Abstract
Prion diseases are caused by the propagation of misfolded cellular prion proteins (PrPs). A completely prion disease-resistant genotype, V127M129, has been identified in Papua New Guinea and verified in transgenic mice. To disclose the structural basis of the disease-resistant effect of the G127V mutant, we determined and compared the structural and dynamic features of the G127V-mutated human PrP (residues 91–231) and the wild-type PrP in solution. HuPrP(G127V) contains α1, α2 and α3 helices and a stretch-strand (SS) pattern comprising residues Tyr128-Gly131 (SS1) and Val161-Arg164 (SS2), with extending atomic distances between the SS1 and SS2 strands, and a structural rearrangement of the Tyr128 side chain due to steric hindrance of the larger hydrophobic side chain of Val127. The extended α1 helix gets closer to the α2 and α3 helices. NMR dynamics analysis revealed that Tyr128, Gly131 and Tyr163 underwent significant conformational exchanges. Molecular dynamics simulations suggest that HuPrP(G127V) prevents the formation of stable β-sheets and dimers. Unique structural and dynamic features potentially inhibit the conformational conversion of the G127V mutant. This work is beneficial for understanding the molecular mechanisms underlying the complete resistance of the G127V mutant to prion disease and for developing new therapeutics for prion disease.
Similar content being viewed by others
Introduction
Prion diseases, the notorious transmissible spongiform encephalopathies (TSEs), are infectious and fatal central nervous system (CNS) degenerative diseases in some mammals1. In humans, prion diseases manifest as a variety of clinical symptoms: Creutzfeldt-Jakob diseases (CJD, including sporadic, iatrogenic, variant, and familial/genetic CJD)2,3,4,5, Gerstmann-Sträussler-Scheinker syndrome (GSS)2,6, fatal familial insomnia (FFI)2,5,7 and Kuru2,8,9. These diseases are caused by the propagation of the insoluble scrapie isoform of the prion protein (PrPSc), a β-sheet-rich form, which is originated from the normal cellular prion protein (PrPC) through conformational conversion and is resistant to proteinases2,10,11. The α-helical PrPC is encoded by the endogenous prion protein gene (PRNP)1 and acts as an agonistic ligand of the Adgrg612. To date, more than 50 pathogenic point mutations of PRNP have been associated with a variety of prion diseases13,14,15. Fortunately, two frail sCJD-resistant mutants, V209M13,16 and E219K17,18,19, have been clinically observed, and their disease-resistance mechanisms have been addressed. Furthermore, a completely prion disease-resistant mutant G127V has also been identified in the bodies of Papua New Guinea’s Fore tribe and verified in transgenic mice studies20,21. However, the underlying disease-resistance mechanisms remain elusive.
Protein structure and dynamics primarily decide function. Structural biology is used extensively to exploit the molecular mechanisms of protein function. Previous work has shown that wild-type (WT) human prion protein (HuPrP) contains an N-terminal random coil (residues 23–124) and a C-terminal globular domain (residues 125–231)22 associated with many pathogenic and protective mutations13,14,15. The three-dimensional (3D) structure of the C-terminal domain in the WT protein is comprised of three α-helices (α1: 144–154, α2: 173–194, α3: 200–228), two short β-strands (β1: 128–131, β2: 161–164) and a disulfide bridge (Cys179-Cys214) even though under different pH conditions16,22,23,24. Almost all the HuPrP mutants adopt similar 3D structures to the WT protein16,22,23,24,25,26,27,28,29.
Previous 15N backbone dynamic studies of WT HuPrP show that the rigid globular C-terminal core has lower 15N longitudinal relaxation rates, higher 15N transverse relaxation rates, and much more positive {1H}-15N heteronuclear steady-state nuclear Overhauser effects (NOEs) than the N-terminal random coil23,29,30,31,32. Conformational exchanges on the μs-ms timescale were exhibited by the residues at or near the short anti-parallel β-sheet30 but were too small to be detected by Carr-Purcell-Meiboom-Gill (CPMG) relaxation dispersion (RD) experiments29,30,31. Due to the dramatic flexibility of the long N-terminal coil of the prion protein, the conventional model-free approach cannot be used to analyse NMR dynamics data29,30,32.
Based on the crystal structure of the WT HuPrP, 3D domain swapping with an intermolecular β-sheet may be an important step in the conversion of PrPC to PrPSc33,34. However, growing evidence has shown that the β1- and β2-strands, especially the β1-strand, and the interchain between them might trigger PrPSc propagation34,35,36. A series of β1-strands can form steric zippers with different packing geometries depending on the presence of either the Met129 or Val129 PRNP alleles35,36, even though the monomeric structures of the two genotypes do not show striking differences34,37,38. Importantly, molecular dynamics (MD) simulations confirmed that the slight protective heterozygosity of 129MV assembled extremely unstable intermolecular β-sheets due to the Val129 side chain, conflicting with spatially adjacent residues39,40,41,42. The different genotypes of residue 129 could cause the different stabilities of the β-sheet43,44,45, the different conformational conversion of the N-terminal flexible segment46, and the intrinsic conformational heterogeneity of the α1 helix47. For instance, D178N/M129 associated with FFI5,7 and D178N/V129 related to fCJD5,7 showed different intermolecular tetramers in crystal structures34, different dynamic features43,44,48 and different rates of the conversion to amyloid fibrils which were larger than WT in vitro39.
Furthermore, the frailly protective mutant HuPrP(V209M) against sCJD changes the geometric packing of the α1 and α3 helices and impairs the tendency of amyloid fibre formation13,16. The V209M mutation decreased the fibrillization rate relative to WT in vitro16. Quite differently, although the V210I mutant alters the geometry of the α2 and α3 helices27,49, it is fully susceptible to gCJD50. In addition, another frailly protective mutant against sCJD, HuPrP(E219K)17,18,19, has a slightly altered 3D structure, changed backbone dynamics, and redistributed surface electrostatic potentials23. The mutated Lys219 residue exhibits one of the highest 15N R2 rates, indicative of decreased backbone flexibility or/and increased conformational exchange on the μs-ms timescale23. The incompatible structures and dynamics of the heterozygous 219EK mutant potentially prevent hetero-dimerization51, which resists the development of sCJD17,18,19,23,51. Interestingly, even though the E200K mutant also redistributed the surface electrostatic potentials, it still leads to gCJD19,26,27,52. Thus, whether or not the alterations in the geometric packing of the α helices or the surface electrostatic potentials resist disease requires further studies.
In the completely disease-resistant G127V mutant, Val127 was observed exclusively on a Met129 PRNP allele21. In transgenic mice experiments, heterozygous 127 GV mice were able to resist Kuru and all of the CJDs except vCJD. Moreover, the protein expression ratios of the Gly127 to Val127 genotypes greatly affected the ability to resist prion disease. More significantly, homozygous 127VV mice were entirely resistant to all the prion diseases21. To identify the molecular mechanisms underlying the significant disease-resistant effect of the G127V mutant, we determined solution structures of both the recombinant HuPrP (residues 91–231) with the V127M129 genotype (termed HuPrP(G127V)) and the WT HuPrP with a G127M129 genotype (termed WT HuPrP) at pH 4.522. We performed NMR dynamics analysis and implemented MD simulations on the determined structures for the two proteins. We observed several significant differences in the structural and dynamic properties of the two proteins. These results provide novel insight into the molecular mechanisms of the disease-resistant effect of the G127V mutant. This work may be of benefit to both a mechanistic understanding of prion propagation and the development of effective therapeutics.
Results
Resonance assignments of HuPrP(G127V) and WT HuPrP
The well-dispersed 1H-15N HSQC spectra illustrate that both HuPrP(G127V) and WT HuPrP adopt well-folded structures (Figs 1 and S1). All resonance assignments were verified by 3D 15N-edited NOSEY-HSQC and 13C-edited NOESY-HSQC spectra. Overall, 91% resonances were assigned for the two proteins. In total, for the 136 backbone N-H resonances of HuPrP (residues 91–231) (141 residues minus 5 prolines), 127 and 128 resonances were unambiguously assigned to HuPrP(G127V) (BMRB ID: 27259) and WT HuPrP (BMRB ID: 27264), respectively. Backbone N-H resonances could not be obtained for Gly94, Arg164, Asp167, Glu168, Tyr169, Ser170, Asn171 and Phe175. Notably, the amine resonance of Tyr128 disappeared from the HSQC spectrum of HuPrP(G127V), but was visible as an isolated peak in that of WT HuPrP (Figs 1 and S1). The 1H-15N HSQC spectra between the mutant and WT proteins illustrate that the G127V mutant introduced notably changed chemical shifts for Met129, Val161, and Tyr162, with distinct peak broadening for Gly131 (Figs 1 and S1).
Solution Structures of HuPrP(G127V) and WT HuPrP at pH 4.5
Based on the resonance assignments and experimentally conformational restraints, we determined the solution structures of HuPrP(G127V) (PDB ID: 5YJ4) and WT HuPrP (PDB ID: 5YJ5) at pH 4.5 and 298K. The complete structural statistics are summarized in Table S1.
