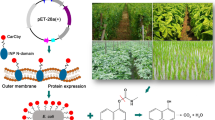
Abstract
Many sites are often co-contaminated with multiple pesticides. To date, there are no reports on simultaneous degradation of different classes of pesticides by a natural microorganism. In this work, we aim at constructing a live biocatalyst able to simultaneously hydrolyze carbaryl and chlorpyrifos. For this purpose, carbaryl hydrolase (CH) was displayed on the cell surface of a chlorpyrifos-degrading bacterium Stenotrophomonas sp. strain YC-1 using N- and C-terminal domain of ice nucleation protein (INPNC) from Pseudomonas syringae INA5 as an anchoring motif. The localization of INPNC-CH fusion protein in the outer membrane fraction was demonstrated by cell fractionation followed by Western blot analysis. Surface display of INPNC-CH was further confirmed by proteinase accessibility experiment and immunofluorescence microscope. CH was present in an active form on cell surface without causing any growth inhibition, suggesting that the INP-based display system is a useful tool for surface expression of macromolecular heterologous proteins on the bacterial cell surface. Because surface-displayed CH has free access to pesticides, this bacterium can be used as a whole-cell biocatalyst for efficient hydrolysis of pesticides.
Similar content being viewed by others
Introduction
Chlorpyrifos is a moderately toxic organophosphorus pesticide that has played an important role in controlling major agricultural pests and raising agricultural productivity. Chlorpyrifos leads to the loss of nerve function of vertebrates by irreversibly inhibiting acetylcholine esterase (AChE) in the central nervous system1,2. Carbaryl is a broad-spectrum carbamate insecticide and is a potent AChE inhibitor3,4. Carbaryl and chlorpyrifos are often used simultaneously to control agricultural pests; thus, many sites are co-contaminated with carbaryl and chlorpyrifos. Especially, the co-existence of different classes of pesticides can potentiate the toxicity of individual pesticides and the joint toxicity of pesticides poses a great threat to human health5.
Microorganisms have evolved degradation pathways for the removal of synthesized pesticides. So far, several carbaryl-degrading bacteria, such as Rhizobium sp. strain AC1006, Arthrobacter sp. strain RC1004,7, Blastobacter sp. strain M5013, and Pseudomonas sp. strains C4, C5, and C68,9,10, have been isolated. Moreover, a carbaryl hydrolase gene (cehA) was cloned from Rhizobium sp. strain AC1006. To date, several chlorpyrifos-degrading bacteria, such as Enterobacter strain B-1411, Alcaligenes faecalis strain DSP312, Stenotrophomonas sp. strain YC-113, Sphingomonas sp. strain Dsp-214, Paracoccus sp. strain TRP15, Bacillus pumilus strain C2A116, and Cupriavidus sp. strain DT-117, have been isolated by selective enrichment with chlorpyrifos. Unfortunately, microorganisms with the capability to simultaneously degrade chlorpyrifos and carbaryl have not yet been isolated from the environment. Therefore, the construction of recombinant microorganisms is useful for the remediation of multiple pesticides-contaminated sites.
Carbaryl-degrading bacteria have been shown to produce intracellular carbaryl hydrolase (CH), and therefore intracellular CH cannot directly bind to extracellular substrates3,6,7. This bottleneck, however, could be eliminated if CH is displayed onto the surface of cells. Previous studies have shown that surface-displayed organophosphorus hydrolase (OPH) can directly interact with substrates without the membrane permeability barrier, thereby improving the overall catalytic efficiency18,19,20.
Various surface-anchoring motifs, such as Lpp-OmpA chimera, ice nucleation protein (INP), and autotransporter, have been widely used to display proteins on the surface of bacterial cell21,22. INP, an outer membrane protein found in Pseudomonas syringae, Xanthomonas campestris, and Erwinia herbicola, accelerates the formation of ice crystal in supercooled water. INP is composed of three distinct domains, including an N-terminal domain, a C-terminal domain, and a highly repetitive central domain23. INP-mediated cell surface display can be achieved using either full-length INP or truncated INP through C-terminal fusion18,24,25. Various types of INPs (e.g., InaK, InaPb, InaQ, InaV and InaZ) have been successfully used to display heterologous proteins, such as levansucrase24, synthetic phytochelatins26, OPH18,19,20,27, green fluorescent protein (GFP)25, chitinase28, NADPH-cytochrome P450 oxidoreductase (CPR)29, human poliovirus receptor30, carbonic anhydrase31, glucose dehydrogenase32, glutamate dehydrogenase33, xylose dehydrogenase34,35, mycoplasma adhesion proteins36, human norovirus capsid proteins37, formate dehydrogenase38, and human arginase-139, on the bacterial cell surface.
