-CGLuc and similar constructs at the Flow Cytometry Core Facility of the University of Bonn using a BD FACSAria III (Becton Dickinson). RK13 cells expressing high surface protein levels were FACS sorted and maintained in 700 μg/mL G418 (Sigma) to obtain stably expressing RK13 cell lines.
Live cell imaging
For live cell imaging, N2a and RK13-DC cells were cultured in DMEM high glucose medium (Thermo Fisher Scientific) supplemented with 10% (v/v) foetal bovine serum (FBS; Thermo Fisher Scientific) and 1% (v/v) penicillin and streptomycin (Sigma) in a 35 mm high µ-Dish (Ibidi) and then incubated with the SAF32 antibody against PrP for 1 h at 37 °C. After three washes with PBS (pH 7.4), cells were incubated for 1 h with a secondary goat anti-mouse antibody conjugated to Alexa Fluor 488 (Thermo Fisher Scientific) followed by three washes with medium. Live images were visualized and recorded using an LSM700 confocal microscope (Carl Zeiss).
Survival Assay
Five thousand ScN2a cells or 10,000 stably transfected RK13 cells per well were cultured in flat-bottomed 96-well plates in DMEM medium without phenol red (Thermo Fisher Scientific) containing 10% FBS and 1% penicillin and streptomycin. After 6 h the cells were treated either with antibodies or selected compounds. Treatment with 10% dimethyl sulfoxide (DMSO; Sigma) served as a positive control for cell toxicity. After 5 days of incubation cell viability was measured in three replicates using the MTT Cell Proliferation Assay Kit (Trevigen) according to the manufacturer’s instructions. Briefly, after addition of 10 μL 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) reagent to each well, the cells were incubated for 3 h at 37 °C. Afterwards 100 μL detergent reagent was added to each well and the plates were incubated for another 2 h at 37 °C to stop the reaction, to lyse the cells, and to solubilize the formazan dye. Absorbance was measured at 570 nm from each well using a Fluostar Omega microplate reader (BMG Labtech) with a reference wavelength set to 690 nm.
Biochemical analysis
To prepare cell lysates for biochemical analysis, cells were lysed for 5 to 10 min in lysis buffer containing 10 mM Tris-HCl (pH 7.4), 150 mM NaCl, 0.5% (w/v) deoxycholate, and 0.5% (v/v) Triton X-100. Lysates were centrifuged at 1,500 x g for 5 min at 4 °C to remove cell debris. The protein concentration of supernatants was measured with the Pierce BCA Protein Assay Kit (Thermo Fisher Scientific). To detect Proteinase K (PK)-resistant PrP, cell lysates and brain homogenates were incubated with PK (New England Biolabs) at an enzyme to protein ration of 1:50 for 1 h at 37 °C with shaking at 300 rpm. PK activity was quenched by adding PMSF to a final concentration of 2 mM. To detect proper glycosylation, samples were treated with Peptide N-Glycosidase F (PNGaseF; New England Biolabs) according to the manufacturer’s instructions. Samples were further centrifuged at 100,000 × g for 1 h at 4 °C and pellets dissolved in lysis buffer and NuPAGE LDS sample buffer, boiled for 10 min, and loaded onto 4–12% NuPAGE Bis-Tris gels (Thermo Fisher Scientific). SDS-PAGE (sodium dodecyl sulfate-polyacrylamide gel electrophoresis) was processed in a MOPS buffer system (Thermo Fisher Scientific). Gels were blotted to PVDF membranes (Merck Millipore), blocked with 5% (w/v) milk in Tris-buffered saline supplemented with 0.05% Tween 20 for 1 h at room temperature and then probed with the Sha31 (Bertin Pharma) or 3F4 (BioLegend) antibody against PrP, or the anti-GLuc (New England Biolabs) antibody against Gaussia luciferase at a 1:2,000 dilution and an antibody against β-tubulin (Thermo Fisher Scientific) at a 1:2,000 dilution over night at 4 °C. Blots were developed with