Similar to WT HuPrP, the solution structure of HuPrP(G127V) (Fig. 2a,b) consisted of the N-terminal flexible segment (Gln91-Gly124) and the C-terminal structural core (Leu125-Ser231) containing three α helices (α1: Asp144-Arg156, α2: Gln172-Lys194, α3: Glu200-Arg228) and a disulfide bond (Cys179-Cys214). The backbone root-mean-square deviation (RMSD) between the average structures of the mutant and WT proteins was 2.27 Å (Fig. S2). The G127V mutant formed a stretch-strand (SS) pattern with two segments (SS1: Tyr128-Gly131; SS2: Val161-Arg164), while the WT protein formed a stable β-sheet with two strands (β1: Tyr128-Gly131; β2: Val161-Arg164). Compared with WT HuPrP, HuPrP(G127V) exhibited unique structural characteristics (Fig. S2a,d), including the following: (I) a smaller distance between the α1 helix and α2/α3 helices; (II) a larger curvature of the α1-SS2 loop (Tyr157-Gln160); (III) a more flexible SS2-α2 loop (Pro165-Asn171) (backbone RMSD of 1.28 Å in the mutant vs. 0.88 Å in WT HuPrP); and (IV) a bigger bend in the α2 helix such that its C-terminal is closer to the N-terminal end of the α3 helix.
Structural analysis of HuPrP(G127V) and WT HuPrP. (a,b) Cartoons of the 20 lowest-energy conformers for the mutant and WT proteins. (c,d) Details of the structural alteration at Tyr128 resulting from the G127V mutation (showing only dHε/Tyr128-Hγ/Ile182 and dHε/Tyr128-Hγ/Gln186 < 5.0 Å). (e,f) Distances (dαα, dαN, dNN < 5.0 Å) between the two strands (stretch-strands for G127V, β-strands for WT). (g,h) Local conformational alterations caused by the change of the dihedral angle psi(N-Cα-C-N) in Tyr157. These results demonstrate that the C-terminal structural cores of the mutant and WT proteins adopt different conformations.
In HuPrP(G127V), the hydrophilic side chain of Tyr128 is rotated sharply so that its dihedral angle, chi (N-Cα-Cβ-Cγ), is reduced from 90° to 0° in WT HuPrP. This structural rearrangement might be introduced by the steric hindrance of the relatively larger hydrophobic side chain of Val127. This rotation pushes the phenyl ring of Tyr128 away from Ile182 and brings it closer to Gln186 (Fig. 2c,d). Although the HNMet129-HNTyr163 distance is identical for both structures (2.6 ± 0.1 Å), the HαLeu130-HαTyr162 distance is larger in the mutant (3.7 ± 0.2 Å vs. 2.2 ± 0.2 Å), as shown in Fig. 2e,f. The backbone atomic distances between SS1 and SS2 or between β1 and β2 are summarized in Table S2.
In the mutant protein, the C-terminal end of the α1 helix is extended to Arg156 (Figs 2a,b and S2a,b). In this configuration, the Arg156 side chain is closer to Thr190 and Thr191 at the C-terminal end of the α2 helix (Fig. S2c,d). In the α1-SS2 loop of HuPrP(G127V), the dihedral angle psi(N-Cα-C-N) of Tyr157 is nearly 60°, causing the pyrrolidine of Pro158 to retroflex approximately 180° (Fig. 2g,h). This alteration changes the atomic distances between Tyr157, Pro158, Val209 and Val210. Tyr157 becomes close to two α3-located residues, Val209 and Val210, and Pro158 moves away from Val209 and Val210. The retroflexion of the Pro158 side chain increases the curvature of the α1-SS2 loop in the mutant protein compared with the α1-β2 loop (His155-Gln160) in the WT (Fig. 2g,h). Additionally, atomic distances between SS2 and the disulfide bridge in the mutant are shorter than those between β2 and the disulfide bridge in the WT (Fig. S2g,h).
Furthermore, the G127V mutant also leads to a redistribution of the surface electrostatic potentials of the protein (Fig. S3). HuPrP(G127V) exhibits neutral potentials near residues Gly126-Ser135, while WT HuPrP shows positive potentials in this segment, except for Met129-Gly131. Additionally, compared with the WT protein, the mutant displays more positive potentials in the region near Arg146 and Arg151 of the α1 helix, and more negative potentials on the N-termini of the α1 and α3 helices.
The two different structures were calculated from their own different NOESY restraints originated from 3D 15N-edited NOESY-HSQC and 13C-edited NOESY-HSQC spectra (Fig. S4, Table S3). Several 1H-1H NOE peaks were missing, and many peaks were weaker in the SS segments from the HuPrP(G127V) than in those from the WT HuPrP (Fig. S4, Table S3). Furthermore, the structural differences were validated by backbone amide residual dipolar couplings (RDCs) measured from 2D 1H-15N IPAP-HSQC spectra (Figs S5 and S6). As indicated by the Q-values, the experimental RDCs from HuPrP(G127V) fitted better with the HuPrP(G127V) structure (Q = 0.532) than those with the WT HuPrP structure (Q = 0.798) and vice versa for the RDCs of the WT HuPrP (G127V vs WT: 0.856 vs 0.564)(Figs S5 and S6). In addition, the differences were confirmed with H/D exchanges based on 2D 1H-15N Fast-HSQC experiments (Figs S7 and S8). Remarkably, the amide proton of Gly131 in the SS segments from the HuPrP(G127V) was exchanged completely with D2O and became invisible in the HSQC spectrum than in those from the WT HuPrP (Figs S7 and S8). However, the amide protons of Met154 and His155 in the extended α1 helix from the HuPrP(G127V) became more stable than in those from the WT HuPrP (Figs S7 and S8).
Backbone amide relaxation analysis
To compare the dynamic features of the HuPrP(G127V) and WT HuPrP backbones, we performed a series of NMR relaxation experiments to obtain 15N longitudinal relaxation rates (R1), 15N transverse relaxation rates (R2) and {1H}-15N heteronuclear steady-state NOEs ({1H}-15N NOEs) at two magnetic field strengths, 14.10 T and 19.97 T (Fig. 3). A total of 106 and 112 backbone amide resonances were used to analyse the dynamic features of the mutant and WT proteins.
For HuPrP(G127V), residues in the N-terminal flexible segment did not show distinct differences in the R1 and R2 rates between the two magnetic fields but exhibited more negative {1H}-15N NOEs at 14.10 T than those at 19.97 T. In contrast, the residues in the C-terminal structural core, except for those in the α2-α3 loop (Gly195-Thr199) and the C-terminus (Gly229-Ser231), displayed significant differences in the R1 and R2 rates and similar {1H}-15N NOEs values (>0.6) between the two magnetic fields. The average R1 rate at 14.10 T was larger than that at 19.97 T (1.1 s−1 vs. 0.8 s−1). All residues, except for Gly131 and Gln172 in the C-terminal structural core, displayed R2 rates varying between 13.0 s−1 and 30 s−1 for both magnetic fields, with slightly higher values at 19.97 T. Furthermore, residues in the α2-α3 loop and the C-terminus showed larger R1 rates and smaller R2 rates as well as smaller {1H}-15N NOEs. Overall, WT HuPrP showed R1 rates, R2 rates and {1H}-15N NOEs roughly similar to HuPrP(G127V) for the two magnetic fields (Fig. 3). Both proteins exhibited larger differences in the R2/R1 ratios between 14.10 T and 19.97 T.
Interestingly, the G127V mutant showed distinctly changed R2 rates for the residues located in the SS1 and SS2 segments. The R2 rates of the SS1-located residue Gly131 in the mutant protein were 38.5 s−1 at 14.10 T and 44.7 s−1 at 19.97 T, which were much larger than those in the WT (24.6 s−1 at 14.10 T and 33.4 s−1 at 19.97 T). Moreover, Gly131 also displayed significantly different R2/R1 ratios between the two proteins. Furthermore, the R2 rates of the SS2-located residue Tyr163 in the mutant protein were slightly larger than those in the WT (14.10 T: 21.8 s−1 vs. 20.3 s−1; 19.97 T: 27.9 s−1 vs. 25.4 s−1). However, the R2 rates for the SS1-located residues Met129 and Leu130 in the mutant protein were extremely similar to those in the WT. Regrettably, the relaxation data of the SS2-located residues Val161 and Tyr162 were not suitable for relaxation analysis because of resonance overlapping in the mutant protein.
As expected, the G127V mutation more or less altered the R2 rates of the α2 helix residues. Although the R2 rate of Gln172 at 14.10 T was almost identical for both the mutant and WT proteins (28.5 s−1 vs. 27.6 s−1), this value at 19.97 T was smaller in the mutant than that in the WT (34.4 s−1 vs. 37.4 s−1). Furthermore, Ile182 in the mutant displayed slightly increased R2 rates compared to that in the WT protein (24.9 s−1 vs. 22.9 s−1 at 14.10 T; 29.4 s−1 vs. 27.5 s−1 at 19.97 T). In addition, the R2 rate of Gln186 in the mutant was much larger than that in the WT protein (21.6 s−1 vs. 16.4 s−1 at 14.10 T; 21.8 s−1 vs. 17.9 s−1 at 19.97 T). These alterations might be caused by the rotation of the Tyr128 side chain, as described above.