We previously isolated a chlorpyrifos-degrading bacterium Stenotrophomonas sp. strain YC-1, and this strain could utilize chlorpyrifos as the sole source of carbon for growth. Moreover, a chromosome-based mpd gene coding for chlorpyrifos hydrolase (CPH) was cloned from strain YC-113. CH and CPH require no cofactors for their activity. In this work, CH encoded by the cehA gene (GenBank accession no. AB069723) from Rhizobium sp. strain AC100 was functionally displayed on the cell surface of Stenotrophomonas sp. strain YC-1 using the truncated InaV (GenBank accession no. AJ001086) from P. syringae INA5 as an anchoring motif, resulting in a recombinant strain capable of simultaneously hydrolyzing carbaryl and chlorpyrifos.
Results and Discussion
Surface localization of INPNC-CH fusion protein
In this study, the feasibility of targeting CH onto the cell surface of Stenotrophomonas sp. strain YC-1 was investigated using the N- and C-terminal domain of InaV (INPNC)40 as an anchoring motif. To create the surface expression vector pVIC3, the inpnc-cehA fusion gene was subcloned into a medium-copy-number vector, pVLT33. The broad-host-range vector, pVLT33, has an ability to replicate in a wide variety of Gram-negative bacteria because it is an RSF1010 derivative41. Expression of INPNC-CH was tightly regulated by an inducible tac promoter and the lacI q gene on the plasmid pVLT33.
To verify whether INPNC-CH fusion protein was produced in the recombinant strain YC-1 carrying pVIC3, Western blot analysis was performed using rabbit polyclonal anti-CH antibodies. As a result, a specific immunoreactive band appeared at the position of ~120 kDa was detected in whole-cell lysates of YC-1 cells carrying pVIC3 (Fig. 1A, lane 2), which matches well with the molecular weight of INPNC-CH fusion protein. However, the immunoreactive band (~120 kDa) was not detected in the control cells carrying the empty vector pVLT33. These results demonstrated the successful synthesis of INPNC-CH fusion protein in the recombinant strain YC-1 carrying pVIC3. To demonstrate the localization of INPNC-CH fusion protein in the outer membrane, the outer membrane and soluble fractions collected from cells carrying pVIC3 were probed with polyclonal anti-CH antibodies. As expected, the vast majority of INPNC-CH fusion proteins (accounting for ~90% of total amount) were associated with the outer membrane fraction as judged by the intensity of the protein bands (Fig. 1A, lane 4). To evaluate the quantity of CH displayed on the cell surface, CH activity was determined using whole cell, the outer membrane and total cell lysate. Over 90% of CH activity was detected in the outer membrane fraction of cells carrying pVIC3. In parallel, more than 90% of CH activity was present on the cell surface as judged by the ratio of whole-cell activity (0.241 U/OD600) to cell lysate activity (0.267 U/OD600).
Western blot analysis for subcellular localization of INPNC-CH fusion protein in Stenotrophomonas sp. strain YC-1/pVIC3 (A) and E. coli DH5α/pVIC3 (B). Lane 1, protein marker; lane 2, whole-cell lysates; lane 3, soluble fraction; lane 4, outer membrane fraction; lane 5, whole-cell lysates of YC-1/pVLT33 (or DH5α/pVLT33).
Since proteinases can hardly penetrate the outer membrane, only proteins displayed on the cell surface can be degraded by proteinases19,25. In this study, the surface localization of INPNC-CH was also confirmed by proteinase accessibility experiment. After treatment with proteinase K for 3 h, whole-cell CH activity of the recombinant strain YC-1 carrying pVIC3 was decreased by 90% relative to proteinase untreated cells, which contrasted with the slight reduction (7%) for YC-1/pVC3 cells expressing cytosolic CH. The outer membrane fraction from YC-1/pVIC3 cells treated with proteinase K for 3 or 2.5 h was probed with polyclonal anti-CH antibodies. As expected, only ~10% or ~20% of INPNC-CH fusion proteins (relative to proteinase untreated cells) were detected in the outer membrane fraction as judged by the intensity of the protein bands because the vast majority of surface-displayed CH molecules were degraded by proteinase K (Fig. S1).
