Abstract
Due to organ shortage, clinicians are prone to consider alternative type of organ donors among them donors deceased after circulatory death (DCD). However, especially using these organs which are more prone to graft dysfunction, there is a need to better understand mechanistic events ocuring during ischemia phase and leading to ischemia/reperfusion injuries (IRI). The aim of this study is to provide a dynamic transcriptomic analysis of preclinical porcine model kidneys subjected to ischemic stress mimicking DCD donor. We compared cortex and corticomedullary junction (CMJ) tissues from porcine kidneys submitted to 60 min warm ischemia (WI) followed by 0, 6 or 24 hours of cold storage in University of Wisconsin solution versus control non-ischemic kidneys (n = 5 per group). 29 cortex genes and 113 CMJ genes were significantly up or down-regulated after WI versus healthy kidneys, and up to 400 genes were regulated after WI followed by 6 or 24 hours of cold storage (p < 0.05). Functionnal enrichment analysis (home selected gene kinetic classification, Gene-ontology-biological processes and Gene-ontology-molecular-function) revealed relevant genes implication during WI and cold storage. We uncovered targets which we will further validate as biomarkers and new therapeutic targets to optimize graft kidney quality before transplantation and improve whole transplantation outcome.
Similar content being viewed by others
Introduction
Transplantation remains the only efficient therapeutic option for end-stage renal diseases. However, this success led to a worldwide organ shortage, and only 30% of patients on the waiting list have access to an organ. This situation, combined with unavoidable demographic change in donor population, has led to the growing use of organs coming from marginal or “extended-criteria” donors, including deceased after circulatory death donors (DCD)1. However, organs coming from marginal donors are more prone to develop ischemia injuries, harmful during reperfusion for organ quality and outcome. Importantly, ischemia-reperfusion (IR) injuries (IRI) are correlated with delayed graft function rate and primary non function rate1. Hence the need to consider new strategies to improve organ preservation quality. Indeed, from cardiac death of the donor, through organ procurement and its cold storage until graft revascularistion in the recipient, ischemic injuries lay the groundwork for secondary lesions resulting in worsening the outcome. In transplantation, severe reperfusion injuries after transplantation are mostly caused by the initiation of ischemic lesions. However, the molecular mechanisms underlying severe ischemia are currently not fully determined and this knowledge gap limits the current effort to design better approach for organ preservation. While several solutions using technological or pharmacological improvements have been tested2, the lack of specific targets as well as the reduced number of available biomarkers are slowing down the development and transfer to the clinic. In addition, IRI- focusing studies mostly rely on the use of small animal models where the application of ischemia protocols similar to those used in clinic is challenging and/or studies are focused on a single pathway in a hypothesis-driven fashion. We propose here an open ended approach based on microarray technology to understand IRI occurring in DCD-like kidney submitted to warm-ischemia (WI) followed by cold ischemia (CS), using a pre-clinical porcine model with the main advantage that porcine and human kidneys are extremely similar in size, structure and function3. The specific objective of this study is to evaluate gene expression profile of kidney submitted to ischemic injury similarly to what is observed in clinic with kidneys coming from DCD donors i.e submitted to WI followed by cold ischemia. Indeed, we investigated differential gene expression patterns in kidneys after a period of WI followed or not by a cold storage (CS) of 6 h (WI + CS6h) or 24 h (WI + CS24h) versus control non-ischemic kidneys (Ctl) in a reproducible model. Identified gene expression profiles were submitted to functional enrichment analysis and a comprehensive bibliographic review was performed to understand the role of each marker in the biological serie of events occurring during ischemia. Our aim is to determine specific inhibited/activated genes which could become pharmacological target and define markers useful to evaluate organs before transplantation.
Results
Ischemia impact on global gene expression profile
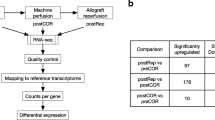
From microarray datasets, we generated heatmaps to depict mRNA expression between our groups and we submitted our data to Principal Component Analysis (PCA). Numbers in parentheses represent the percentage of total variance explained by the first and second principal components, explaining respectively 80.27% and 13.15% of the variability of our results for cortex (C) genes (Fig. 1A), and 59.55 and 23.77% of the variability of our results for CMJ genes (Fig. 1B), showing that our experimental groups have well distanced transcriptomic profiles. Moreover, for both cortex and CMJ genes, PCA shows that both experimental groups submitted to cold storage are clustering closely, and are well separated from the two others groups i.e control and WI groups, highlighting that the duration of the cold storage itself has moderate impact on the variability of our results and therefore on sample gene expression profiles.
To further decipher transcriptomic changes occurring during our ischemia treatments, we then focused on heatmaps-extracted differentially expressed genes between selected groups. Precisely, we found that the number of differentially expressed genes between different ischemia-treated groups and control group (Ctl) was: 29 cortex genes and 113 CMJ genes between 60 minutes of WI vs Ctl group, 1145 cortex genes and 456 CMJ genes between 60 minutes of WI followed by 6 hours of cold storage (WI + CS6h) vs Ctl group and 805 cortex genes and 485 CMJ genes between 60 minutes of WI followed by 24 hours of cold storage (WI + CS24h) vs Ctl group. Additionally, our heatmap results showed that 561 cortex genes and 462 CMJ genes were differentially expressed between WI and WI + CS6 groups, and 398 cortex genes and 446 CMJ genes were differentially expressed between WI and WI + CS24 groups (Supplementary Tables S1 and S2). The criteria for significance being |log2 fold change| ≥ 0.5 and a corrected p-value < 0.05.
Home-made panel gene expression dynamics
We extracted from heatmaps the kinetic expression of several relevant genes significantly differentially expressed and classed in categories having an important role in IRI (Figs 2–7). Heatmaps are available on Supplementary Figures S1–S10.
Kinetic expression of relevant genes from (A) “Mitochondria and redox state regulation” and (B) “Protein folding and proteasome” categories which are significantly differentially expressed from heatmaps. Results are expressed in mean ± SEM, **p < 0.01, ***p < 0.001 versus Ctl cortex, ##p < 0.01, ###p < 0.001 versus Ctl CMJ, ¥¥p < 0.01, ¥¥¥p < 0.001 versus WI Cortex, $$p < 0.001, $$$p < 0.001 versus WI CMJ.
Kinetic expression of relevant genes from “Inflammation and apoptosis” category which are significantly differentially expressed from heatmaps. Results are expressed in mean ± SEM, **p < 0.01, ***p < 0.001 versus Ctl cortex, ##p < 0.01, ###p < 0.001 versus Ctl CMJ, ¥¥p < 0.01, ¥¥¥p < 0.001 versus WI Cortex, $$p < 0.001, $$$p < 0.001 versus WI CMJ.
Kinetic expression of relevant genes from “Cell cycle, cellular differentiation and proliferation” category which are significantly differentially expressed from heatmaps. Results are expressed in mean ± SEM, **p < 0.01, ***p < 0.001 versus Ctl cortex, ##p < 0.01, ###p < 0.001 versus Ctl CMJ, ¥¥p < 0.01, ¥¥¥p < 0.001 versus WI Cortex, $$p < 0.001, $$$p < 0.001 versus WI CMJ.