The G127V mutation also changed the R2 rates of the residues located within the α3 helix. Because of the retroflexion of the Pro158 pyrrolidine, as described above, the R2 rates of Val209 and Val210 subtly fluctuated at 14.10 T and were markedly disturbed at 19.97 T. Compared with the WT protein, the mutant showed slightly larger R2 rates for the two residues at 14.10 T (Val209: 24.6 s−1 vs. 23.8 s−1; Val210: 24.6 s−1 vs. 23.6 s−1) and displayed smaller R2 rates at 19.97 T (Val209: 23.8 s−1 vs. 25.1 s−1; Val210: 23.5 s−1 vs. 28.2 s−1) (Fig. 3).
Reduced spectral density mapping
To explicitly explore the internal motion of the amide backbone, we calculated the reduced spectral density functions at three frequencies, J(0), J(ωN) and J(0.87ωH), based on experimentally derived 15N relaxation data for both HuPrP(G127V) and WT HuPrP (Fig. 4). For HuPrP(G127V), the J(0) values of the N-terminal flexible segment and the C-terminus were less than 2.5 ns/rad for both magnetic fields. However, the C-terminal structural core displayed J(0) values varying from 5.0 ns/rad to 10.0 ns/rad. The α3 helix exhibited higher J(0) values than the α1 and α2 helices, but the α2-α3 loop displayed relatively smaller J(0) values than the α1 and α2 helices. Moreover, the N-terminal flexible segment showed J(ωN) values scattering from 0.05 ns/rad to 0.35 ns/rad at the two magnetic fields, but the C-terminal structural core exhibited J(ωN) values fluctuating near 0.27 ± 0.03 ns/rad at 14.10 T and 0.20 ± 0.03 ns/rad at 19.97 T. Furthermore, the N-terminal flexible segment showed J(0.87ωH) values between 0.014 ns/rad and 0.045 ns/rad at 14.10 T, which changed to 0.011 ns/rad and 0.026 ns/rad at 19.97 T. The C-terminal structural core displayed J(0.87ωH) values varying near 0.006 ns/rad at 14.10 T and 0.003 ns/rad at 19.97 T (Fig. 4). On the whole, compared with HuPrP(G127V), WT HuPrP did not show distinctly different J(0), J(ωN), J(0.87ωH) values or trends.
A comparison of the reduced spectral density functions between HuPrP(G127V) and WT HuPrP. All spectral densities were calculated from the corresponding backbone dynamics parameters (R1, R2, {1H}-15N NOEs), which were measured at magnetic field strengths of 14.10 T (red for G127V, violet for WT) and 19.97 T (blue for G127V, olive for WT).
Compared with WT HuPrP, HuPrP(G127V) showed much larger J(0) values for the SS1-located residue Gly131 (14.5 ns/rad vs. 9.2 ns/rad at 14.10 T; 16.7 ns/rad vs. 12.5 ns/rad at 19.97 T). The J(0) values for Tyr163 located in the SS2 of the mutant were only subtly larger than those in the WT protein (8.1 ns/rad vs. 7.5 ns/rad at 14.10 T; 10.4 ns/rad vs. 9.5 ns/rad at 19.97 T), similar to the R2 rate for Tyr163, which was slightly higher in the mutant than that in the WT protein. These results suggest that the two residues in the mutant underwent slow conformational fluctuations.
Gln172, located in the α2 helix of the mutant protein, displayed almost the same J(0) value as that of the WT protein at 14.10 T (10.7 ns/rad vs. 10.3 ns/rad) but exhibited a smaller J(0) value than that of the WT protein at 19.97 T (12.8 ns/rad vs. 14.0 ns/rad). The anomalous alteration in the J(0) values might have been caused by significant conformational fluctuations. Furthermore, the mutant showed subtly different J(0) values from the WT protein for several residues, including the following: Ile182 (14.10 T: 9.3 ns/rad vs. 8.5 ns/rad; 19.97 T: 11.0 ns/rad vs. 10.2 ns/rad), Gln186 (14.10 T: 8.0 ns/rad vs. 6.0 ns/rad; 19.97 T: 8.1 ns/rad vs. 6.6 ns/rad), Val209 (14.10 T: 9.2 ns/rad vs. 8.8 ns/rad; 19.97 T: 8.9 ns/rad vs. 9.3 ns/rad) and Val210 (14.10 T: 9.2 ns/rad vs. 8.9 ns/rad; 19.97 T: 8.8 ns/rad vs. 10.5 ns/rad). For these residues, the trends in the J(0) values (Fig. 4) were similar to those of the R2 rates (Fig. 3).
Relaxation dispersion measurements
To compare in detail the dynamic features between HuPrP(G127V) and WT HuPrP, especially for Gly131 and Tyr163 with large J(0) values, we performed CPMG RD experiments at pH 4.5 on both the mutant and WT proteins (1.0 mM) at two magnetic field strengths (14.10 T and 19.97 T). The resulting individual interconversion rates (kex) are shown in Table S4.
For HuPrP(G127V), Gly131 and Tyr163 in the SS1 and SS2 segments displayed kex rates of 1295 ± 122 s−1 and 2842 ± 186 s−1, respectively (Fig. 5, Table S4). Notably, residue Tyr128 disappears from the 1H-15N HSQC spectrum because of peak broadening that may be caused by conformational exchange. The relaxation dispersion data from the Val161 and Tyr162 residues were not suitable for the CPMG RD analysis because of resonance overlapping. Interestingly, Met129 and Leu130 did not display observable conformational fluctuations (Fig. S9, Table S4). Gln172 and Gln186, located in the α2 helix, exhibited significant conformational exchanges, with kex rates of 3171 ± 302 s−1 and 2143 ± 328 s−1, respectively (Fig. 5, Table S4).
A comparison of μs-ms timescale conformational exchanges between HuPrP(G127V) and WT HuPrP. All CPMG RD experiments were conducted at magnetic field strengths of 14.10 T (red for G127V, violet for WT) and 19.97 T (blue for G127V, olive for WT). The individual interconversion rates (kex) were mapped onto the 3D structures of the mutant and WT proteins. Gly131, Tyr163, Gln172 and Gln186 in the mutant are coloured light blue, TextView blue, blue and marine, respectively. Gln172 and Ile182 from the WT protein are coloured smudge and chartreuse, respectively. The colours of the spheres on the 3D structures show the changing tendency from bright to dark, corresponding to the declining tendency of kex for the related residues.
For WT HuPrP, Gln172 in the nearby the α2 helix displayed conformational exchange with a kex rate of 1815 ± 281 s−1 (Fig. 5; Table S4). Interestingly, Ile182 showed significant conformational exchange (kex = 1009 ± 486 s−1), but Gln186 exhibited negligible conformational exchange (Fig. 5, Table S4), although they are both located in the α2 helix. Furthermore, all residues in the β-sheet displayed insignificant conformational exchanges, excluding the unassigned Arg164 (Fig. S9, Table S4).
Additionally, the α3 helix in both the mutant and WT protein displayed substantial magnetic field strength-dependent conformational fluctuations on the μs-ms timescale (Fig. S10, Table S4). For example, two residues located in the α3 helix, Met205 and Thr216, exhibited significant conformational exchanges at 19.97 T rather than at 14.10 T.
Molecular dynamics simulations
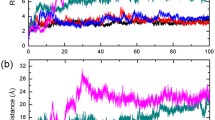
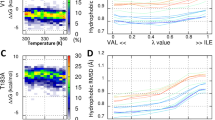
To further disclose the differences in dynamic structural properties between HuPrP(G127V) and WT HuPrP, we performed MD simulations based on the identified protein structures (Fig. 6 and S11). By analysing secondary structure elements and the geometric relationship of residues Leu125-Asp167, we summarized five primary distinctions between the mutant and WT proteins: the SS1 and SS2 segments rarely form a β-sheet in the mutant, instead, two β-strands always formed a stable β-sheet in the WT during the entire MD simulation (Fig. 6a); the α1 helix in the mutant is extended compared with that in the WT protein (Fig. 6a); the Tyr128 side chain adopts either the “mediate” or “out” conformation in the mutant instead of the “in” conformation found in the WT protein (Fig. 6b,c); the dynamic distance between the mass centres of Val127 and Pro165 in the mutant is smaller than that of Gly127 and Pro165 in the WT protein (Fig. 6d); the dihedral angle psi(N-Cα-C-N) of Tyr157 is approximately 60° for the mutant but is approximately 180° in the WT protein (Fig. 6e). These distinct dynamic properties might derive from the difference in hydrophobicity between Val127 and Gly127. Compared with glycine, valine is more hydrophobic and tends to be near the hydrophobic Pro165 in the SS2-α2 loop, as supported by the 3D structures of the mutant and WT proteins. This spatial alteration might introduce alterations in the conformation of Tyr128 and Tyr157 and the feasibility of β-sheet formation.