Immunolabeling with specific antibodies is a useful tool to detect proteins displayed on cell surface19,42. To confirm the presence of INPNC-CH on the cell surface of Stenotrophomonas, cells were probed with rabbit polyclonal anti-CH antibodies and then fluorescently stained with tetramethylrhodamine isothiocyanate (TRITC)-labeled goat anti-rabbit IgG antibody. Since antibodies cannot diffuse through the outer membrane, only surface-exposed CH can interact with its specific antibodies. Under a fluorescence microscope, YC-1/pVIC3 cells with surface-displayed CH were brightly fluorescent, whereas YC-1/pVC3 cells expressing cytosolic CH were not immunostained (data not shown), indicating that the cell surface of Stenotrophomonas was covered with antibody-TRITC complex.
After treatment with proteinase K for 3.5 h, cells carrying pVIC3 were probed with polyclonal anti-CH antibodies and TRITC-conjugated IgG secondary antibody. However, cells were not immunostained when observed with a fluorescence microscope, which indicated that surface-exposed CH had been removed by proteinase K. From all of these results, we concluded that CH was displayed functionally on the cell surface of Stenotrophomonas using the truncated InaV as an anchoring motif.
CH activity and cell growth
Whole-cell CH activity of the recombinant Stenotrophomonas with surface-displayed CH (pVIC3) was 7.7-fold higher than that of the same strain with cytosolic CH (pVC3) (Fig. 2). This improvement in whole-cell CH activity may be attributed to the fact that the binding of CH to substrates is reinforced by the presentation of CH on cell surface. The recombinant strain with surface-displayed CH overcomes the mass transport limitation, and therefore it can be employed as a whole-cell biocatalyst for the hydrolysis of carbaryl. Moreover, the resting-cell suspension of the recombinant strain YC-1 with surface-displayed CH maintained the original CH activity during a 14-day incubation period (Fig. S2), suggesting that the enzyme is immobilized with enhanced structural stability on the outer cell membrane through the INP anchoring motif.
In this study, the effects of host strains on the efficiency of surface expression were also investigated. Whole-cell CH activity of the recombinant Stenotrophomonas carrying pVIC3 was 3.6-fold higher than that of E. coli DH5α carrying the same plasmid (Fig. 2). Subcellular fractionated samples of E. coli DH5α carrying pVIC3 were probed with polyclonal anti-CH antibodies. As a result, ~25% and ~75% of INPNC-CH fusion proteins were associated with the outer membrane and soluble fractions of DH5α/pVIC3 cells, respectively, as judged by the intensity of the protein bands (Fig. 1B), consistent with the activity distribution between the outer membrane (0.069 U/OD600) and soluble fractions (0.202 U/OD600). This high-level surface expression of CH in Stenotrophomonas is in accordance with previous report in which the INP system was used for cell surface display of OPH in Stenotrophomonas 42. Our results suggest that improved membrane translocation may occur with the INP system in Stenotrophomonas. This may be attributed to compatibility of the INP anchoring motif with the membrane structure of Stenotrophomonas since INP was originally isolated from strains of the genus Pseudomonas that has a close phylogenetic relationship with Stenotrophomonas 43. This study highlights the potential of Stenotrophomonas to be used as host strain for INP-mediated cell surface display.
Prior to IPTG induction, CH activity was not detected. Whole-cell CH activity increased gradually after induction with 0.5 mM IPTG and reached a maximum (0.24 U/OD600) at 24 h. Effects of different levels of induction on whole-cell CH activity were also investigated. Whole-cell CH activity increased gradually with increasing concentrations of IPTG and reached a maximum (0.239 U/OD600) at an IPTG concentration of 0.5 mM. However, induction with higher IPTG concentrations (0.6 to 1 mM) caused declines in whole-cell CH activity (Fig. 3), probably because a high transcription rate can block the translocation pathway of a secreted protein. The inhibitory effects of overexpression on protein translocation have been reported in previous studies on INP-mediated cell surface display of GFP and OPH20,25. Here, the optimal balance between protein expression and translocation was accomplished by the use of an inducible expression system.