Kinetic expression of relevant genes from (A) “Nucleus genes and transcriptional regulation” and (B) “Transporters” categories which are significantly differentially expressed from heatmaps. Results are expressed in mean ± SEM, **p < 0.01, ***p < 0.001 versus Ctl cortex, ##p < 0.01, ###p < 0.001 versus Ctl CMJ, ¥¥p < 0.01, ¥¥¥p < 0.001 versus WI Cortex, $$p < 0.001, $$$p < 0.001 versus WI CMJ.
Kinetic expression of relevant genes from “Metabolism regulation” category which are significantly differentially expressed from heatmaps. Results are expressed in mean ± SEM, **p < 0.01, ***p < 0.001 versus Ctl cortex, ##p < 0.01, ###p < 0.001 versus Ctl CMJ, ¥¥p < 0.01, ¥¥¥p < 0.001 versus WI Cortex, $$p < 0.001, $$$p < 0.001 versus WI CMJ.
Kinetic expression of relevant genes from “MAPK and GTPase activity” category which are significantly differentially expressed from heatmaps. Results are expressed in mean ± SEM, **p < 0.01, ***p < 0.001 versus Ctl cortex, ##p < 0.01, ###p < 0.001 versus Ctl CMJ, ¥¥p < 0.01, ¥¥¥p < 0.001 versus WI Cortex, $$p < 0.001, $$$p < 0.001 versus WI CMJ.
Functional enrichment analysis
The genes we identified as differentially expressed were further submitted to enrichement analysis based on Gene-Ontology Biological-Process (GO-BP) and Gene-Ontology Molecular-Function (GO-MF) categories (Gene Ontology Families). We detected up- and down-regulated categories in cortex tissues represented respectively in Supplementary Tables S3 and S5, and in CMJ represented respectively in Supplementary Tables S4 and S6 (p < 0.05). Each family contained one or more differentially expressed genes. Most significative GO-BP and GO-MF categories of genes identified by the functionnal enrichment analysis are available on Supplementary Figures S12.
Confirmation by Real-Time Polymerase Chain Reaction
In order to confirm microarray results, we chose genes which where differentially expressed in our analysis and performed RT-qPCR to validate our results (Supplementary Table S7). We found significant correlations between Agilent heatmaps and RT-qPCR results, for both cortex (spearman r = 0.84, p < 001) and CMJ (spearman r = 0.88, p < 001) genes (Supplementary Figure S11).
Discussion
While attempting to better understand ischemia mechanisms, we performed a high throughput transciptomic analysis of pig kidneys subjected to intense ischemia injury observed in DCD organ transplantation. Our model has significant advantage over other transcriptomic analyses of rodent kidneys submitted to warm-ischemia4,5, since these latters are far to be anatomically and physiologically similar to their human counterparts3. A clinical study of Naesens et al. reported transcriptomic analysis of human kidneys coming from DCD donors versus living donors at the end of preservation period, but they did not compare their data with control healthy kidneys, leading to difficulties to isolate consequences of ischemia phase itself6. In a similar study comparing the same type of samples, authors submitted raw data to Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis and highlighted several variability factors between samples such as cause of death, donor age, warm-ischemia time, cold ischemia duration and preservation solution7. We investigated differential gene expression patterns in each kidney after a period of WI followed by cold storage (WI + CS) versus Control non-ischemic kidneys (Ctl) in a reproducible model. Our experimental design allowed us to avoid the high variability reported in human studies, using yet a preclinical model of kidney transplantation highly transposable to clinic3.
Identified gene expression profiles were submitted to functional enrichment analysis and a comprehensive bibliographic review was performed to understand the role of each marker in the biological serie of events occurring during ischemia. We analyzed the kinetics of the most differentially expressed genes and families from GO-BP and GO-MF families and we hypothesize that these genes could play a key role in IRI, and are subdivided in the following 8 categories:
-
1.
“Mitochondria and redox state regulation” such as CYP2C42, CYP2C34, ACOX1, GSTA2, GSTT1, HMOX1, HMOX2, CDO1 and EPO (Fig. 2A). Mitochondria, involved in the redox state regulation, are the most important organel affected by ischemia. CYP2C42 and CYP2C34 are proteins of the Cytochromes P450 family containing heme as a cofactor. They usually are the terminal oxidase enzymes in electron transfer chains. Similarly to what we found, prolonged ischemia decrease cytochromes P450 isoforms levels in a rat kidney IRI model8. Acyl-CoA oxidase (ACOX) regulation may be a reflection of the loss of acyl-CoA oxidase activity9. Glutathione S-transferase Alpha (GSTA) and Theta (GSTT) enzymes are major actors of oxidative stress products detoxification. They are dowregulated in cold-ischemia, conforting the fact that the oxidative stress response is acting at the reperfusion time. Heme oxygenases (HMOX) are enzymes catalyzing heme degradation. HMOX1 encodes Heme oxygenase 1 (HO-1), an inducible isoform responding to hypoxia. We indeed observe HMOX1 upregulation in our study, likely trigerring heme degradation and IRI protection10, probably via the NRF2-AKT interactions11. HMOX2 encodes HO-2, a constitutive isoform that is expressed under homeostatic conditions. Cysteine dioxygenase (CDO1) is a non-heme iron enzyme catalyzing conversion of L-cysteine to cysteine sulfinic acid by dioxygen incorporation. Cysteine residues maintained and transduced redox signals in the mitochondria12,13. Erythropoietin (EPO), activated by HIF-1a transcription factor, is upregulated by the kidney in response to cellular hypoxia14. In our study, 30 min of WI seems to be too short to upregulate significantly cytoprotectives EPO and CDO1 RNAs.
-
2.
“Protein folding and proteasome” such as TMX4, PAAF1, PLK3, HSP70 and HSPH1 (Fig. 2B). Like mitochondria, the endoplasmic reticulum and the proteasome are important cellular compartiments altered by ischemia. In accordance with our results, oxidative stress induces mitochondrial dysfunction and protective unfolded protein response in epithelial cells, with upregulation of thioredoxin-related transmembrane protein 4 (TMX4) mRNA15, as well as lower expression of catalytic and structural subunits of the proteasome, as Proteasomal ATPase-associated factor 1 (PAAF1), contributes to decreased proteolysis16. Polo-like kinase 3 (PLK3), herein upregulated, is involved in the GO-BP family “regulation of proteasomal ubiquitin-dependent protein catabolic” at WI + CS6h vs Ctl (Supplementary Table S3). Heat shock protein 70 kDa (Hsp70) family play a major role in cell machinery for protein folding, helpful for stress protection17. Hsp105 (HSPH1) interacts with Cofilin-118 preventing the aggregation of denatured proteins in cells under severe stress where the ATP levels decrease markedly19. Similarly to other studies where ischemia activates protective mRNA transcripts for heat shock proteins in rat heart20, our results confort that HSP expression upregulation is protective against injury.
-
3.