MD simulations of HuPrP(G127V) and WT HuPrP. (a) Time evolutions of the secondary structure elements of both proteins. (b) Time evolutions of the side chain dihedral angle chi(N-Cα-Cβ-Cγ) of Tyr128. The Tyr128 side chain can adopt three orientations: “out” - pointing to the solvent, “in” - pointing to the protein, and “mediate” - an orientation between “out” and “in”. (c) Several unique residues are identified in the 3D structures of the two proteins. (d) Time evolutions of the centroid distances between Gly127/Val127 and Pro165. (e) Time evolutions of the backbone dihedral angle psi(N-Cα-C-N) of Tyr157.
More meaningfully, the G127V mutant induces intramolecular steric hindrance in the relatively larger Val127 side chain, leading to a striking structural rearrangement and conformational alternation of the Tyr128 side chain. The MD simulations suggest that the orientation of the Tyr128 side chain directly determined the feasibility of the intermolecular dimerization. For the “exposed” case (either the “mediate” conformation or “out” conformation) in HuPrP(G127V), the steric hindrance closely associated with the Tyr128 side chain potentially prevents the monomeric prion protein from forming intermolecular interactions and might thus prohibit prion dimerization. In contrast, for the “buried” case (the “in” conformation) in WT HuPrP, the Tyr128 side chain likely does not reduce the feasibility of intermolecular dimerization (Fig. 6c). Hence, the mutation-induced structural rearrangement and dramatic conformational exchange of the Tyr128 side chain might be unfavourable for the dimerization and conformational conversion of HuPrP(G127V).
Discussion
Prion disease pathogenesis is closely associated with the conformational conversion of prion proteins from PrPC to PrPSc. The α2 and α3 helices33,53,54, octarepeats55, the N-terminal flexible segment56, and the glycophosphatidylinositol (GPI) anchor57 contained in PrPs might be involved in conformational conversions58,59,60,61. Moreover, conformational conversion is triggered at the two β-strands, the α1 helix, the α2 helix, the β1-α1 loop, the α1-β2 loop, and the β2-α2 loop34,35,62,63,64,65,66,67. Notably, the more stable β-structure is formed by the segment of the N-terminus (residues 120–144), the earlier stages of misfolding are caused by43,68,69 in MD, and a relatively short β-sheet core (residues 112–139) is capable of seeding the conversion to fibrils in vitro70. Nevertheless, the molecular mechanism underlying the disease-resistant effect of the G127V mutation still remains elusive. To reveal the molecular mechanisms, we determined the solution structures of both the HuPrP(G127V) and WT HuPrP under identical experimental conditions. We then analysed the backbone dynamics using 15N relaxation experiments and conducted MD simulations for both proteins. We focused primarily on the dynamic structural properties of the two SS segments and adjacent regions, including intramolecular interactions between SS1 and SS2, SS1/SS2 and α2, SS1/SS2 and α3, α1/α1-SS2 loop and α3, and SS2-α2 loop/α2 and α3.
The primary structural distinction between HuPrP(G127V) and WT HuPrP22,33 or other pathogenic mutants16,23,25,26,27,28 is that HuPrP(G127V) extends atomic distances between SS1 and SS2, increases the solvent accessibility surface of SS1-located residues (Figs S5 and S6), and exhibits significant μs-ms timescale conformational fluctuations at Tyr128, Gly131 and Tyr163. These properties indicate that the SS region is more flexible than the β-sheet and is not prone to conversion to a stable β-sheet conformation. Moreover, the striking structural rearrangement and alternate conformation of the Tyr128 side chain potentially induces the intermolecular steric hindrance effect, prevents the formation of intermolecular hydrogen bonds and prohibits prion protein dimerization. Notably, our result is fundamentally different from a previously published result, which suggested that the intermolecular steric hindrance was closely associated with the bulky sidechain of Val12771. The previous MD simulation work was based on the modelled structures of the G127V mutant using the solution structure of WT HuPrP (125–228) determined at pH 7.0 (PDB ID: 1HJN) and the crystal structure of the β1-strand fragment (PDB ID: 4TUT) as the templates71. Additionally, HuPrP(G127V) also alters the local electrostatic potential distribution near the SS1 and SS2 segments to influence potentially electrostatic interactions.
Previous studies suggest that pathogenic and protective mutants of PrPs have similar structures and dynamics29,30,35,36,42,44. However, our results confirmed that the structural and dynamic alterations caused by G127V are tremendously different from the changes caused by Met129, Val129 or any other known mutants29,30,35,36,42,44. Furthermore, as previously hypothesized, the β-sheet in the prion protein, and especially the β1-strand, might be the cornerstone on which prion protein aggregation is triggered34,35,36,63,64,66. For instance, D178N/M129 and F198S form intermolecular antiparallel four-strand β-sheets based on β1-strands in crystal structures34, and the β1-strand fragments form a steric zipper conformation35,36. However, HuPrP(G127V) possesses flexible SSs with structural rearrangement and conformational fluctuations, rearrangement and alternate conformation of the Tyr128 side chain as well as surface electrostatic potential redistribution that destroys the prion protein aggregation trigger and prohibits prion protein fibrillization.
On the other hand, HuPrP(G127V) and WT HuPrP have similar atomic distances between Met129 and Tyr163 (Table S2), similar H/D exchanges of Met129, Leu130, Val161, Tyr162 and Tyr163 (Figs S7 and S8), and similar dynamic properties for Met129 and Leu130 (Figs 3, 4 and S9). The similar structural and dynamic properties between the SSs in the mutant protein and the β-sheet in the WT protein imply that HuPrP(G127V) might partially reserve the structural and dynamic properties of the β-sheet in the WT protein via the SS pattern.
Regarding the intramolecular interactions between the SSs and the α2/α3 helices in the G127V mutant, we found that (I) the G127V mutant changes the orientation of the Tyr128 side chain and leads to different conformational exchanges for Ile182 and Gln186 (Table S4); (II) the mutation-induced steric hindrance effect between the side chains of Val127 and Arg164 pushes the Arg164 side chain close to Asp178, strengthens the electrostatic interaction between Arg164 and Asp178 (Fig. S2e,f) and enhances the hydrophobic interaction between Val127 and Pro165 (Fig. 6d); (III) the G127V mutant positions two SS2-located residues, Tyr163 and Arg164, slightly closer to Cys179 (Fig. S2g,h). These structural alternations reveal that the G127V mutation changes the local circumstances around the SSs and α2/α3 regions in HuPrP(G127V), which are distinctly different from those in the WT protein and several other HuPrP mutants such as D178N34. These unique structural features of HuPrP(G127V) potentially reduce the feasibility of prion protein aggregation54.
Furthermore, distinguishing structural features are also identified in the regions around the α1 helix and the α1-SS2 loop and the α2 and α3 helices in HuPrP(G127V), which may be responsible for the prion disease-resistance effects of the G127V mutant. Overall, the G127V mutation extends the α1 helix and induces the retroflexion of the Pro158 pyrrolidine, thus increasing the curvature of the α1-SS2 loop. These structural alterations potentially prevent the unwinding of the α1 helix. As previously suggested, the α1 helix could be converted to the β-strand to form fibrils via a despiralization process63,64. In the G127V mutant, the atomic distances between Tyr157, Pro158, Val209 and Val210 changed (Fig. 2g,h) and introduced magnetic field strength-dependent fluctuations in the R2 and R2/R1 rates of Val209 and Val210 (Fig. 3). These alterations might correspondingly change the local environment of the Val210 mutable site (the V210I mutant is associated with fCJD27,49), and may promote the protective effect of the G127V mutant.
Additionally, the extended α1 helix, the bent α2 helix, and the α3 helix are packed more compactly in HuPrP(G127V) than those in the WT protein. The unique geometric packing in the G127V mutant is similar to the protective packing in the V209M mutant16 and might slow the initial fibrillization rate in a manner similar to that in HuPrP(V209M)16 and the G126V mutant of the mouse prion protein (moPrP)72. The moPrP(G126V) is equivalent to HuPrP(G127V), slows initial fibril growth and increases the critical concentration72. The compact geometric packing might also change the local environment of the α2-α3 loop near the α1 helix. Note that the fCJD-associated F198S34 mutant occurs in the α2-α3 loop. In the G127V mutant, the surface electrostatic potential distribution on the region encompassing the α1 and α3 helices is diametrically distinct from those in the WT and the fCJD-associated E200K mutant26. The alterations of HuPrP(G127V) in both the geometric packing and electrostatic potential distribution combined with the close atomic distances between SS2 and the disulfide bridge, might prohibit rearrangement of the disulfide bridge, aggregation and fibrillization as previously published results16,33,54,72.