Display of macromolecular heterologous proteins on the bacterial cell surface may result in instability of the outer membrane and growth inhibition of the cell21,22. To test whether surface display of CH inhibits cell growth, the growth kinetics of YC-1/pVIC3 and YC-1/pVC3 cells were compared. As expected, no growth inhibition was observed for YC-1/pVIC3 cells. YC-1/pVIC3 and YC-1/pVC3 cells showed similar growth profiles and reached a maximum OD600 of 1.681 and 1.787 at 32 h, respectively (Fig. 4). These results indicated that surface display of CH did not disturb the membrane structure or cause host growth defects. In previous studies, the INP-based display systems have been used to display several large heterologous proteins, such as 90 kDa chitinase28 and 77 kDa CPR29, on the cell surface of E. coli. In this study, we successfully expressed 88 kDa CH on the Stenotrophomonas cell surface using the truncated InaV as an anchoring motif, suggesting that the INP system is well suited as a carrier of relatively large inserts for functional expression of macromolecular foreign proteins on the surface of bacterial cell. To our knowledge, CH is the largest protein displayed with the INP system in bacteria other than E. coli so far.
Simultaneous hydrolysis of carbaryl and chlorpyrifos by the recombinant strain YC-1
In order to evaluate the pesticide hydrolysis capability of the recombinant strain YC-1 with surface-displayed CH, the hydrolysis experiments for carbaryl and chlorpyrifos were carried out with growing and resting cells of YC-1/pVIC3. As shown in Fig. 5A, 0.4 mM carbaryl and chlorpyrifos were completely hydrolyzed by growing cells (with an initial cell density of OD600 = 0.1) of YC-1/pVIC3 within 16 and 24 h, respectively. Moreover, 0.4 mM carbaryl and chlorpyrifos were completely hydrolyzed by resting-cell suspension (OD600 = 1.0) of YC-1/pVIC3 within 8 and 12 h, respectively (Fig. 5B). As expected, carbaryl could not be hydrolyzed by wild-type strain YC-1 (Fig. 5A), indicating that the recombinant strain YC-1 acquired the hydrolysis capability for carbaryl through the heterologous expression of CH. The recombinant strain YC-1 hydrolyzed chlorpyrifos as fast as wild-type strain YC-1 (Fig. 5A), indicating that surface expression of CH did not influence the intrinsic hydrolysis capability of strain YC-1 for chlorpyrifos. As expected, the concentration of carbaryl and chlorpyrifos did not change in non-inoculated control. These results suggest that the recombinant strain YC-1 can be employed for simultaneous hydrolysis of carbaryl and chlorpyrifos.
(A) Hydrolysis of carbaryl and chlorpyrifos by growing cells (with an initial cell density of OD600 = 0.1) of the wild-type and recombinant strain YC-1. (B) Hydrolysis of carbaryl and chlorpyrifos by resting-cell suspension (OD600 = 1.0) of the recombinant strain YC-1. Symbols: ▲, hydrolysis of chlorpyrifos by YC-1/pVIC3 cell; ■, hydrolysis of chlorpyrifos by YC-1 cell; ●, hydrolysis of carbaryl by YC-1/pVIC3 cell; ♦, hydrolysis of carbaryl by YC-1 cell. The data are mean values ± standard deviations of three replicates.
The recombinant strain YC-1 with endogenous CPH can transform chlorpyrifos to 3,5,6-trichloro-2-pyridinol (TCP) and diethylthiophosphoric acid (DETP) and utilize DETP as a carbon source, but TCP cannot be further degraded. The recombinant strain YC-1 with exogenous CH can hydrolyze carbaryl to 1-naphthol and methylamine, but the products cannot be further metabolized due to lack of other carbaryl-degrading genes.
Conclusions
Practical applications of large-scale enzymatic degradation have always been limited by the cost of purification and stability of the enzyme. The use of whole-cell biocatalysts is an alternative strategy for degradation of toxic pollutants. In this work, a whole-cell biocatalyst was developed for efficient hydrolysis of carbaryl and chlorpyrifos by the display of CH on the cell surface of a chlorpyrifos-degrading bacterium using an INP-based surface-anchoring system. Owing to its high activity and superior stability, the live biocatalyst is ideal for the decontamination of sites polluted with pesticides. Moreover, our findings highlight the potential of the INP system for the display of large proteins on cell surfaces of various environmental bacteria, and suggest that the species Stenotrophomonas may serve as a promising candidate for the development of versatile whole-cell biocatalysis systems by the cell surface display of a wide variety of novel enzymes.