“Inflammation and apoptosis” such as CD83, CCL2, CCL26, GATA3, TLR4, ZFAND5, SMAD6, BCL6, TRAF3, IER3 and TMEM14A (Fig. 3). Inflammation and apoptosis are important consequences of IR with large impact on graft function and outcome. Cluster of differentiation 83 (CD83) play a significant role in antigen presentation or cellular interactions following lymphocyte activation. The chemokine (C-C motif) ligand 2 (CCL2) is also referred to monocyte chemoattractant protein-1 (MCP1). CCL2 recruits monocytes, memory T cells and dendritic cells to inflammation sites related to tissue injury21. CCL26 is expressed on endothelial cell surface22 and inhibits CCL2 mediated response23. Here, CCL2/CCL26 expression ratio during cold storage is in favour to CCL2. GATA-3 is a transcriptional activator which binds to the T-cell receptor genes enhancer and is required for T-helper 2 (Th2) differentiation process following immune and inflammatory responses24. Toll-like receptor 4 (TLR4) plays a fundamental role in damage-associated molecular patterns (DAMPs) recognition and innate immunity activation. Upregulation/release of DAMPs molecule, exacerbates renal IRI by stimulating inflammatory and immune responses through TLR4 signaling pathway25,26. Thus, TLR4 RNA upregulation could be associated with innate immune response stimulation. Zinc finger, AN1-type domain-5 (ZFAND5, also called ZNF216), inhibits TNF, IL-1 and TLR4-induced NF-kappa-B activation27. Mothers against decapentaplegic homolog-6 (SMAD6) acts as negative mediator of TGF-β and BMP antiflammatory activity, preventing NF-kappa-B activation28. In our study, SMAD6 expression downregulation (except in CMJ WI + CS24h) is in favour of fibrosis and inflammatory pathways. B-cell lymphoma-6 protein (BCL6) modulates STAT-dependent Interleukin 4 (IL-4) responses of B cells. During cold storage, BCL6 upregulation leads to the differentiation of naive helper T cells in Follicular Helper T cells29. Our data suggest that upregulation of TNF receptor-associated factor-3 (TRAF3), could represent a novel mechanism for preserving the functional integrity of the endothelial monolayer30. This protein is involved in the signal transduction of CD40, a TNFR family member important for the activation of the immune response. Immediate early response-3 (IER3) is member of the NUPR1/RELB/IER3 survival pathway31. Nevertheless, IER3, which is here down-regulated during WI, plays a complex and to some extent contradictory role in cell cycle control and apoptosis32. Transmembrane protein-14A (TMEM14A) inhibits apoptosis via negative regulation of the mitochondrial outer membrane permeabilization involved in apoptotic signaling pathway33. Herein, except CD83 and IER3, inflammation and apoptosis genes were all regulated during cold ischemia without differential expression between cortex and CMJ. These markers expression are evidences of an important immune response enhancing due to ischemia-induced stress.
-
4.
“Cell cycle, cellular differentiation and proliferation” such as CEBPA, ADIPOQ, EGR1, COL1A1, ANXA3, ANGPTL4, ADAM9, VAV3, DYRK2, JUN, FOS, CYR61, OAS, CDC42EP1 and KLF4 (Fig. 4). Phenotypic alterations due to ischemia resulted in cellular differentiation and proliferation pathway regulation. CCAAT/enhancer-binding protein alpha (CEBPA) is a transcription factor which coordinates proliferation arrest and differentiation34. To modulate lipogenesis, CEBPA interacts and transcriptionally synergizes with SREBP1 (Sterol regulatory element-binding transcription factor-1) in promoter activation of specific lipogenic target genes35. To regulate gluconeogenesis, CEBPA seems to act as FOXO1 coactivator accessing to Adiponectin (ADIPOQ) promoter36. ADIPOQ regulates glucose regulation and fatty acid oxidation37. Early growth response protein 1 (EGR1) is a transcriptional regulator38 playing a role in cell differentiation, survival, proliferation and death. It regulates the transcription of numerous target genes, and thereby regulates the response to growth factors, DNA damage and ischemia39. Here, upregulation of EGR1 and CEBPA greatly increase plasminogen activator inhibitor-1 transcriptional response in hypoxia independently of HIF1-alpha40. Collagen alpha-1(I) chain (COL1A1) is a major constituent of the connective tissue, interacting with platelet-derived growth factor-B (PDGFB) and von Willebrand factor41, which act respectively on angiogenesis and hemostasis. Annexin A3 (ANXA3) is a inhibitor of phospholipase A2, cleaves inositol 1,2-cyclic phosphate to form inositol 1-phosphate and also possesses anti-coagulant properties. ANXA3 is expressed on healthy epithelial cells42 and neutrophils granules43. Studies show that ANXA3 enhances hypoxia-inducible factor-1 (HIF-1) transactivation activity and acts as angiogenic factor inducing VEGF production through HIF-1 pathway44. Angiopoietin-like-4 (ANGPTL4), involved in glucose homeostasis and lipid metabolism regulation, inhibits endothelial cells proliferation, migration, tubule formation and reduces vascular leakage45. Disintegrin and metalloproteinase domain-containing protein 9 (ADAM9) plays a role on angiogenesis46 and mediates cell-cell and cell-matrix interactions47. VAV3 is a guanine nucleotide exchange factor for Rho family GTPases that activate pathways leading to actin cytoskeletal rearrangements and integrin-mediated signalling. VAV3 is regulated during cell cycle48 and promots angiogenesis. Dual specificity tyrosine-phosphorylation-regulated kinase 2 (DYRK2) presumed to be involved in cellular growth and/or development49. The transcription factor c-FOS is a proto-oncogene and hence referred to be an immediate early gene expressed after stimuli50. It encodes a 62 kDa protein, which forms heterodimer with c-JUN, resulting in the formation of Activator Protein-1 (AP1, also called JUN)51 and is involved in cell proliferation, differentiation and survival associated with hypoxia and angiogenesis52. Alltogether, upregulation of COL1A1, ADAM9, VAV3, DYRK2, JUN and c-FOS as well as downregulation of ANGPTL4 are in favour of an angiogenic response due to ischemia.The interferon-induced 2′-5′-oligoadenylate synthetases (OAS)53 play a role in cellular processes such as apoptosis, cell growth, differentiation and gene regulation. Cell division control protein-42 homolog effector protein-1 (CDC42EP1), encodes CDC42 which have an essential role in organism survival, growth and development54. CDC42 activity in primary cells displayed a slow proliferation rate by modulating the JNK-mediated apoptotic machinery55. Kruppel-like factor-4 (KLF4) is involved in the regulation of proliferation, differentiation and apoptosis. KLF4 is highly expressed in non-dividing cells and its overexpression induces cell cycle arrest56. KLF4 mediates p53-dependent G1/S cell cycle arrest in response to DNA damage57, preventing entry into mitosis58. In our study, it was demonstrated that CDC2 (other cell division control protein) kinase measurements showed an inverse correlation between CDC2 kinase activities and KLF4 levels58. Most of “cell cycle, cellular differentiation and proliferation” genes, as ADIPOQ, EGR1, COL1A1, JUN, FOS and KLF4, were more regulated in CMJ than in cortex after WI. However, cold ischemia time seems to reduce this differential expression. It is interesting to note that the expression of two “immediate early” genes, EGR1 and c-FOS, have been described to be involved in response to renal ischemia after 30 min of WI59.
-
5.