Compared with the WT HuPrP, the SS2-α2 loop (Pro165-Asn171) of the HuPrP(G127V) exhibits more flexibility. The G127V mutation allows Met166 at the SS2-α2 loop to be closer to Tyr218, which is located in the α3 helix (Fig. S2i,j). In HuPrP(G127V), Gln172, next to the SS2-α2 loop, undergoes a more significant conformational exchange than that in the WT HuPrP (Table S4). These results indicate that the SS2-α2 loop has dynamic structural features distinct from the β2-α2 loop (Pro165-Gln172), which is probably correlated to the susceptibility to prion disease65. The unique dynamic structural properties of the SS2-α2 loop might contribute to the prion disease resistance of the G127V mutant as well.
Astonishingly, the α3 helix in HuPrP(G127V) showed R2/R1 ratios that were dramatically different from those of the WT HuPrP at 19.97 T (Fig. 3). Moreover, the α3 helix exhibited varying J(0) values, similar to the R2/R1 ratios. In addition, Met205 and Thr216 in both proteins experienced slow conformational exchange, which was observable only at 19.97 T. Unexpectedly, the α3-located Glu219 in both proteins displayed large R2 rates and J(0) values but did not exhibit observable conformational exchanges (Fig. S10, Table S4). Furthermore, Glu219 in HuPrP(G127V) showed the dynamic property, distinct from HuPrP(E219K)23. Thus, the dramatically altered dynamic structural properties relevant to the α3 helix could potentially influence the intermolecular interactions of the prion protein with the so called “protein X”73,74.
Besides, our fibrillization experiments showed that HuPrP(G127V) had significantly slower initial fibril growth than WT HuPrP. The measured lag phases were 61 ± 2 h for HuPrP(G127V) and 25 ± 2 h for WT HuPrP as showed in Fig. S12. Moreover, the mixing samples of WT HuPrP and HuPrP(G127V) (at a mixing ratio of 1:1) exhibited a slower fibrillization rate than WT HuPrP but faster than HuPrP(G127V). The measured lag phase was 47 ± 2 h for the mixing sample. These kinetic analyses are similar to the quantitative comparison of moPrP(G126V) and WT moPrP72. These unique dynamic structural features might be responsible for the prion disease-resistance effect of the G127V mutant20,21. As expected, the further study of the exploitation of the structural and dynamic features of the GSS-associated mutant G131V75,76 (GSS), which was confirmed to enhance the stability of the β-sheet and drive conformational conversion by MD simulation77,78, would greatly help to address the crucial role of the SS1 segment in conformational conversion and propagation.
Summarily, we performed solution structure determinations, NMR dynamics analysis and MD simulations on both HuPrP(G127V) and WT HuPrP. We addressed the G127V mutation-induced significant distinct alterations in structural and dynamic properties in detail. The G127V mutation extends atomic distances between the SS1 and SS2 segments and enhances the conformational exchange of the two strands, leading to the formation of the SS pattern instead of the stable β-sheet. The relatively larger hydrophobic side chain of Val127 introduces steric hindrance and a striking structural rearrangement in the Tyr128 side chain. Additionally, the G127V mutation also subtly alters the geometric stacking of the three α helices. These structural and dynamic features might prevent the SS1 (Tyr128-Gly131) and SS2 (Val161-Arg164) segments and adjacent regions from being converted into a stable β-sheet under certain circumstances. Furthermore, the steric hindrance effect of the rearrangement of the Tyr128 side chain, together with the dramatic conformational alternation, could potentially prohibit the prion protein intermolecular interaction and dimerization, and thus inhibit prion protein aggregation and fibrillization. Moreover, HuPrP(G127V) had significantly slower initial fibril growth than WT HuPrP. Although more researches are required to clarify completely the molecular mechanisms of the prion disease-resistance of HuPrP(G127V), our results provide several important evidences regarding the differences in structure and dynamics between HuPrP(G127V) and WT HuPrP. These structural and dynamic differences substantially contribute to the different conversion of monomer to dimer in MD and of monomer to fibril in fibrillization between the two proteins. This work may be helpful for mechanistically understanding the pathogenesis of prion diseases and for developing effective drugs against prion diseases.
Methods
NMR sample preparation
Recombination of the pET30a plasmids without any tag bearing the DNA of the WT HuPrP (residues 91–231 with G127M129) was prepared as previously described79,80,81. The recombination plasmids for HuPrP(G127V) (residues 91–231 with the genotype of V127M129) were cloned by PCR using site-directed mutagenesis. The forward primer used in the PCR was: 5′-AGTGGTGGGGGGCCTTGGCGTTTACATGCTGGGAA-3′ and the reverse primer used was: 5′-ATGGCACTTCCCAGCATGTAAACGCCAAGGCCCCCCA-3′. The uniformly labelled protein was overexpressed in E. coli Bl21(DE3) grown in M9 medium. 15NH4Cl was added to the M9 medium to prepare 15N-labelled proteins and both 15NH4Cl and 13C6-glucose were added to prepare 13C/15N-labelled proteins. After the cells were sonicated and the lysates were centrifuged, the inclusion bodies were denatured in 6 M guanidine hydrochloride and refolded by dialysing against the NMR buffer (20 mM NaOAc, 0.02% NaN3, pH 4.5) as previously described79,80,81. Thereafter, the protein was purified in NMR buffer through size exclusion chromatography with Superdex-75 on an ÄKTA FPLC system (GE Healthcare). Finally, the protein solution was concentrated to approximately 0.5 mM with 10% D2O (v/v).
NMR spectroscopy
To perform backbone and side chain resonances and to determine the solution structures of the HuPrP(G127V) and WT HuPrP proteins, we recorded a suite of 2D/3D heteronuclear NMR spectra at 25 °C on a Bruker Avance III 850-MHz spectrometer (magnetic field strength is 19.97 T) with a 1H/13C/15N triple-resonance cryogenic probe (TCI). These 3D NMR spectra included HNCACB, CBCA(CO)NH, HNCA, HNCOCA, HNCO, HN(CA)CO, HBHA(CO)NH, H(CCCO)NH, CC(CO)NH and (H)CCH-TOCSY. A mixing time of 120 ms was used for both 15N-edited NOSEY-HSQC and 13C-edited NOESY-HSQC experiments. All NMR spectra were processed by NMRPipe software82 and analysed with CARA software83.
Structure calculations
Distance constraints were generated from the 1H-1H NOEs of both 13C and 15N-labelled NOESY-HSQC spectra. Dihedral angle restraints were obtained based on chemical shifts of the backbone atoms including HN, Hα, Cα, Cβ, C(O), and N using the TALOS+ programme84. The 3D structures were calculated and refined with the XPLOR-NIH package85. Then, the qualities of the calculated structures were evaluated by the PROCHECK programme86. Ultimately, the 20 lowest-energy conformers were selected as representative solution structures of each protein. The tertiary structures of the proteins were displayed using PyMOL87 and MOLMOL88.
Backbone dynamics
Backbone amide relaxation measurements
A complete set of backbone amide R1, R2 and {1H}-15N NOEs spectra were acquired on 15N labelled samples on both a Bruker Avance III 850-MHz (magnetic field strength of 19.97 T with a TCI cryogenic probe) and a Bruker Avance III 600-MHz (magnetic field strength of 14.10 T with a BBO cryogenic probe) spectrometer at 25 °C and pH 4.5. All spectra were recorded with 1024 × 128 complex points. R1 values were measured from 2D 1H-1N HSQC spectra with relaxation delays of 10, 50, 100 (×2), 250, 500, 800, 1200 (×2), 1600 and 2000 ms. R2 values were determined with relaxation delays of 16.32, 32.64 (×2), 48.96, 65.28, 81.6, 97.92, 114.24(×2), 130.56, 146.88, and 163.2 ms. The repeated spectra were used for experimental error analysis. {1H}-15N NOEs were obtained by recording spectra with a 1H pre-saturation of 3 s plus a 2-s relaxation delay and without a pre-saturation of a 5-s relaxation delay. All NMR spectra were processed using NMRPipe software82 and analysed using CcpNmr software89. Peak heights were used to represent peak intensities. Standard errors of the fitted parameters were obtained by Monte Carlo simulations. For HuPrP(G127V), 21 residues were unavailable for NMR dynamics analysis due to resonance overlapping: 101, 121, 127, 146, 147, 155, 159, 161, 162, 173, 177, 178, 183, 185, 187, 189, 190, 204, 213, 218 and 231. Thus, 106 backbone amide resonances were used to analyse the dynamic behaviour of the molecular backbone. For WT HuPrP, 16 residues could not be used for NMR dynamics analysis due to resonance overlapping: 101, 140, 146, 149, 154, 155, 159, 173, 177, 185, 189, 190, 204, 213, 218, and 231. Thus, 112 backbone amide resonances were employed to analyse the dynamic behaviour of the molecular backbone.