Materials and Methods
Bacterial strains, plasmids, and culture conditions
A chlorpyrifos-degrading bacterium Stenotrophomonas sp. strain YC-1, which was isolated from activated sludge of the wastewater treating system of an organophosphorus pesticides manufacturer13, was used as host strain for cell surface display of CH. A surface expression vector, pVIC3, coding for INPNC-CH was used for targeting CH onto the cell surface of strain YC-1. An inpnc-cehA fusion gene was chemically synthesized by BGI Inc., Beijing, China and then ligated into pUC57-simple vector. The synthetic gene was released from pUC57-simple with EcoRI and HindIII and then subcloned into the same restriction sites of pVLT33, an Escherichia coli-Pseudomonas shuttle vector, to generate pVIC3. To construct a control plasmid expressing cytosolic CH, the cehA gene was amplified by PCR from pVIC3, digested with EcoRI and HindIII, and subcloned into the same restriction sites of pVLT33 to generate pVC3.
Strains bearing plasmids were grown in Luria-Bertani (LB) medium44 or M9 minimal medium45 supplemented with 50 μg/ml kanamycin at 30 °C. Transformation of plasmid into Stenotrophomonas was carried out using the electroporation method46. For the inducible expression of INPNC-CH fusion protein, 0.5 mM IPTG was added when the culture reached an OD600 of 0.4. After induction, the cultivation continued for additional 24 h at 30 °C.
Western blot analysis
Cells carrying pVIC3 were divided into total cell lysate and the outer membrane and soluble fractions by differential centrifugation19. Subcellular fractionated samples were analyzed by 12% SDS-PAGE44, after which protein samples were electroblotted using a semidry transfer system (Bio-Rad) onto a PVDF membrane (Roche). The membrane was blocked for 1 h in TBST (20 mM Tris-HCl, pH 7.5, 150 mM NaCl, 0.05% Tween 20) containing 3% BSA. Subsequently, the membrane was incubated with rabbit polyclonal anti-CH antibodies at a dilution of 1:1,000 for 2 h, after which the membrane was washed with TBST for 30 min and then incubated with a horseradish peroxidase-conjugated goat anti-rabbit IgG secondary antibody (Abcam) at a dilution of 1:2,000 for 1.5 h. Immunoreactive bands were detected by enhanced chemiluminescence using an ECL Plus kit (Amersham).
Immunofluorescence microscope
Cells carrying pVIC3 or pVC3 were resuspended (OD600 = 0.5) in 100 mM phosphate-buffered saline (PBS; pH 7.4) containing 3% BSA and then incubated with rabbit polyclonal anti-CH antibodies diluted (1:500) in PBS for 1.5 h at room temperature. After being washed with PBS, the cells were resuspended in PBS with goat anti-rabbit IgG antibody conjugated with TRITC (1:2,000 dilution, Invitrogen) and incubated for 1 h at room temperature. Prior to microscopic observation, cells were washed five times with PBS and mounted on poly(L-lysine)-coated microscopic slides. Photographs were taken using a Nikon fluorescence microscope equipped with TRITC and FITC filters.
CH activity assay
CH activity was assayed in a reaction mixture containing 0.5 mM carbaryl, 50 mM sodium phosphate buffer (pH 7.0), and 100 μl of cells (OD600 = 1.0) in a total volume of 1.0 ml. The reaction was started by adding the cells and incubated at 30 °C. The reaction was stopped by adding 100 μl of 2 mM HgCl2, and 1-naphthol produced was quantified by reacting with the fast blue B salt using colorimetric analysis6. Activities are expressed as units (1 μmol of 1-naphthol formed per minute at 30 °C) per OD600 of whole cells.