IRDye 680- or IRDye 800-conjugated goat anti-mouse or goat anti-rabbit secondary antibodies (LI-COR Biosciences), and imaged with an Odyssey infrared imaging system (LI-COR Biosciences). Image cropping was done in Adobe Photoshop CS5.1. For release of GPI-linked proteins from the cell surface, approximately 0.5–1.0 × 106 cells were cultured in 60 mm diameter dishes, rinsed twice with cold PBS, and then treated with 0, 0.5, or 3.0 units of phosphoinositide phospholipase C (PI-PLC) from B. cereus (Thermo Fisher Scientific) in 1.5 mL of PBS for 2 h at 4 °C. Bovine albumin from the recovered buffer was removed by overnight incubation with anti-bovine albumin (BSA)-agarose antibody (Sigma) and subsequent centrifugation at 14,000 × g for 2 min at 4 °C. The recovered supernatant was concentrated to 50 µL (20-fold) using an Amicon Ultra 0.5 mL centrifugal filter with a 3 kDa pore size. For analysis of released proteins, 32.5 µL of concentrated sample was analyzed on a 4–12% NuPAGE Bis-Tris gel (Thermo Fisher Scientific) as described above.
Bioluminescence assay and compound screening
For bioluminescence measurements, 5,000 stably transfected RK13 cells were plated in 100 µL per well in 96-well half-area plates in DMEM without phenol red containing 10% FBS, 1% penicillin and streptomycin, and 700 μg/mL G418. Depending on the experiment, cells were cultured in the presence of antibodies, brain homogenate, or ethylenediaminetetraacetic acid (EDTA; Sigma) for up to 5 days at 37 °C and 5% CO2. Immediately before starting the assay a 1.2 mM stock solution of coelenterazine (Biosynth) in methanol (Sigma) was diluted to a final concentration of 40 μM in DMEM/F-12 without phenol red. Bioluminescence was measured from each well 1 s after 30 µL of diluted coelenterazine was added with an automated pump to a well with a Fluostar Omega microplate reader (BMG Labtech), or after 10 s when the Envision Multilabel platereader (PerkinElmer) was used. Wells without cells served as blanks. Antibodies against PrP used in the bioluminescence assay were reconstituted or diluted in water and used without further purification. All monoclonal PrP antibodies from Bertin Pharma, SAF32, SAF34, SAF61, SAF83, Sha31, and BAR236, were provided as lyophilized IgG with BSA. The monoclonal 6D11 antibody against PrP was produced in the Integra flask system using serum free media and was purified using protein A affinity chromatography and provided in PBS (Covance). The monoclonal EP1802Y antibody against PrP was obtained in 50 mM Tris-Glycine (pH 7.4), 0.15 M NaCl, 40% Glycerol, 0.01% sodium azide and 0.05% BSA (Genetex). Unspecific mouse IgG antibodies were obtained as lyophil from 0.01 M phosphate buffer, pH 7.2, with 0.015 M NaCl after by fractionation and ion-exchange chromatography of pooled normal mouse serum (Sigma). Unless otherwise stated antibodies were used at 1 µg/mL.
For compound screening, 1,650 compounds of the Pharmacologically Active Compound Library (Selleckchem) were screened in triplicates in RK13-DC cells. Screening was carried out in 384-well plates (Nunc) with 5,000 cells per well in 35 µL DMEM without phenol red containing 10% FBS, 1% penicillin and streptomycin, 700 μg/mL G418, and 10 μM compound. Compounds reducing bioluminescence in RK13-DC cells were counter screened in RK13 cells stably expressing full-length Gaussia luciferase to identify toxic compounds and those negatively affecting luciferase activity. After 24 h of incubation the bioluminescence was measured from each well after adding 45 µL of 40 µM coelenterazine in DMEM without phenol red using an Envision Multilabel plate reader (PekinElmer). Wells with untreated cells were used as positive controls and wells without cells as blanks.