Reduced spectral density mapping
Reduced spectral density mapping is usually employed to characterize the internal motions of the N-H bonds with the assumption that J(0.87ωH) is approaching J(ωH + ωN) and J(ωH + ωN) at high frequencies90. Therefore, the values of the relaxation rates R1, R2 and {1H}-15N NOEs are taken to map the spectral density using the following formula:
where
and
where μ0 is the permeability of the free space; h is Planck’s constant; γN and γH are the gyromagnetic ratios of 15N and 1H, respectively; ωN and ωH are the Larmor frequencies of 15N and 1H, respectively; rNH is the length of the N–H bond; and \({\Delta }\sigma \) = \({\sigma }_{\parallel }-{\sigma }_{\perp }\) is the chemical shift anisotropy for 15N. The calculations were implemented using Mathematica software91.
Relaxation dispersion measurements
Single quantum CPMG RD experiments were performed on the same NMR instruments described above (850 MHz at 19.97 T with a TCI cryogenic probe, 600 MHz at 14.10 T with a BBO cryogenic probe). The CPMG RD spectra were recorded on 15N-edited HuPrP(G127V) and WT HuPrP proteins at 25 °C and pH 4.5 using a constant relaxation time of 40 ms and under thirteen νCPMG values of 0, 100(×2), 200, 300, 400, 500, 600, 700(×2), 800, 900, and 1000 Hz. All spectra were recorded with complex points of 1024 × 128. The νCPMG is defined by the following formula92:
Here, τcp is the time between refocusing pulses during the CPMG pulse train. We used the following equation92 to calculate the effective transverse relaxation rates, \({R}_{2}^{eff}\)
where Tcp is the constant transverse relaxation time and I(νCPMG) and I0 are the intensity with or without different νCPMG. The RD of \({R}_{2}^{eff}\) relies on νCPMG if the residue undergoes conformational exchange at the μs-ms timescale. All spectra were processed in NMRPipe82 and the integrals of the peaks were obtained in NMRFAM-Sparky93. The dispersion data were fitted with a Carver-Richards two-state exchange model94 in NESSY software95. Similar to the 15N backbone dynamics analysis, overlapping amide resonances were not used for CPMG RD analysis.
Molecular Dynamics Simulations
All MD simulations were performed with the AMBER99SB96 force field in AMBER1297. All systems were solvated within a cubic box of TIP3P98 water molecules by extending 10 Å from the protein surface. The initial coordinates and topology files were generated using the tleap programme contained in AMBER12. First, energy minimizations were performed to relax the solvent and optimize the system. Then, each system was gradually heated from 0 to 300 K under the NVT ensemble for 100 ps and another 100 ps of NPT ensemble MD simulation was performed at 300 K and a target pressure of 1.0 atm. Finally, a 100 ns MD simulation under the NVT ensemble was performed for each model. The system temperature was controlled by the Langevin thermostat method. During the MD simulations, all hydrogen-containing bonds were constrained using the SHAKE algorithm99. A cut-off of 12 Å was set for both the van der Waals and electrostatic interactions. The DSSP algorithm was employed to assign the secondary structure of the protein100.
Residual Dipolar Couplings
Initially, both 15N-labeled HuPrP(G127V) and WT HuPrP were dissolved in H2O buffer (90% H2O, 10% D2O, 20 mM NaOAc, 0.02% NaN3, pH 4.5) to a final concentration of 0.4 mM. As reference spectra, 2D 1H-15N IPAP-HSQC spectra were recorded at 25 °C on a Bruker Avance III 600-MHz spectrometer (magnetic field strength of 14.10 T with a triple-resonance TCI cryogenic probe) at the University of Science and Technology of China. All spectra were recorded with complex points of 1024 × 400. Then, the two proteins were dissolved in C12E5/n-hexanol alignment media101. The final concentration of C12E5 was 3% (r = 0.96)101. 2D 1H-15N IPAP-HSQC spectra were recorded under the same experimental conditions. All data were processed on NMRPipe82, analysed on NMRFAM-Sparky93 and fitted on PALES102. The Q-value was fitted by PALES, which is normally used to assess the agreement between the experimental RDCs and calculated RDCs based on the structure102,103. When fitted using the PALES program, the experimental RDCs were just from the residues of the C-terminal structural core minus the overlapping resonance, as described above.
Amide Hydrogen/Deuterium Exchange
Both 15N labelled HuPrP(G127V) and WT HuPrP were initially dissolved in H2O buffer (90% H2O, 10% D2O, 20 mM NaOAc, 0.02% NaN3, pH 4.5). As reference spectra, 2D Fast-1H-15N HSQC104 were recorded at 25 °C on a Bruker Avance III 850-MHz spectrometer (magnetic field strength of 19.97 T with a triple-resonance TCI cryogenic probe). All spectra were recorded with complex points of 1024 × 128. Through buffer exchange with centrifugal filter devices (Amicon® Ultra 3 K device) at 2,555 × g and 4 °C for 3 h, the proteins were re-dissolved in equal volumes of D2O buffer (99.9% D2O, 20 mM NaOAc, 0.02% NaN3, pH 4.5). Then, 2D Fast-1H-15N HSQC spectra were recorded on the re-dissolved proteins as amide proton exchange spectra under the same experimental conditions. All data were processed on Topspin 3.2 (Bruker) and analysed in CcpNmr89. This approach allowed the quantitative analysis of peak intensity decreases caused by the mutation but could not be used to measure amide protection factors for the protein105.
Accession codes
Chemical shift data were deposited in the Biological Magnetic Resonance Data Bank (http://www.bmrb.wisc.edu) under accession numbers 27259 for HuPrP(G127V) and 27264 for WT HuPrP. The atomic coordinates were deposited in the Protein Data Bank under the accession codes 5YJ4 for HuPrP(G127V) and 5YJ5 for WT HuPrP.
References
Prusiner, S. B. Novel proteinaceous infectious particles cause scrapie. Science 216, 136–144 (1982).
Prusiner, S. B. Prions. Proc Natl Acad Sci USA 95, 13363–13383 (1998).
Masters, C. L., Gajdusek, D. C. & Gibbs, C. J. Jr. The familial occurrence of Creutzfeldt-Jakob disease and Alzheimer’s disease. Brain 104, 535–558 (1981).
Fornai, F. et al. A hypothesis on prion disorders: are infectious, inherited, and sporadic causes so distinct? Brain Res Bull 69, 95–100 (2006).
Goldfarb, L. G. et al. Fatal familial insomnia and familial Creutzfeldt-Jakob disease disease phenotype determined by a DNA polymorphism. Science 258, 806–808 (1992).
Hsiao, K. et al. Linkage of a prion protein missense variant to Gerstmann-Sträussler syndrome. Nature 338, 342–345 (1989).
Petersen, R. B., Parchi, P., Richardson, S. L., Urig, C. B. & Gambetti, P. Effect of the D178N mutation and the codon 129 polymorphism on the metabolism of the prion protein. J Biol Chem 271, 12661–12668 (1996).
Gibbs, C. J. Jr, Amyx, H. L., Bacote, A., Masters, C. L. & Gajdusek, D. C. Oral Transmission of Kuru, Creutzfeldt-Jakob Disease, and Scrapie to Nonhuman Primates. J Infect Dis 142, 205–208 (1980).
Field, E. J. The significance of astroglial hypertrophy in Scrapie, Kuru, Multiple Sclerosis and old age together with a note on the possible nature of the scrapie agent. Deutsche Zeitschrift fur Nervenheilkunde 192, 265–274 (1967).
Pan, K. M. et al. Conversion of alpha helices into beta sheets features in the formation of the scrapie prion protein. Proc Natl Acad Sci USA 90, 10962–10966 (1993).
Meyer, R. K. et al. Separation and properties of cellular and scrapie prion proteins. Proc Natl Acad Sci USA 83, 2310–2314 (1986).
Küffer, A. et al. The prion protein is an agonistic ligand of the G protein-coupled receptor Adgrg6. Nature 536, 464–468 (2016).
Beck, J. A. et al. PRNP allelic series from 19 years of prion protein gene sequencing at the MRC Prion Unit. Hum Mutat 31, E1551–1563 (2010).
Jeong, B. H. & Kim, Y. S. Genetic studies in human prion diseases. J Korean Med Sci 29, 623–632 (2014).
Giachin, G., Biljan, I., Ilc, G., Plavec, J. & Legname, G. Probing early misfolding events in prion protein mutants by NMR spectroscopy. Molecules 18, 9451–9476 (2013).
Kong, Q. et al. Thermodynamic stabilization of the folded domain of prion protein inhibits prion infection in vivo. Cell Rep 4, 248–254 (2013).
Petraroli, R. & Pocchiari, M. Codon 219 polymorphism of PRNP in healthy Caucasians and Creutzfeldt-Jakob disease patients. Am J Hum Genet 58, 888–889 (1996).
Shibuya, S., Higuchi, J., Shin, R. W., Tateishi, J. & Kimamoto, T. Codon 219 lys allele of prnp is not found in sporadic creutzfeldt-jakob disease. Ann Neurol 43, 826–828 (1998).