Proteinase accessibility experiment
Cells carrying pVIC3 or pVC3 were resuspended in 1 ml of 15 mM Tris-HCl buffer (pH 7.8) supplemented with 15% sucrose and 0.1 mM EDTA. Samples were incubated for 2.5, 3 or 3.5 h with 5 μl of 20 mg/ml proteinase K at room temperature. Proteinase treated and untreated cells were used for CH activity assay, Western blot analysis, and immunofluorescence microscope as described above.
Pesticide hydrolysis studies
YC-1/pVIC3 cells were harvested after incubation in LB medium plus kanamycin and resuspended (OD600 = 1.0) in M9 minimal medium. Subsequently, 10 ml of cell suspension was inoculated into 90 ml of M9 minimal medium supplemented with 0.2% glucose, 0.05% yeast extract, 50 μg/ml kanamycin, 0.5 mM IPTG, and 0.4 mM carbaryl and chlorpyrifos. Samples were incubated at 30 °C and 200 rpm and removed every 4 h for the quantitative analysis of carbaryl and chlorpyrifos using high performance liquid chromatography (HPLC). Pesticide extraction and HPLC analysis were carried out as described previously45.
YC-1/pVIC3 cells were inoculated at OD600 = 0.1 into M9 minimal medium supplemented with 0.2% glucose, 0.05% yeast extract, 50 μg/ml kanamycin, 0.5 mM IPTG, and 0.2 mM chlorpyrifos. After incubation for 24 h at 30 °C, cells were harvested, washed twice with 50 mM sodium phosphate buffer (pH 7.0), and resuspended (OD600 = 1.0) in the same buffer. Subsequently, the cell suspension was incubated with 0.4 mM carbaryl and chlorpyrifos at 200 rpm and 30 °C. The samples were withdrawn every 2 h and carbaryl and chlorpyrifos were quantified by HPLC.
References
Singh, B. K. & Walker, A. Microbial degradation of organophosphorus compounds. FEMS Microbiol. Rev. 30, 428–471 (2006).
Singh, B. K. Organophosphorus-degrading bacteria: ecology and industrial applications. Nat. Rev. Microbiol. 7, 156–164 (2009).
Hayatsu, M. & Nagata, T. Purification and characterization of carbaryl hydrolase from Blastobacter sp. strain M501. Appl. Environ. Microbiol. 59, 2121–2125 (1993).
Hayatsu, M., Hirano, M. & Nagata, T. Involvement of two plasmids in the degradation of carbaryl by Arthrobacter sp. strain RC100. Appl. Environ. Microbiol. 65, 1015–1019 (1999).
Hernándeza, A. F. et al. Toxic effects of pesticide mixtures at a molecular level: their relevance to human health. Toxicology 307, 136–145 (2013).
Hashimoto, M., Fukui, M., Hayano, K. & Hayatsu, M. Nucleotide sequence and genetic structure of a novel carbaryl hydrolase gene (cehA) from Rhizobium sp. strain AC100. Appl. Environ. Microbiol. 68, 1220–1227 (2002).
Hayatsu, M., Mizutani, A., Hashimoto, M., Sato, K. & Hayano, K. Purification and characterization of carbaryl hydrolase from Arthrobacter sp. RC100. FEMS Microbiol. Lett. 201, 99–103 (2001).
Swetha, V. P. & Phale, P. S. Metabolism of carbaryl via 1,2-dihydroxynaphthalene by soil isolates Pseudomonas sp. strains C4, C5, and C6. Appl. Environ. Microbiol. 71, 5951–5956 (2005).
Singh, R., Trivedi, V. D. & Phale, P. S. Metabolic regulation and chromosomal localization of carbaryl degradation pathway in Pseudomonas sp. strains C4, C5 and C6. Arch. Microbiol. 195, 521–535 (2013).
Trivedi, V. D., Jangir, P. K., Sharma, R. & Phale, P. S. Insights into functional and evolutionary analysis of carbaryl metabolic pathway from Pseudomonas sp. strain C5pp. Sci. Rep. 6, 38430 (2016).
Singh, B. K., Walker, A., Morgan, J. A. W. & Wright, D. J. Biodegradation of chlorpyrifos by Enterobacter strain B-14 and its use in bioremediation of contaminated soils. Appl. Environ. Microbiol. 70, 4855–4863 (2004).
Yang, L., Zhao, Y., Zhang, B., Yang, C. & Zhang, X. Isolation and characterization of a chlorpyrifos and 3,5,6-trichloro-2-pyridinol degrading bacterium. FEMS Microbiol. Lett. 251, 67–73 (2005).