Challenge of RK13-DC cells with prions
To expose RK13-DC cells to infectious inocula, 5,000 cells were plated per well in a 96-well half-area plate in 97.5 µL DMEM without phenol red (Thermo Fisher Scientific) containing 10% FBS, 1% penicillin and streptomycin, and 700 μg/mL G418. Brain tissue samples from healthy C57BL6/J mice (NBH, The Jackson Laboratory) and RML-infected C57BL6/J mice (kindly provided by Dr. Walker Jackson, German Center for Neurodegenerative Diseases) were homogenized in Ca2+- and Mg2+-free phosphate-buffered saline (PBS) by three 60-s-cycles in a Precellys 24-Dual homogenizer (Peqlab) to obtain 10% (w/v) brain homogenates. The mouse-adapted scrapie strains 22 L, ME7, 79 A, 87 V, and 22 A were obtained as 10% (w/v) brain homogenates in PBS (kindly provided by Dr. Jean Manson, the Roslin TSE Resource Centre, University of Edinburgh). The 263 K hamster prion strain was obtained as a 10% (w/v) hamster brain homogenate in PBS (kindly provided by Dr. Walker Jackson). Brain homogenates were centrifuged at 1,500 x g for 5 min at 4 °C to remove cell debris. Before plating the cells, 2.5 µL of a 10% (w/v) brain homogenate in PBS from healthy mice or animals infected either with RML, 87 V, 22 A, 22 L, ME7, 79 A, or 263 K prions was added to each well. Bioluminescence of cells exposed to RML or 263 K prions was measured after 1, 3, and 5 days of incubation, while bioluminescence of cells exposed to other prion strains was measured only after 5 days. Statistical significance was assessed by one-way ANOVA, followed by Dunnett’s multiple comparisons test. All experiments were conducted in accordance with legal regulations in Germany and all experimental protocols were approved by the DZNE’s Committee for Animal Experiments.
References
Colby, D. W. & Prusiner, S. B. Prions. Cold Spring Harb Perspect Biol 3, a006833, https://doi.org/10.1101/cshperspect.a006833 (2011).
Riek, R. et al. NMR structure of the mouse prion protein domain PrP(121–231). Nature 382, 180–182, https://doi.org/10.1038/382180a0 (1996).
Haire, L. F. et al. The crystal structure of the globular domain of sheep prion protein. J Mol Biol 336, 1175–1183, https://doi.org/10.1016/j.jmb.2003.12.059 (2004).
Stahl, N., Borchelt, D. R., Hsiao, K. & Prusiner, S. B. Scrapie prion protein contains a phosphatidylinositol glycolipid. Cell 51, 229–240 (1987).
Caughey, B. W. et al. Secondary structure analysis of the scrapie-associated protein PrP 27–30 in water by infrared spectroscopy. Biochemistry 30, 7672–7680 (1991).
Vazquez-Fernandez, E. et al. The Structural Architecture of an Infectious Mammalian Prion Using Electron Cryomicroscopy. PLoS Pathog 12, e1005835, https://doi.org/10.1371/journal.ppat.1005835 (2016).
Herrmann, U. S. et al. Prion infections and anti-PrP antibodies trigger converging neurotoxic pathways. PLoS Pathog 11, e1004662, https://doi.org/10.1371/journal.ppat.1004662 (2015).
Safar, J. et al. Eight prion strains have PrP(Sc) molecules with different conformations. Nat Med 4, 1157–1165, https://doi.org/10.1038/2654 (1998).
Wadsworth, J. D. et al. Review. The origin of the prion agent of kuru: molecular and biological strain typing. Philosophical transactions of the Royal Society of London. Series B, Biological sciences 363, 3747–3753, https://doi.org/10.1098/rstb.2008.0069 (2008).