Seno, H. et al. New haplotype of familial creutzfeldt-jakob disease with a codon 200 mutation and a codon 219 polymorphism of the prion protein gene in a japanese family. Acta Neuropathol 99, 125–130 (2000).
Mead, S. et al. A novel protective prion protein variant that colocalizes with kuru exposure.pdf. N Engl J Med 361, 2056–2065 (2009).
Asante, E. A. et al. A naturally occurring variant of the human prion protein completely prevents prion disease. Nature 522, 478–481 (2015).
Zahn, R. et al. NMR solution structure of the human prion protein. Proc Natl Acad Sci USA 97, 145–150 (2000).
Biljan, I. et al. Structural basis for the protective effect of the human prion protein carrying the dominantnegative E219K polymorphism. Biochem J 446, 243–251 (2012).
Calzolai, L. & Zahn, R. Influence of pH on NMR structure and stability of the human prion protein globular domain. J Biol Chem 278, 35592–35596 (2003).
Ilc, G. et al. NMR structure of the human prion protein with the pathological Q212P mutation reveals unique structural features. PloS one 5, e11715 (2010).
Zhang, Y., Swietnicki, W., Zagorski, M. G., Surewicz, W. K. & Sonnichsen, F. D. Solution structure of the E200K variant of human prion protein. Implications for the mechanism of pathogenesis in familial prion diseases. J Biol Chem 275, 33650–33654 (2000).
Biljan, I. et al. Toward the molecular basis of inherited prion diseases: NMR structure of the human prion protein with V210I mutation. J Mol Biol 412, 660–673 (2011).
Kovac, V., Zupancic, B., Ilc, G., Plavec, J. & Curin Serbec, V. Truncated prion protein PrP226* - A structural view on its role in amyloid disease. Biochem Biophys Res Commun 484, 45–50 (2017).
Bae, S. H. et al. Prion proteins with pathogenic and protective mutations show similar structure and dynamics. Biochemistry 48, 8120–8128 (2009).
Viles, H. et al. Local structural plasticity of the prion protein. Analysis of NMR relaxation dynamics. Biochemistry 40, 2743–2753 (2001).
Kuwata, K., Kamatari, Y. O., Akasaka, K. & James, T. L. Slow Conformational Dynamics in the Hamster Prion Protein. Biochemistry 43, 4439–4446 (2004).
O’Sullivan, D. B. et al. Dynamics of a truncated prion protein, PrP(113-231), from (15)N NMR relaxation: order parameters calculated and slow conformational fluctuations localized to a distinct region. Protein Sci 18, 410–423 (2009).
Knaus, K. J. et al. Crystal structure of the human prion protein reveals a mechanism for oligomerization. Nat Struct Biol 8, 5 (2001).
Lee, S. et al. Conformational diversity in prion protein variants influences intermolecular beta-sheet formation. The EMBO journal 29, 251–262 (2010).
Yu, L., Lee, S. J. & Yee, V. C. Crystal Structures of Polymorphic Prion Protein beta1 Peptides Reveal Variable Steric Zipper Conformations. Biochemistry 54, 3640–3648 (2015).
Apostol, M. I., Sawaya, M. R., Cascio, D. & Eisenberg, D. Crystallographic studies of prion protein (PrP) segments suggest how structural changes encoded by polymorphism at residue 129 modulate susceptibility to human prion disease. J Biol Chem 285, 29671–29675 (2010).
Hosszu, L. L. et al. The residue 129 polymorphism in human prion protein does not confer susceptibility to Creutzfeldt-Jakob disease by altering the structure or global stability of PrPC. J Biol Chem 279, 28515–28521 (2004).
Owen, F., Poulter, M., Collinge, J. & Crow, T. J. Codon 129 changes in the prion protein gene in caucasians. Am J Hum Genet 46, 1215–1216 (1990).
Apetri, A. C., Vanik, D. L. & Surewicz, W. K. Polymorphism at Residue 129 Modulates the Conformational Conversion of the D178N Variant of Human Prion Protein 90–231. Biochemistry 44, 9 (2005).
Palmer, M. S., Dryden, A. J., Hughes, J. T. & Collinge, J. Homozygous prion protein genotype predisposes to sporadic Creutzfeldt-Jakob Disease. Nature 352, 340–342 (1991).
Collinge, J. Molecular neurology of prion disease. J Neurol Neurosurg Psychiatry 76, 906–919 (2005).
Fernandez-Borges, N. et al. Protective Effect of Val129-PrP against Bovine Spongiform Encephalopathy but not Variant Creutzfeldt-Jakob Disease. Emerg Infect Dis 23, 1522–1530 (2017).
Barducci, A. et al. Metadynamics Simulation of Prion Protein beta Structure Stability and the Early Stages of Misfolding. J Am Chem Soc 128, 2705–2710 (2006).
Paz, S. A., Vanden-Eijnden, E. & Abrams, C. F. Polymorphism at 129 dictates metastable conformations of the human prion protein N-terminal β-sheet. Chem Sci 8, 1225–1232 (2017).
Baskakov, I. et al. The presence of valine at residue 129 in human prion protein accelerates amyloid formation. FEBS Lett 579, 2589–2596 (2005).
Chen, W., van der Kamp, M. W. & Daggett, V. Structural and dynamic properties of the human prion protein. Biophys J 106, 1152–1163 (2014).
Guo, J., Ren, H., Ning, L., Liu, H. & Yao, X. Exploring structural and thermodynamic stabilities of human prion protein pathogenic mutants D202N, E211Q and Q217R. J Struct Biol 178, 225–232 (2012).
Barducci, A., Chelli, R., Procacci, P. & Schettino, V. Misfolding pathways of the prion protein probed by molecular dynamics simulations. Biophys J 88, 1334–1343 (2005).
Biljan, I., Ilc, G., Giachin, G., Plavec, J. & Legname, G. Structural rearrangements at physiological pH: nuclear magnetic resonance insights from the V210I human prion protein mutant. Biochemistry 51, 7465–7474 (2012).
Pocchiari, M. et al. A new point mutation of the prion protein gene in Creutzfeldt-Jakob disease. Ann Neurol 34, 802–807 (1993).
Jahandideh, S., Jamalan, M. & Faridounnia, M. Molecular dynamics study of the dominant-negative E219K polymorphism in human prion protein. J Biomol Struct Dyn 33, 1315–1325 (2015).
Gabizon, R. et al. Mutation in codon 200 in scrapie amyloid precursor gene linked to Creutzfeldt-Jakob disease in Sephardic Jews of Lybian and non-Lybian origin. Phil. Trans R Soc Lond B 343, 385–390 (1994).
Chakroun, N. et al. Decrypting Prion Protein Conversion into a beta-Rich Conformer by Molecular Dynamics. J Chem Theory Comput 9, 2455–2465 (2013).
Apostol, M. I., Perry, K. & Surewicz, W. K. Crystal structure of a human prion protein fragment reveals a motif for oligomer formation. J Am Chem Soc 135, 10202–10205 (2013).
Benetti, F. & Legname, G. New insights into structural determinants of prion protein folding and stability. Prion 9, 119–124 (2015).
DeMarco, M. L. & Daggett, V. From conversion to aggregation: Protofibril formation of the prion protein. Proc Natl Acad Sci USA 101, 2293–2298 (2004).
Johannes, Z. & Jill, E. G. Molecular dynamics simulation of human prion protein including both N-linked oligosaccharides and the GPI anchor. Glycobiology 10, 959–974 (2000).
Campos, S. R. R., Machuqueiro, M. & Baptista, A. Constant-pH Molecular Dynamics Simulations Reveal a β-Rich Form of the Human Prion Protein. J Phys Chem B 114, 12692–12700 (2010).
Zhang, J. & Zhang, Y. Molecular dynamics studies on the NMR and X-ray structures of rabbit prion protein. J Theor Biol 342, 70–82 (2014).
Cheng, C. J. & Daggett, V. Molecular dynamics simulations capture the misfolding of the bovine prion protein at acidic pH. Biomolecules 4, 181–201 (2014).
Ye, W., Wang, W., Jiang, C., Yu, Q. & Chen, H. Molecular dynamics simulations of amyloid fibrils: an in silico approach. Acta Biochim Biophys Sin (Shanghai) 45, 503–508 (2013).
Swietnicki, W., Morillas, M., Chen, S. G., Gambetti, P. & Surewicz, W. K. Aggregation and fibrillization of the recombinant human prion protein huPrP90-231. Biochemistry 39, 8 (2000).
Samson, A. O. & Levitt, M. Normal modes of prion proteins: from native to infectious particle. Biochemistry 50, 2243–2248 (2011).
Singh, J. & Udgaonkar, J. B. Molecular Mechanism of the Misfolding and Oligomerization of the Prion Protein: Current Understanding and Its Implications. Biochemistry 54, 4431–4442 (2015).