Yang, C., Liu, N., Guo, X. & Qiao, C. Cloning of mpd gene from a chlorpyrifos-degrading bacterium and use of this strain in bioremediation of contaminated soil. FEMS Microbiol. Lett. 265, 118–125 (2006).
Li, X., He, J. & Li, S. Isolation of chlorpyrifos degrading bacterium Sphingomonas sp. strain Dsp-2 and cloning of the mpd gene. Res. Microbiol. 158, 143–149 (2007).
Xu, G. et al. Biodegradation of chlorpyrifos and 3,5,6-tricholoro-2-pyridinol by a newly isolated Paracoccus sp. strain TRP. Int. Biodeterior. Biodegrad. 62, 51–56 (2008).
Anwar, S., Liaquat, F., Khan, Q. M., Khalid, Z. M. & Iqbal, S. Biodegradation of chlorpyrifos and its hydrolysis product 3,5,6-tricholoro-2-pyridinol by Bacillus pumilus strain C2A1. J. Hazard. Mater. 168, 400–405 (2009).
Lu, P. et al. Biodegradation of chlorpyrifos and 3,5,6-trichloro-2-pyridinol by Cupriavidus sp. DT-1. Bioresour. Technol. 127, 337–342 (2013).
Shimazu, M., Mulchandani, A. & Chen, W. Simultaneous degradation of organophosphorus pesticides and p-nitrophenol by a genetically engineered Moraxella sp. with surface-expressed organophosphorus hydrolase. Biotechnol. Bioeng. 76, 318–324 (2001).
Yang, C. et al. Surface display of MPH on Pseudomonas putida JS444 using ice nucleation protein and its application in detoxification of organophosphates. Biotechnol. Bioeng. 99, 30–37 (2008).
Cao, X. et al. Simultaneous degradation of organophosphate and organochlorine pesticides by Sphingobium japonicum UT26 with surface-displayed organophosphorus hydrolase. Biodegradation 24, 295–303 (2013).
Wu, C. H., Mulchandani, A. & Chen, W. Versatile microbial surface-display for environmental remediation and biofuels production. Trends Microbiol. 16, 181–188 (2008).
Kim, C. S., Seo, J. H., Kang, D. G. & Cha, H. J. Engineered whole-cell biocatalyst-based detoxification and detection of neurotoxic organophosphate compounds. Biotechnol. Adv. 32, 652–662 (2014).
Kozloff, L. M., Turner, M. A. & Arellano, F. Formation of bacterial membrane ice-nucleation lipoglycoprotein complexes. J. Bacteriol. 173, 6528–6536 (1991).
Jung, H. C., Lebeault, J. M. & Pan, J. G. Surface display of Zymomonas mobilis levansucrase by using the ice-nucleation protein of Pseudomonas syringae. Nat. Biotechnol. 16, 576–580 (1998).
Li, L., Kang, D. G. & Cha, H. J. Functional display of foreign protein on surface of Escherichia coli using N-terminal domain of ice nucleation protein. Biotechnol. Bioeng. 85, 214–221 (2004).
Bae, W., Chen, W., Mulchandani, A. & Mehra, R. K. Enhanced bioaccumulation of heavy metals by bacterial cells displaying synthetic phytochelatins. Biotechnol. Bioeng. 70, 518–524 (2000).
Tang, X. et al. Cell surface display of organophosphorus hydrolase for sensitive spectrophotometric detection of p-nitrophenol substituted organophosphates. Enzyme Microb. Technol. 55, 107–112 (2014).
Wu, M. L., Tsai, C. Y. & Chen, T. H. Cell surface display of Chi92 on Escherichia coli using ice nucleation protein for improved catalytic and antifungal activity. FEMS Microbiol. Lett. 256, 119–125 (2006).
Yim, S. K. et al. Functional expression of mammalian NADPH-cytochrome P450 oxidoreductase on the cell surface of Escherichia coli. Protein Expr. Purif. 49, 292–298 (2006).
Abbaszadegan, M., Alum, A., Abbaszadegan, H. & Stout, V. Cell surface display of poliovirus receptor on Escherichia coli, a novel method for concentrating viral particles in water. Appl. Environ. Microbiol. 77, 5141–5148 (2011).