Aguzzi, A., Baumann, F. & Bremer, J. The prion’s elusive reason for being. Annu Rev Neurosci 31, 439–477, https://doi.org/10.1146/annurev.neuro.31.060407.125620 (2008).
Watts, J. C. & Westaway, D. The prion protein family: diversity, rivalry, and dysfunction. Biochimica et biophysica acta 1772, 654–672, https://doi.org/10.1016/j.bbadis.2007.05.001 (2007).
Bremer, J. et al. Axonal prion protein is required for peripheral myelin maintenance. Nat Neurosci 13, 310–318, https://doi.org/10.1038/nn.2483 (2010).
Kuffer, A. et al. The prion protein is an agonistic ligand of the G protein-coupled receptor Adgrg6. Nature 536, 464–468, https://doi.org/10.1038/nature19312 (2016).
Schmitt-Ulms, G., Ehsani, S., Watts, J. C., Westaway, D. & Wille, H. Evolutionary descent of prion genes from the ZIP family of metal ion transporters. PLoS One 4, e7208, https://doi.org/10.1371/journal.pone.0007208 (2009).
Mehrabian, M. et al. The Prion Protein Controls Polysialylation of Neural Cell Adhesion Molecule 1 during Cellular Morphogenesis. PLoS One 10, e0133741, https://doi.org/10.1371/journal.pone.0133741 (2015).
Priola, S. A., Caughey, B., Wehrly, K. & Chesebro, B. A 60-kDa prion protein (PrP) with properties of both the normal and scrapie-associated forms of PrP. J Biol Chem 270, 3299–3305 (1995).
Meyer, R. K. et al. A monomer-dimer equilibrium of a cellular prion protein (PrPC) not observed with recombinant PrP. J Biol Chem 275, 38081–38087, https://doi.org/10.1074/jbc.M007114200 (2000).
Teruya, K. & Doh-Ura, K. Insights from Therapeutic Studies for PrP Prion Disease. Cold Spring Harb Perspect Med, https://doi.org/10.1101/cshperspect.a024430 (2016).
Saborio, G. P., Permanne, B. & Soto, C. Sensitive detection of pathological prion protein by cyclic amplification of protein misfolding. Nature 411, 810–813, https://doi.org/10.1038/35081095 (2001).
Klöhn, P. C., Stoltze, L., Flechsig, E., Enari, M. & Weissmann, C. A quantitative, highly sensitive cell-based infectivity assay for mouse scrapie prions. Proc Natl Acad Sci USA 100, 11666–11671, https://doi.org/10.1073/pnas.1834432100 (2003).
Wilham, J. M. et al. Rapid end-point quantitation of prion seeding activity with sensitivity comparable to bioassays. PLoS Pathog 6, e1001217, https://doi.org/10.1371/journal.ppat.1001217 (2010).
Remy, I. & Michnick, S. W. A highly sensitive protein-protein interaction assay based on Gaussia luciferase. Nat Methods 3, 977–979, https://doi.org/10.1038/nmeth979 (2006).
Vilette, D. et al. Ex vivo propagation of infectious sheep scrapie agent in heterologous epithelial cells expressing ovine prion protein. Proc Natl Acad Sci USA 98, 4055–4059, https://doi.org/10.1073/pnas.061337998 (2001).
Rambold, A. S. et al. Stress-protective signalling of prion protein is corrupted by scrapie prions. EMBO J 27, 1974–1984, https://doi.org/10.1038/emboj.2008.122 (2008).
Mouillet-Richard, S. et al. Signal transduction through prion protein. Science 289, 1925–1928 (2000).
Santuccione, A., Sytnyk, V., Leshchyns’ka, I. & Schachner, M. Prion protein recruits its neuronal receptor NCAM to lipid rafts to activate p59fyn and to enhance neurite outgrowth. J Cell Biol 169, 341–354, https://doi.org/10.1083/jcb.200409127 (2005).