“Nucleus genes and transcriptional regulation” such as HIRA, HIST1H2AB, HIST1H4A, AURKB, pSNORD3, NR1H4, LEO1 and DICER1 (Fig. 5A). Histone cell cycle regulator (HIRA) is required for the periodic repression/regulation of histone gene transcription during cell cycle60. Histone H2A (HIST1H2AB), Histone H4 (HIST1H4A) and Aurora-B kinase (AURKB) are important for DNA regulation and replication. AURKB is a key regulator of mitosis61 interacting with histones. Small nucleolar RNA C/D box 3 cluster (pSNORD3 cluster snoRNA) is associated with RNA methylation. Nuclear Receptor Subfamily 1H4 (NR1H4) is a ligand-activated transcription factor. RNA polymerase-associated protein LEO1 (upregulated during cold-stoarge) is a component of the PAF1 complex (PAF1C) which has multiple functions during transcription and is implicated in regulation of development and maintenance of embryonic stem cell pluripotency and required for transcription of Hox and Wnt target genes62. Finally, Dicer-1 ribonuclease type-III (DICER1) is involved in mi-RNA production and its inhibition triggers resistance of tubular cells in a mouse model of kidney IRI63. This gene appears as upregulated in our model during cold storage, showing the importance of RNA interference contribution during ischemic stress.
-
6.
“Transporters” such as ABCC2, ATP2B1, KCNIP4, KCNJ10, MCOLN3 and AQP3 (Fig. 5B). Organ preservation induces membrane tansporters activation to “physiologic balance” reestablishement. ATP-binding cassette (ABC) transporters, like ABCC2, utilize the energy of ATP binding and hydrolysis to transport various substrates across cellular membranes. ABCC2, also called multidrug resistance protein MRP2, acts as an ATP-dependent conjugate export pump in apical membranes of polarized cells64. Plasma membrane calcium-transporting ATPase-1 (ATP2B1) is a magnesium-dependent enzyme catalyzing the hydrolysis of ATP coupled with calcium transport out of the cell. Potassium voltage-gated (Kv) channel interacting protein-4 (KCNIP4) is a subunit component of native Kv4 channel complexes. ATP-sensitive inward rectifier potassium channel-10 (KCNJ10) has a greater tendency to allow potassium to flow into the cell. Regulation of these RNA during cold storage for ABBC2, ATP2B1, KCNIP4 and KCNJ10 could be due to ion balance of the preservation solution and ATP availability. Mucolipin-3 (MCOLN3) is an inwardly-rectifying cation channel65 which mediates Ca2+ release from endosomes to cytoplasm (contributing to endosomal acidification) and is involved in the regulation of membrane trafficking and fusion in the endosomal pathway66. Here, ATP2B1 and MCOLN3 are upregulated during cold storage due to Ca2+ trafficking deregulation, one major consequence of ischemia67. Aquaporin-3 (AQP3) provides kidney medullary collecting duct with high permeability to water, thereby enabling water toward an osmotic gradient. In our study, during WI and in response to water deprivation, AQP3 expression increases in kidney cortex and medulla68.
-
7.
“Metabolism regulation” such as MTHFD1, LRCH1, ACBP, ACADVL, HADHA, CPT2, FUT2, CKM, ARG2 and PDK4 (Fig. 6). Ischemia alters cell metabolism, however the full range of alterations remains to be defined. Methylenetetrahydrofolate dehydrogenase-1 (MTHFD1) gene encodes the C-1-tetrahydrofolate synthase cytoplasmic protein which is involved in the pathway of tetrahydrofolate interconversion. Leucine-rich repeat and calponin homology domain-containing protein-1 (LRCH1) prevents CDC42 activation and negatively regulates CD4+ T-cell migration69. Acyl-CoA-binding domain-containing protein (ACBP) likely participates in intermembrane lipid transport from the ER to the plasma membrane, where it could maintain a membrane-associated acyl pool70. Acyl-CoA dehydrogenase very long chain (ACADVL) is targeted to the inner mitochondrial membrane, where it catalyzes the first step of the mitochondrial fatty acid beta-oxidation pathway71. Hydroxyacyl-CoA dehydrogenase trifunctional multienzyme complex subunit alpha (HADHA) catalyzes the last three steps of mitochondrial beta-oxidation of long chain fatty acids72. Carnitine O-palmitoyltransferase-2 (CPT2) oxidizes long-chain fatty acids in the mitochondria73. CPT promotes the binding of Acyl-CoA to Carnitine. ACBP, ACADVL, HADHA and CPT2 are mostly dowregulated during cold ischemia likely due to regulation of fatty-acids beta-oxydation. Fucosyltransferase-2 (FUT2) mediates glycosylation of cell surface glycoproteins and glycolipids74. Glycosylation, acts in rough endoplasmic reticulum, induces tissue aging but also may have protective effects75. Our results show that FUT2 is upregulated during cold storage-induced ischemia. Creatine kinase M-type (CKM) play a central role in energy transduction. CKM regenerates ATP from ADP, using phosphocreatine76. Creatine kinase is a marker of damage of CK-rich tissue such as in acute kidney injury77. We observed that CKM is expressed only during cold ischemia. Arginase-2 (ARG2) has multiple fonctions as it play a role in the regulation of polyamine metabolism and also in down-regulation of nitric-oxide synthesis, it is involved in the negative regulation of the survival capacity of activated CD4+ and CD8+ T cells78 and it inhibit endothelial autophagy independently of its enzymatic activity implicating mTORC2 signaling79. Blocking ARG2 expression attenuated lesions in a mice model of IRI80. Our results are comforting these observations and the dowregulation of ARG2 seems to be protective. Pyruvate dehydrogenase lipoamide kinase isozyme-4 (PDK4) is located in the mitochondria matrix and inhibits the pyruvate dehydrogenase complex by phosphorylating one of its subunits, reducing the conversion of pyruvate, which is produced from the oxydation of glucose and amino acids to acetyl-CoA and contributing to glucose metabolism regulation. PDK4 helps to decrease metabolism and conserve glucose by limiting its conversion to acetyl-CoA, which enters the citric acid cycle and is converted to ATP81.
-
8.