Caldarulo, E., Barducci, A., Wuthrich, K. & Parrinello, M. Prion protein beta2-alpha2 loop conformational landscape. Proc Natl Acad Sci USA 114, 9617–9622 (2017).
Abskharon, R. N. et al. Probing the N-terminal beta-sheet conversion in the crystal structure of the human prion protein bound to a nanobody. J Am Chem Soc 136, 937–944 (2014).
Bamdad, K. Sequence-dependent dynamical instability of the human prion protein: A comparative simulation study. J Biomol Struct Dyn, 1–11 (2017).
Wang, Y., Shao, Q. & Hall, C. K. N-terminal Prion Protein Peptides (PrP(120–144)) Form Parallel In-register β-Sheets via Multiple Nucleation-dependent Pathways. J Biol Chem 291, 22093–22105 (2016).
Wang, Y. & Hall, C. K. Seeding and Cross-seeding Fibrillation of N-terminal Prion Protein Peptides PrP(120–144). Protein Sci 27, 1304–1313 (2018).
Choi, J. K. et al. Amyloid fibrils from the N-terminal prion protein fragment are infectious. Proc Natl Acad Sci USA 113, 13851–13856 (2016).
Zhou, S., Shi, D., Liu, X., Liu, H. & Yao, X. Protective V127 prion variant prevents prion disease by interrupting the formation of dimer and fibril from molecular dynamics simulations. Sci Rep 6, 21804 (2016).
Sabareesan, A. T. & Udgaonkar, J. B. The G126V Mutation in the Mouse Prion Protein Hinders Nucleation-Dependent Fibril Formation by Slowing Initial Fibril Growth and by Increasing the Critical Concentration. Biochemistry 56, 5931–5942 (2017).
Kaneko, K. et al. Evidence for protein X binding to a discontinuous epitope on the cellular prion protein during scrapie prion propagation. Proc Natl Acad Sci USA 94, 10069–10074 (1997).
Perrier, V. et al. Dominant-negative inhibition of prion replication in transgenic mice. Proc Natl Acad Sci USA 99, 13079–13084 (2002).
Panegyres, P. K. et al. A new PRNP mutation (G131V) associated with Gerstmann-Sträussler-Scheinker disease. Arch Neurol 58, 1899–1902 (2001).
Race, B. et al. Familial human prion diseases associated with prion protein mutations Y226X and G131V are transmissible to transgenic mice expressing human prion protein. Acta Neuropathol Commun 6, 13 (2018).
Santini, S., Claude, J.-B., Audic, S. & Derreumaux, P. Impact of the tail and mutations G131V and M129V on prion protein flexibility. Proteins 51, 258–265 (2003).
Chen, W., van der Kamp, M. W. & Daggett, V. Diverse Effects on the Native β-Sheet of the Human Prion Protein Due to Disease-Associated Mutations. Biochemistry 49, 9874–9881 (2010).
Wen, Y. et al. Unique structural characteristics of the rabbit prion protein. J Biol Chem 285, 31682–31693 (2010).
Yu, Y. et al. Distinct effects of mutations on biophysical properties of human prion protein monomers and oligomers. Acta Biochim Biophys Sin (Shanghai) 48, 1016–1025 (2016).
Yu, Z. et al. Unique Properties of the Rabbit Prion Protein Oligomer. PloS one 11, e0160874 (2016).
Delaglio, F. et al. NMRPipe a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6, 277–293 (1995).
Keller, R. L. J. The Computer Aided Resonance Assignment Tutorial. (CANTINA Verlag, 2004).
Shen, Y., Delaglio, F., Cornilescu, G. & Bax, A. TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44, 213–223 (2009).
Schwieters, C., Kuszewski, J. & Mariusclore, G. Using Xplor–NIH for NMR molecular structure determination. Prog Nucl Magn Reson Spectrosc 48, 47–62 (2006).
Laskowski, R. A., Rullmannn, J. A. C., MacArthur, M. W. & Thornton, J. AQUA and PROCHECK-NMR programs for checking the quality of protein structures solved by NMR. J Biomol NMR 8, 477–486 (1996).
Schrodinger, LLC. The PyMOL Molecular Graphics System, Version 1.8 (2015).
Reto, K., Martin, B. & Wüthrich, K. MOLMOL: A program for display and analysis of macromolecular structures. J Mol Graph 14, 51–55 (1996).
Vranken, W. F. et al. The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59, 687–696 (2005).
Farrow, N. A., Zhang, O., Szabo, A., Torchia, D. A. & Kay, L. E. Spectral density function mapping using 15N relaxation data exclusively. J Biomol NMR 6, 153–162 (1995).
Spyracopoulos, L. A suite of Mathematica notebooks for the analysis of protein main chain 15N NMR relaxation data. J Biomol NMR 36, 215–224 (2006).
Mulder, F. A., Skrynnikov, N. R., Hon, B., Dahlquist, F. W. & Kay, L. E. Measurement of slow (micros-ms) time scale dynamics in protein side chains by (15)N relaxation dispersion NMR spectroscopy application to Asn and Gln residues in a cavity mutant of T4 lysozyme. J Am Chem Soc 123, 967–975 (2001).
Lee, W., Tonelli, M. & Markley, J. L. NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31, 1325–1327 (2015).
Carver, J. P. & Richards, R. E. A General Two-Site Solution for the Chemical Exchange Produced Dependence of T2 Upon the Carr-Purcell Pulse Separation. J Magn Reson 6, 89–105 (1972).
Bieri, M. & Gooley, P. R. Automated NMR relaxation dispersion data analysis using NESSY. BMC bioinformatics 12, 421–431 (2011).
Duan, Y. et al. A point-charge force field for molecular mechanics simulations of proteins based on condensedphase quantum mechanical calculations. J Comput Chem 24, 1999–2012 (2003).
Case, D. A. et al. Amber12. University of California, San Francisco (2012).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J Chem Phys 79, 926–935 (1983).
Ryckaert, J.-P., Ciccotti, G. & Berendsen, H. J. C. Numerical Integration of the Cartesian Equations of Motion of a System with Constrains Molecular Dynamics of n Alkanes. J Comput Phys 23, 327–341 (1977).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Rückert, M. & Otting, G. Alignment of Biological Macromolecules in Novel Nonionic Liquid Crystalline Media for NMR Experiments. J Am Chem Soc 122, 7793–7797 (2000).
Zweckstetter, M. & Bax, A. Prediction of Sterically Induced Alignment in a Dilute Liquid Crystalline Phase: Aid to Protein Structure Determination by NMR. J Am Chem Soc 122, 3791–3792 (2000).
Tolman, J. R. & Ruan, K. NMR Residual Dipolar Couplings as Probes of Biomolecular Dynamics. Chem Rev 106, 1720–1736 (2006).
Mori, S., Abeygunawardana, C., Johnson, M. O. & Vanzijl, P. C. M. Improved Sensitivity of HSQC Spectra of Exchanging Protons at Short Interscan Delays Using a New Fast HSQC (FHSQC) Detection Scheme That Avoids Water Saturation. J Magn Reson B 108, 94–98 (1995).
Laszlo, L. P. H. et al. Structural mobility of the human prion protein probed by backbone hydrogen exchange. Nat Struct Biol 6, 740–743 (1999).
Acknowledgements
This research was financially supported by the National Natural Science Foundation of China (Nos. 31670741, 31170717 and 31470034). The RDCs experiments were performed at the University of Science and Technology of China, and all other NMR experiments were performed at the High-Field Nuclear Magnetic Resonance Research Centre of Xiamen University. In particular, Zhen Zheng wants to thank the invaluable support from Jun Wan over the years. I love you.
Author information
Authors and Affiliations
Contributions
Z.Z., C.G., and D.L. devised all the experiments. Z.Z. and M.Z. prepared the samples. Z.Z. and L.F. implemented the NMR experiments and analysed the data. Z.Z. performed the structural calculations, dynamic analyses and fibrillization experiments. Y.W. executed the molecular dynamics simulation. Z.Z., R.M., and J.W. performed the RDC experiments and re-implemented the CPMG RD experiments. Z.Z., Y.W., C.G., H.Y. and D.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zheng, Z., Zhang, M., Wang, Y. et al. Structural basis for the complete resistance of the human prion protein mutant G127V to prion disease. Sci Rep 8, 13211 (2018). https://doi.org/10.1038/s41598-018-31394-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-31394-6
- Springer Nature Limited
This article is cited by
-
Prion protein gene mutation detection using long-read Nanopore sequencing
Scientific Reports (2022)
-
Homodimeric complexes of the 90–231 human prion: a multilayered computational study based on FMO/GRID-DRY approach
Journal of Molecular Modeling (2022)
-
The G127V variant of the prion protein interferes with dimer formation in vitro but not in cellulo
Scientific Reports (2021)
-
Zn(II) binding causes interdomain changes in the structure and flexibility of the human prion protein
Scientific Reports (2021)
-
1H, 13C, 15N backbone and side‐chain resonance assignments of the pathogenic G131V mutant of human prion protein (91–231)
Biomolecular NMR Assignments (2021)