Schmitt-Ulms, G. et al. Binding of neural cell adhesion molecules (N-CAMs) to the cellular prion protein. J Mol Biol 314, 1209–1225, https://doi.org/10.1006/jmbi.2000.5183 (2001).
Brown, D. R. et al. The cellular prion protein binds copper in vivo. Nature 390, 684–687, https://doi.org/10.1038/37783 (1997).
Walter, E. D., Stevens, D. J., Visconte, M. P. & Millhauser, G. L. The prion protein is a combined zinc and copper binding protein: Zn2+ alters the distribution of Cu2+ coordination modes. J Am Chem Soc 129, 15440–15441, https://doi.org/10.1021/ja077146j (2007).
Rayner, M. H. & Suzuki, K. T. A simple and effective method for the removal of trace metal cations from a mammalian culture medium supplemented with 10% fetal calf serum. Biometals 8, 188–192 (1995).
Vey, M. et al. Subcellular colocalization of the cellular and scrapie prion proteins in caveolae-like membranous domains. Proc Natl Acad Sci USA 93, 14945–14949 (1996).
Gorodinsky, A. & Harris, D. A. Glycolipid-anchored proteins in neuroblastoma cells form detergent-resistant complexes without caveolin. J Cell Biol 129, 619–627 (1995).
Hugel, B. et al. Modulation of signal transduction through the cellular prion protein is linked to its incorporation in lipid rafts. Cell Mol Life Sci 61, 2998–3007, https://doi.org/10.1007/s00018-004-4318-2 (2004).
Mahal, S. P. et al. Prion strain discrimination in cell culture: the cell panel assay. Proc Natl Acad Sci USA 104, 20908–20913, https://doi.org/10.1073/pnas.0710054104 (2007).
Bosque, P. J. & Prusiner, S. B. Cultured cell sublines highly susceptible to prion infection. J Virol 74, 4377–4386 (2000).
Vorberg, I., Raines, A., Story, B. & Priola, S. A. Susceptibility of common fibroblast cell lines to transmissible spongiform encephalopathy agents. The Journal of infectious diseases 189, 431–439, https://doi.org/10.1086/381166 (2004).
Nishida, N. et al. Successful transmission of three mouse-adapted scrapie strains to murine neuroblastoma cell lines overexpressing wild-type mouse prion protein. J Virol 74, 320–325 (2000).
Rubenstein, R. et al. Demonstration of scrapie strain diversity in infected PC12 cells. J Gen Virol 73(Pt 11), 3027–3031, https://doi.org/10.1099/0022-1317-73-11-3027 (1992).
Kimberlin, R. H. & Walker, C. A. Evidence that the transmission of one source of scrapie agent to hamsters involves separation of agent strains from a mixture. J Gen Virol 39, 487–496, https://doi.org/10.1099/0022-1317-39-3-487 (1978).
Race, R. & Chesebro, B. Scrapie infectivity found in resistant species. Nature 392, 770, https://doi.org/10.1038/33834 (1998).
Race, R., Raines, A., Raymond, G. J., Caughey, B. & Chesebro, B. Long-term subclinical carrier state precedes scrapie replication and adaptation in a resistant species: analogies to bovine spongiform encephalopathy and variant Creutzfeldt-Jakob disease in humans. J Virol 75, 10106–10112, https://doi.org/10.1128/JVI.75.21.10106-10112.2001 (2001).
Berry, D. B. et al. Drug resistance confounding prion therapeutics. Proc Natl Acad Sci USA 110, E4160–4169, https://doi.org/10.1073/pnas.1317164110 (2013).
Bolognesi, M. L. & Legname, G. Approaches for discovering anti-prion compounds: lessons learned and challenges ahead. Expert Opin Drug Discov 10, 389–397, https://doi.org/10.1517/17460441.2015.1016498 (2015).
Shinkai, H. et al. 4-Aminoquinolines: novel nociceptin antagonists with analgesic activity. J Med Chem 43, 4667–4677 (2000).