“MAPK and GTPase activity” such as RABGAP1, PARP4, MAP3K8, MLK4, MAP3K5, RHOB, PLK3, MX2, RHOU, JUN and EXPH5 (Fig. 7). Ischemia activates intracellular enzymes as G Proteins and MAPK to regulate several pathways. CDC42 effector protein 1 (CDC42EP1) is a member of the Rho GTPase family regulating multiple cellular activities, including organization of the actin cytoskeleton82. Rab GTPase-activating protein-1 (RABGAP1), upregulated herein during cold storage, acts as a GTPase-activating protein in the RAB6A-mediated pathway involved in the mitotic metaphase-anaphase transition83. Poly [ADP-ribose] polymerase-4 (PARP4) is able to catalyze a poly(ADP-ribosyl)ation reaction. PARP4 interacts with Major vault protein, which interacts with the inactive PERK (protein involved in endoplasmic reticulum stress). Mitogen-activated protein kinase-kinase-kinase-8 (MAP3K8, also called TPL-2) can activate both the ERK1/2 and p38 MAP kinases84. MAP3K8 regulates renal cell apoptosis in ischemia/reperfusion injury85. Herein, MAP3K8 is dowregulated from WI to end of cold storage. Mitogen-activated protein kinase kinase kinase 21 (MAP3K21 also called MLK4), herein downregulated during cold storage, is a negative regulator of TLR4 signaling86. This is in favour of the TLR4 sigaling activation, one of the major pathway involved during IRI87. Thus, activation of MLK4 could be an interesting target for new therapy. Mitogen-activated protein kinase kinase kinase 5 (MAP3K5), also called apoptosis signal-regulating kinase-1 (ASK1), is pivotal component in cell apoptosis and can be activated by a variety of death stimuli including TNF-alpha and oxidative stress88. Herein, the downregulation of MAP3K5 during cold ischemia is protective. Ras homolog gene family member-B (RHOB), upregulated during WI, enhances cytokine-induced transcription of inductible nitric-oxide synthase-2 (iNOS)89 inducing a oxidative environment. PLK3, upregulated from WI to cold storage end, is involved in cell cycle regulation, response to stress, Golgi disassembly and DNA damage response90. PLK3 is rapidly activated upon stress stimulation, such as ROS, hyperosmotic stress and hypoxia. PLK3 is important for the downregulation of apoptosis and regulation of microtubule dynamics and centrosomal function91. Interferon-induced GTP-binding protein MX2, also dowregulated during cold storage, regulates nucleocytoplasmic transport and cell-cycle progression92. Rho-related GTP-binding protein RHOU is encoded by a non-canonical Wnt induced gene93. RHOU/Wrch delineates with RhoV/Chp a Rho subclass related to RAC and CDC42, which emerged in early multicellular organisms during evolution94. Similarly to CDC42, RHOU and Exophilin-5 (EXPH5) are also dowregulated during cold storage. EXPH5 may act as Rab effector protein and play a role in intracellular vesicle trafficking. Reduced expression of this gene results in keratin filament defects, in association with collagen structure.
We summarized the kinetic of altered pathways during the experiment timecourse in Fig. 8, taking into account the Gene-Ontology Analysis and our data interpretation in the context of renal ischemia. Several gene families are up- or down-regulated similarly in both cortex and CMJ, but we noted some differences of expression among renal regions at specific timepoints, as reported in Fig. 8.
Alltogether, our study highlight critical genetic alterations induced by ischemia at the cellular/tissular levels, dissecting ischemia mechanism and kinetics using an experimental model extremely close to human conditions. Several of these pathways have “opposite” roles (e.g.: survival/development versus apoptosis/death) which may be linked to the high stress level withstood by the kidneys in our experiments resulting to highly complex responses at the cellular level aimed at counterbalancing stress-induced lesions. Beyond improving our understanding of IRI, our study point out several dysregulated genes which could be used as biomarkers of ischemia injury, allowing a thinner evaluation of kidney quality, one of the major challenges in renal transplantation. In order to validate their quality as biomarkers, further studies are required to evaluate the protein expression level in kidneys subjected to different levels of ischemic injury and correlate these data to kidney transplantation outcome. In conclusion, our data strengthen the fact that ischemia is a key step during the transplantation process with important transcriptional modifications inducing a full reprogramming of the transcriptome of major pathways such those related to oxidative stress responses, cell reprogramming, cell-cycle, inflammation and cell metabolism. These pathways provide interesting research prospects for the development of strategies which could be used during kidney conservation, aimed at improving whole transplantation outcome.
Methods
Animal experimentation
The animal experimental protocol was approved by French Government and institutional Committee on the Ethics of Animal Experiments of the Poitou-Charentes (France) (comity number C2EA-84, protocol number: CE2012-4). Experimentations were performed in accordance with EU Directive 2010/63/EU at the IBiSA MOPICT platform, INRA Magneraud, France. Full methods for animal experimentation are provided in Suplementary Material and Methods.
Microarrays slides
Porcine Gene Expression Microarray G2519F (Agilent) contains 60-mer oligonucleotide probes to 43,803 porcine probes for the pig Sus Scrofa. The data discussed in this article have been deposited in NCBIs Gene Expression Omnibus Database (GEO), accessible through accession number GSE10971995.
RNA Isolation
RNA was extracted using a commercial kit including a DNAse step to remove genomic DNA (Qiagen RNeasy plus mini). The RNA yield and integrity were controlled using a Nanodrop ND-1000 and a Bioanalyzer 2100 Expert from Agilent, then labelled with the Low Input Quick Amp Labeling kit, designed to reliably amplify and label target RNA for the robust generation of complementary RNA.
Hybridizations
Hybridization was performed following the One-Color Microarray-Based Gene Expression Analysis Protocol. Assembled slide chamber was placed in rotisserie in a hybridization oven. Hybridization took place during 17 hours at 65 °C and at 10 rpm.
Slide scanning and image analysis/treatment
The slides were scanned on a Tecan MS200 scanner and analyzed by Feature Extraction V.11.5.1.1. Pre-analysis data quality assessment was performed by visual inspection of individual false color hybridization images and standard diagnostic plots of probe level intensity distributions using Bioconductor (http://www.bioconductor.org/) and R software (version 2.15.2). All data were analyzed using the Bioconductor software project and the statistical language R. After transformed log2 data, the data were normalized by condition using quantile method96 with limma package97 and finally summarized. Significant genes were identified using the limma package. A False Discovery Rate corrected p-value < 0.05 and a log2 fold change >0.5 were used as significance criterion.
Heatmap
The heatmap was generated with R software using Euclidean distance and Ward linkage from the list of differentially expressed genes between Ischemia and Control.
Functional enrichment analysis
The detection of significantly overrepresented Gene Ontology categories was performed using the GOStats package in Bioconductor98. The list of differentially expressed genes was divided in two parts: up-regulated and down-regulated genes. For each sublist, we performed a hypergeometrical statistical test; corresponding to the ratio of differentially expressed genes found in the category over the total number of genes in the category compared to the ratio of the total number of differentially expressed genes over total number of genes on the chip.
Real-Time Quantitative-PCR confirmation
Full methods for Real-Time Quantitative-PCR are provided in Supplementary Material and Methods.
References
Brook, N. R. & Nicholson, M. L. Kidney transplantation from non heart-beating donors. Surg. J. R. Coll. Surg. Edinb. Irel. 1, 311–322 (2003).
Bon, D. et al. New strategies to optimize kidney recovery and preservation in transplantation. Nat. Rev. Nephrol. 8, 339–347 (2012).
Giraud, S. et al. Contribution of large pig for renal ischemia-reperfusion and transplantation studies: the preclinical model. J. Biomed. Biotechnol. 2011, 532127 (2011).
Chen, G. et al. Increased susceptibility of aging kidney to ischemic injury: identification of candidate genes changed during aging, but corrected by caloric restriction. Am. J. Physiol. Renal Physiol. 293, F1272–1281 (2007).
Liu, M. et al. Effect of T cells on vascular permeability in early ischemic acute kidney injury in mice. Microvasc. Res. 77, 340–347 (2009).
Naesens, M. et al. Expression of complement components differs between kidney allografts from living and deceased donors. J. Am. Soc. Nephrol. JASN 20, 1839–1851 (2009).