Fan, L. H., Liu, N., Yu, M. R., Yang, S. T. & Chen, H. L. Cell surface display of carbonic anhydrase on Escherichia coli using ice nucleation protein for CO2 sequestration. Biotechnol. Bioeng. 108, 2853–2864 (2011).
Liang, B., Lang, Q., Tang, X. & Liu, A. Simultaneously improving stability and specificity of cell surface displayed glucose dehydrogenase mutants to construct whole-cell biocatalyst for glucose biosensor application. Bioresour. Technol. 147, 492–498 (2013).
Song, J. et al. Bacterial cell-surface displaying of thermo-tolerant glutamate dehydrogenase and its application in L-glutamate assay. Enzyme Microb. Technol. 70, 72–78 (2015).
Liang, B., Li, L., Mascin, M. & Liu, A. Construction of xylose dehydrogenase displayed on the surface of bacteria using ice nucleation protein for sensitive D-xylose detection. Anal. Chem. 84, 275–282 (2012).
Feng, R. et al. Rational design of xylose dehydrogenase for improved thermostability and its application in development of efficient enzymatic biofuel cell. Enzyme Microb. Technol. 84, 78–85 (2016).
Bao, S. et al. Construction of a cell-surface display system based on the N-terminal domain of ice nucleation protein and its application in identification of mycoplasma adhesion proteins. J. Appl. Microbiol. 119, 236–244 (2015).
Niu, M. et al. Engineering bacterial surface displayed human norovirus capsid proteins: a novel system to explore interaction between norovirus and ligands. Front. Microbiol. 6, 1448 (2015).
Liu, A., Feng, R. & Liang, B. Microbial surface displaying formate dehydrogenase and its application in optical detection of formate. Enzyme Microb. Technol. 91, 59–65 (2016).
Zhang, Z. et al. Surface immobilization of human arginase-1 with an engineered ice nucleation protein display system in E. coli. PLoS One 11, e0160367 (2016).
Schmid, D. et al. Molecular organization of the ice nucleation protein InaV from Pseudomonas syringae. FEBS Lett. 414, 590–594 (1997).
de Lorenzo, V., Eltis, L., Kessler, B. & Timmis, K. N. Analysis of Pseudomonas gene products using lacI q/Ptrp-lac plasmids and transposons that confer conditional phenotypes. Gene 123, 17–24 (1993).
Liu, Z. et al. Simultaneous degradation of organophosphates and 4-substituted phenols by Stenotrophomonas species LZ-1 with surface-displayed organophosphorus hydrolase. J. Agric. Food Chem. 57, 6171–6177 (2009).
Ryan, R. P. et al. The versatility and adaptation of bacteria from the genus. Stenotrophomonas. Nat. Rev. Microbiol. 7, 514–525 (2009).
Green, M. R. & Sambrook, J. Molecular cloning: a laboratory manual (fourth edition). Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, USA (2012).
Gong, T. et al. Engineering Pseudomonas putida KT2440 for simultaneous degradation of carbofuran and chlorpyrifos. Microb. Biotechnol. 9, 792–800 (2016).
Cho, J. H., Kim, E. K. & So, J. S. Improved transfomation of Pseudomonas putida KT2440 by electroporation. Biotechnol. Tech. 9, 41–44 (1995).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 31570035), Open Fund of State Key Laboratory of Microbial Technology, Shandong University (No. M2016-03), and Open Fund of State Key Laboratory of Microbial Metabolism, Shanghai Jiao Tong University (No. MMLKF16-02).
Author information
Authors and Affiliations
Contributions
C.Y., Y.P.L. and R.H.L. designed the research. C.Y., X.Q.X., H.J. and Y.B.W. performed the research. C.Y., X.Q.X., Y.P.L., H.J., Y.B.W., P.X. and R.H.L. analyzed the data. C.Y., Y.P.L. and R.H.L. wrote the paper. All of the authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, C., Xu, X., Liu, Y. et al. Simultaneous hydrolysis of carbaryl and chlorpyrifos by Stenotrophomonas sp. strain YC-1 with surface-displayed carbaryl hydrolase. Sci Rep 7, 13391 (2017). https://doi.org/10.1038/s41598-017-13788-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-13788-0
- Springer Nature Limited