Yamada, H., Nakamoto, H., Suzuki, Y., Ito, T. & Aisaka, K. Pharmacological profiles of a novel opioid receptor-like1 (ORL(1)) receptor antagonist, JTC-801. Br J Pharmacol 135, 323–332, https://doi.org/10.1038/sj.bjp.0704478 (2002).
Korth, C., May, B. C., Cohen, F. E. & Prusiner, S. B. Acridine and phenothiazine derivatives as pharmacotherapeutics for prion disease. Proc Natl Acad Sci USA 98, 9836–9841, https://doi.org/10.1073/pnas.161274798 (2001).
Doh-Ura, K., Iwaki, T. & Caughey, B. Lysosomotropic agents and cysteine protease inhibitors inhibit scrapie-associated prion protein accumulation. J Virol 74, 4894–4897 (2000).
Ahn, M. et al. Pharmacokinetics of quinacrine efflux from mouse brain via the P-glycoprotein efflux transporter. PLoS One 7, e39112, https://doi.org/10.1371/journal.pone.0039112 (2012).
Lambert, D. G. The nociceptin/orphanin FQ receptor: a target with broad therapeutic potential. Nat Rev Drug Discov 7, 694–710, https://doi.org/10.1038/nrd2572 (2008).
Giles, K., Olson, S. H. & Prusiner, S. B. Developing Therapeutics for PrP Prion Diseases. Cold Spring Harb Perspect Med 7, https://doi.org/10.1101/cshperspect.a023747 (2017).
Feraudet, C. et al. Screening of 145 anti-PrP monoclonal antibodies for their capacity to inhibit PrPSc replication in infected cells. J Biol Chem 280, 11247–11258, https://doi.org/10.1074/jbc.M407006200 (2005).
Spinner, D. S. et al. CpG oligodeoxynucleotide-enhanced humoral immune response and production of antibodies to prion protein PrPSc in mice immunized with 139A scrapie-associated fibrils. Journal of leukocyte biology 81, 1374–1385, https://doi.org/10.1189/jlb.1106665 (2007).
Pankiewicz, J. et al. Clearance and prevention of prion infection in cell culture by anti-PrP antibodies. Eur J Neurosci 23, 2635–2647, https://doi.org/10.1111/j.1460-9568.2006.04805.x (2006).
Doolan, K. M. & Colby, D. W. Conformation-dependent epitopes recognized by prion protein antibodies probed using mutational scanning and deep sequencing. J Mol Biol 427, 328–340, https://doi.org/10.1016/j.jmb.2014.10.024 (2015).
Acknowledgements
We thank Olga Sharma, Theresa Hundt, and Rachana Subedi for expert technical support, Dr. David Westaway and Dr. Debbie McKenzie for providing ScN2a and SMB cells, Walker Jackson for providing the mouse-adapted RML and the hamster-adapted 263 K scrapie strains. We thank Dr. Jean Manson and the Roslin TSE Resource Centre for providing the mouse-adapted scrapie strains 87V, 22A, 22L, ME7, and 79A. We thank Dr. Philip Denner and his team at the DZNE’s core facility for Lab Automation Technologies for compound spotting. We thank Dr. Elmar Endl and his team at the University of Bonn’s Flow Cytometry Core Facility for technical support with FACS analysis and sorting. This work was supported financially by DZNE core funding to G.T.
Author information
Authors and Affiliations
Contributions
G.T. conceived the experiments. K.A.W., A.S., M.E.B. and P.P.R. conducted the experiments. K.A.W., A.S., M.E.B., P.P.R. and G.T. analysed the results. K.A.W. and G.T. wrote and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wüsten, K.A., Reddy, P.P., Smiyakin, A. et al. A Bioluminescent Cell Assay to Quantify Prion Protein Dimerization. Sci Rep 8, 14178 (2018). https://doi.org/10.1038/s41598-018-32581-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-32581-1
- Springer Nature Limited