Damman, J. et al. Hypoxia and Complement-and-Coagulation Pathways in the Deceased Organ Donor as the Major Target for Intervention to Improve Renal Allograft Outcome. Transplantation 99, 1293–1300 (2015).
Tamura, Y., Imaoka, S., Gemba, M. & Funae, Y. Effects of ischemia-reperfusion on individual cytochrome P450 isoforms in the rat kidney. Life Sci. 60, 143–149 (1997).
Gulati, S., Ainol, L., Orak, J., Singh, A. K. & Singh, I. Alterations of peroxisomal function in ischemia-reperfusion injury of rat kidney. Biochim. Biophys. Acta 1182, 291–298 (1993).
Thomas, R. A. B. et al. Hemin Preconditioning Upregulates Heme Oxygenase-1 in Deceased Donor Renal Transplant Recipients: A Randomized, Controlled, Phase IIB Trial. Transplantation 100, 176–183 (2016).
Potteti, H. R. et al. Nrf2-AKT interactions regulate heme oxygenase 1 expression in kidney epithelia during hypoxia and hypoxia-reoxygenation. Am. J. Physiol. Renal Physiol. 311, F1025–F1034 (2016).
Mailloux, R. J., Jin, X. & Willmore, W. G. Redox regulation of mitochondrial function with emphasis on cysteine oxidation reactions. Redox Biol. 2, 123–139 (2014).
Bak, D. W. & Weerapana, E. Cysteine-mediated redox signalling in the mitochondria. Mol. Biosyst. 11, 678–697 (2015).
Imamura, R. et al. Erythropoietin protects the kidneys against ischemia reperfusion injury by activating hypoxia inducible factor-1alpha. Transplantation 83, 1371–1379 (2007).
Cano, M. et al. Oxidative stress induces mitochondrial dysfunction and a protective unfolded protein response in RPE cells. Free Radic. Biol. Med. 69, 1–14 (2014).
Ponnappan, S., Ovaa, H. & Ponnappan, U. Lower expression of catalytic and structural subunits of the proteasome contributes to decreased proteolysis in peripheral blood T lymphocytes during aging. Int. J. Biochem. Cell Biol. 39, 799–809 (2007).
Tavaria, M., Gabriele, T., Kola, I. & Anderson, R. L. A hitchhiker’s guide to the human Hsp70 family. Cell Stress Chaperones 1, 23–28 (1996).
Saito, Y., Doi, K., Yamagishi, N., Ishihara, K. & Hatayama, T. Screening of Hsp105alpha-binding proteins using yeast and bacterial two-hybrid systems. Biochem. Biophys. Res. Commun. 314, 396–402 (2004).
Rauch, J. N. & Gestwicki, J. E. Binding of human nucleotide exchange factors to heat shock protein 70 (Hsp70) generates functionally distinct complexes in vitro. J. Biol. Chem. 289, 1402–1414 (2014).
Simkhovich, B. Z., Marjoram, P., Poizat, C., Kedes, L. & Kloner, R. A. Brief episode of ischemia activates protective genetic program in rat heart: a gene chip study. Cardiovasc. Res. 59, 450–459 (2003).
Carr, M. W., Roth, S. J., Luther, E., Rose, S. S. & Springer, T. A. Monocyte chemoattractant protein 1 acts as a T-lymphocyte chemoattractant. Proc. Natl. Acad. Sci. USA 91, 3652–3656 (1994).
Cuvelier, S. L. & Patel, K. D. Shear-dependent eosinophil transmigration on interleukin 4-stimulated endothelial cells: a role for endothelium-associated eotaxin-3. J. Exp. Med. 194, 1699–1709 (2001).
Ogilvie, P., Paoletti, S., Clark-Lewis, I. & Uguccioni, M. Eotaxin-3 is a natural antagonist for CCR2 and exerts a repulsive effect on human monocytes. Blood 102, 789–794 (2003).
Sasaki, T. et al. Genome-Wide Gene Expression Profiling Revealed a Critical Role for GATA3 in the Maintenance of the Th2 Cell Identity. PloS One 8, e66468 (2013).
Chen, C.-B. et al. Up-Regulation of HMGB1 Exacerbates Renal Ischemia-Reperfusion Injury by Stimulating Inflammatory and Immune Responses through the TLR4 Signaling Pathway in Mice. Cell. Physiol. Biochem. Int. J. Exp. Cell. Physiol. Biochem. Pharmacol. 41, 2447–2460 (2017).
Zhang, J. et al. HMGB1-TLR4 signaling participates in renal ischemia reperfusion injury and could be attenuated by dexamethasone-mediated inhibition of the ERK/NF-κB pathway. Am. J. Transl. Res. 8, 4054–4067 (2016).
Huang, J. et al. ZNF216 Is an A20-like and IkappaB kinase gamma-interacting inhibitor of NFkappaB activation. J. Biol. Chem. 279, 16847–16853 (2004).
Choi, K.-C. et al. Smad6 negatively regulates interleukin 1-receptor-Toll-like receptor signaling through direct interaction with the adaptor Pellino-1. Nat. Immunol. 7, 1057–1065 (2006).
Nurieva, R. I. et al. Bcl6 mediates the development of T follicular helper cells. Science 325, 1001–1005 (2009).
Urbich, C. et al. Upregulation of TRAF-3 by shear stress blocks CD40-mediated endothelial activation. J. Clin. Invest. 108, 1451–1458 (2001).
Garcia, J. et al. IEX-1: a new ERK substrate involved in both ERK survival activity and ERK activation. EMBO J. 21, 5151–5163 (2002).
Arlt, A. & Schäfer, H. Role of the immediate early response 3 (IER3) gene in cellular stress response, inflammation and tumorigenesis. Eur. J. Cell Biol. 90, 545–552 (2011).
Woo, I. S. et al. TMEM14A inhibits N-(4-hydroxyphenyl)retinamide-induced apoptosis through the stabilization of mitochondrial membrane potential. Cancer Lett. 309, 190–198 (2011).
Müller, C., Calkhoven, C. F., Sha, X. & Leutz, A. The CCAAT enhancer-binding protein alpha (C/EBPalpha) requires a SWI/SNF complex for proliferation arrest. J. Biol. Chem. 279, 7353–7358 (2004).
Pedersen, T. A. et al. Distinct C/EBPalpha motifs regulate lipogenic and gluconeogenic gene expression in vivo. EMBO J. 26, 1081–1093 (2007).
Qiao, L. & Shao, J. SIRT1 regulates adiponectin gene expression through Foxo1-C/enhancer-binding protein alpha transcriptional complex. J. Biol. Chem. 281, 39915–39924 (2006).
Díez, J. J. & Iglesias, P. The role of the novel adipocyte-derived hormone adiponectin in human disease. Eur. J. Endocrinol. 148, 293–300 (2003).
Hu, C.-T. et al. Snail associates with EGR-1 and SP-1 to upregulate transcriptional activation of p15INK4b. FEBS J. 277, 1202–1218 (2010).
Yan, S. F. et al. Egr-1, a master switch coordinating upregulation of divergent gene families underlying ischemic stress. Nat. Med. 6, 1355–1361 (2000).
Liao, H., Hyman, M. C., Lawrence, D. A. & Pinsky, D. J. Molecular regulation of the PAI-1 gene by hypoxia: contributions of Egr-1, HIF-1alpha, and C/EBPalpha. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 21, 935–949 (2007).
Simon, M. P. et al. Deregulation of the platelet-derived growth factor B-chain gene via fusion with collagen gene COL1A1 in dermatofibrosarcoma protuberans and giant-cell fibroblastoma. Nat. Genet. 15, 95–98 (1997).
Wozny, W. et al. Differential radioactive quantification of protein abundance ratios between benign and malignant prostate tissues: cancer association of annexin A3. Proteomics 7, 313–322 (2007).
Troppmann, C. et al. Delayed graft function, acute rejection, and outcome after cadaver renal. transplantation. The multivariate analysis. Transplantation 59, 962–8 (1995).
Park, J. E. et al. Annexin A3 is a potential angiogenic mediator. Biochem. Biophys. Res. Commun. 337, 1283–1287 (2005).
Ito, Y. et al. Inhibition of angiogenesis and vascular leakiness by angiopoietin-related protein 4. Cancer Res. 63, 6651–6657 (2003).
Guaiquil, V. et al. ADAM9 is involved in pathological retinal neovascularization. Mol. Cell. Biol. 29, 2694–2703 (2009).
Nath, D. et al. Meltrin gamma(ADAM-9) mediates cellular adhesion through alpha(6)beta(1)integrin, leading to a marked induction of fibroblast cell motility. J. Cell Sci. 113(Pt 12), 2319–2328 (2000).
Fujikawa, K. et al. Vav3 is regulated during the cell cycle and effects cell division. Proc. Natl. Acad. Sci. USA 99, 4313–4318 (2002).
Lochhead, P. A. et al. dDYRK2: a novel dual-specificity tyrosine-phosphorylation-regulated kinase in Drosophila. Biochem. J. 374, 381–391 (2003).
Hu, E. et al. Targeted disruption of the c-fos gene demonstrates c-fos-dependent and -independent pathways for gene expression stimulated by growth factors or oncogenes. EMBO J. 13, 3094–3103 (1994).
Chiu, R. et al. The c-Fos protein interacts with c-Jun/AP-1 to stimulate transcription of AP-1 responsive genes. Cell 54, 541–552 (1988).
Tulchinsky, E. Fos family members: regulation, structure and role in oncogenic transformation. Histol. Histopathol. 15, 921–928 (2000).
Eskildsen, S., Justesen, J., Schierup, M. H. & Hartmann, R. Characterization of the 2′-5′-oligoadenylate synthetase ubiquitin-like family. Nucleic Acids Res. 31, 3166–3173 (2003).
Johnson, D. I. & Pringle, J. R. Molecular characterization of CDC42, a Saccharomyces cerevisiae gene involved in the development of cell polarity. J. Cell Biol. 111, 143–152 (1990).
Yang, Q. et al. The BLOS1-interacting protein KXD1 is involved in the biogenesis of lysosome-related organelles. Traffic Cph. Den. 13, 1160–1169 (2012).
Shields, J. M., Christy, R. J. & Yang, V. W. Identification and characterization of a gene encoding a gut-enriched Krüppel-like factor expressed during growth arrest. J. Biol. Chem. 271, 20009–20017 (1996).
Yoon, H. S., Chen, X. & Yang, V. W. Kruppel-like factor 4 mediates p53-dependent G1/S cell cycle arrest in response to DNA damage. J. Biol. Chem. 278, 2101–2105 (2003).
Yoon, H. S. & Yang, V. W. Requirement of Krüppel-like factor 4 in preventing entry into mitosis following DNA damage. J. Biol. Chem. 279, 5035–5041 (2004).
Ouellette, A. J., Malt, R. A., Sukhatme, V. P. & Bonventre, J. V. Expression of two ‘immediate early’ genes, Egr-1 and c-fos, in response to renal ischemia and during compensatory renal hypertrophy in mice. J. Clin. Invest. 85, 766–771 (1990).
Zhang, R. et al. Formation of MacroH2A-containing senescence-associated heterochromatin foci and senescence driven by ASF1a and HIRA. Dev. Cell 8, 19–30 (2005).
Goto, H., Yasui, Y., Nigg, E. A. & Inagaki, M. Aurora-B phosphorylates Histone H3 at serine28 with regard to the mitotic chromosome condensation. Genes Cells Devoted Mol. Cell. Mech. 7, 11–17 (2002).
Rozenblatt-Rosen, O. et al. The parafibromin tumor suppressor protein is part of a human Paf1 complex. Mol. Cell. Biol. 25, 612–620 (2005).
Wei, Q. et al. Targeted deletion of Dicer from proximal tubules protects against renal ischemia-reperfusion injury. J. Am. Soc. Nephrol. JASN 21, 756–761 (2010).
Hagmann, W. et al. Purification of the human apical conjugate export pump MRP2 reconstitution and functional characterization as substrate-stimulated ATPase. Eur. J. Biochem. 265, 281–289 (1999).
Clapham, D. E., Julius, D., Montell, C. & Schultz, G. International Union of Pharmacology. XLIX. Nomenclature and structure-function relationships of transient receptor potential channels. Pharmacol. Rev. 57, 427–450 (2005).
Lelouvier, B. & Puertollano, R. Mucolipin-3 regulates luminal calcium, acidification, and membrane fusion in the endosomal pathway. J. Biol. Chem. 286, 9826–9832 (2011).
Schrier, R. W. & Hensen, J. Cellular mechanism of ischemic acute renal failure: role of Ca2+ and calcium entry blockers. Klin. Wochenschr. 66, 800–807 (1988).
Amlal, H. & Wilke, C. Resistance of mTAL Na+-dependent transporters and collecting duct aquaporins to dehydration in 7-month-old rats. Kidney Int. 64, 544–554 (2003).
Xu, X. et al. LRCH1 interferes with DOCK8-Cdc42-induced T cell migration and ameliorates experimental autoimmune encephalomyelitis. J. Exp. Med. 214, 209–226 (2017).
Chye, M. L., Li, H. Y. & Yung, M. H. Single amino acid substitutions at the acyl-CoA-binding domain interrupt 14[C]palmitoyl-CoA binding of ACBP2, an Arabidopsis acyl-CoA-binding protein with ankyrin repeats. Plant Mol. Biol. 44, 711–721 (2000).
Pougovkina, O. et al. Mitochondrial protein acetylation is driven by acetyl-CoA from fatty acid oxidation. Hum. Mol. Genet. 23, 3513–3522 (2014).
Carpenter, K., Pollitt, R. J. & Middleton, B. Human liver long-chain 3-hydroxyacyl-coenzyme A dehydrogenase is a multifunctional membrane-bound beta-oxidation enzyme of mitochondria. Biochem. Biophys. Res. Commun. 183, 443–448 (1992).
Djouadi, F., Aubey, F., Schlemmer, D. & Bastin, J. Peroxisome proliferator activated receptor delta (PPARdelta) agonist but not PPARalpha corrects carnitine palmitoyl transferase 2 deficiency in human muscle cells. J. Clin. Endocrinol. Metab. 90, 1791–1797 (2005).
Kelly, R. J., Rouquier, S., Giorgi, D., Lennon, G. G. & Lowe, J. B. Sequence and expression of a candidate for the human Secretor blood group alpha(1,2)fucosyltransferase gene (FUT2). Homozygosity for an enzyme-inactivating nonsense mutation commonly correlates with the non-secretor phenotype. J. Biol. Chem. 270, 4640–4649 (1995).
Simm, A. et al. Protein glycation - Between tissue aging and protection. Exp. Gerontol. 68, 71–75 (2015).
Wallimann, T., Wyss, M., Brdiczka, D., Nicolay, K. & Eppenberger, H. M. Intracellular compartmentation, structure and function of creatine kinase isoenzymes in tissues with high and fluctuating energy demands: the ‘phosphocreatine circuit’ for cellular energy homeostasis. Biochem. J. 281(Pt 1), 21–40 (1992).
Moghadam-Kia, S., Oddis, C. V. & Aggarwal, R. Approach to asymptomatic creatine kinase elevation. Cleve. Clin. J. Med. 83, 37–42 (2016).
Geiger, R. et al. L-Arginine Modulates T Cell Metabolism and Enhances Survival and Anti-tumor Activity. Cell 167, 829–842.e13 (2016).
Xiong, Y. et al. ARG2 impairs endothelial autophagy through regulation of MTOR and PRKAA/AMPK signaling in advanced atherosclerosis. Autophagy 10, 2223–2238 (2014).
Raup-Konsavage, W. M. et al. Arginase-2 mediates renal ischemia-reperfusion injury. Am. J. Physiol. Renal Physiol. 313, F522–F534 (2017).
Andrews, M. T., Squire, T. L., Bowen, C. M. & Rollins, M. B. Low-temperature carbon utilization is regulated by novel gene activity in the heart of a hibernating mammal. Proc. Natl. Acad. Sci. USA 95, 8392–8397 (1998).
Burbelo, P. D., Snow, D. M., Bahou, W. & Spiegel, S. MSE55, a Cdc42 effector protein, induces long cellular extensions in fibroblasts. Proc. Natl. Acad. Sci. USA 96, 9083–9088 (1999).
Cuif, M. H. et al. Characterization of GAPCenA, a GTPase activating protein for Rab6, part of which associates with the centrosome. EMBO J. 18, 1772–1782 (1999).
Arthur, J. S. C. & Ley, S. C. Mitogen-activated protein kinases in innate immunity. Nat. Rev. Immunol. 13, 679–692 (2013).
Yaomura, T. et al. Serine/threonine kinase, Cot/Tpl2, regulates renal cell apoptosis in ischaemia/reperfusion injury. Nephrol. Carlton Vic 13, 397–404 (2008).
Seit-Nebi, A., Cheng, W., Xu, H. & Han, J. MLK4 has negative effect on TLR4 signaling. Cell. Mol. Immunol. 9, 27–33 (2012).
Zhao, H., Perez, J. S., Lu, K., George, A. J. T. & Ma, D. Role of Toll-like receptor-4 in renal graft ischemia-reperfusion injury. Am. J. Physiol. Renal Physiol. 306, F801–811 (2014).
Li, X. et al. Tumor necrosis factor alpha-induced desumoylation and cytoplasmic translocation of homeodomain-interacting protein kinase 1 are critical for apoptosis signal-regulating kinase 1-JNK/p38 activation. J. Biol. Chem. 280, 15061–15070 (2005).
Delarue, F. L., Taylor, B. S. & Sebti, S. M. Ras and RhoA suppress whereas RhoB enhances cytokine-induced transcription of nitric oxide synthase-2 in human normal liver AKN-1 cells and lung cancer A-549 cells. Oncogene 20, 6531–6537 (2001).
Smith, P., Syed, N. & Crook, T. Epigenetic inactivation implies a tumor suppressor function in hematologic malignancies for Polo-like kinase 2 but not Polo-like kinase 3. Cell Cycle Georget. Tex 5, 1262–1264 (2006).
Wang, Q. et al. Cell cycle arrest and apoptosis induced by human Polo-like kinase 3 is mediated through perturbation of microtubule integrity. Mol. Cell. Biol. 22, 3450–3459 (2002).
King, M. C., Raposo, G. & Lemmon, M. A. Inhibition of nuclear import and cell-cycle progression by mutated forms of the dynamin-like GTPase MxB. Proc. Natl. Acad. Sci. USA 101, 8957–8962 (2004).
Tao, W., Pennica, D., Xu, L., Kalejta, R. F. & Levine, A. J. Wrch-1, a novel member of the Rho gene family that is regulated by Wnt-1. Genes Dev. 15, 1796–1807 (2001).
Boureux, A., Vignal, E., Faure, S. & Fort, P. Evolution of the Rho family of ras-like GTPases in eukaryotes. Mol. Biol. Evol. 24, 203–216 (2007).
Gene Expression Omnibus Database [http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE109719].
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinforma. Oxf. Engl. 19, 185–193 (2003).
Smyth, G. Limma: linear models for microarray data. Bioinforma. Comput. Biol. Solut. Using R Bioconductor Ed. Gentlem. R Carey V Huber W Irizarry R Dudoit Springer 397–420 (2005).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinforma. Oxf. Engl. 23, 257–258 (2007).
Acknowledgements
The authors thank the sources of funding: this work was supported by laboratory institutional grants from Institut national de la santé et de la recherche médicale (INSERM), Conseil Régional Poitou-Charentes, Université de Poitiers and Centre hospitalier Universitaire (CHU) de Poitiers. We thank Institut national de recherche agronomique (INRA). We thank Lidwine Trouilh from Plateforme GeT-Biopuce (LISBP - INSA de Toulouse, France) and William Hebrard (Plateforme MOPICT, INRA Surgères, France) for their excellent technical support.
Author information
Authors and Affiliations
Contributions
S.G., C.S., G.A., P.C., D.L., S.L., V.A., S.T. performed ex vivo and in vitro experiments, data acquisition and analysis. S.G., C.S., P.H., R.T. and T.H. performed the design of the study, interpretation of data and drafted the manuscript. T.H. coordinated the study. All the authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Giraud, S., Steichen, C., Allain, G. et al. Dynamic transcriptomic analysis of Ischemic Injury in a Porcine Pre-Clinical Model mimicking Donors Deceased after Circulatory Death. Sci Rep 8, 5986 (2018). https://doi.org/10.1038/s41598-018-24282-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-24282-6
- Springer Nature Limited
This article is cited by
-
Exploring the therapeutic mechanisms of Coptidis Rhizoma in gastric precancerous lesions: a network pharmacology approach
Discover Oncology (2024)
-
Predictive value of serum proteomic biomarkers for noise-induced hearing loss
Environmental Science and Pollution Research (2023)
-
Multi-omics Approach in Kidney Transplant: Lessons Learned from COVID-19 Pandemic
Current Transplantation Reports (2023)
-
Hypothermic machine perfusion reduces donation after circulatory death liver ischemia–reperfusion injury through the SERPINA3-mediated PI3Kδ/Akt pathway
Human Cell (2023)