Abstract
The mechanisms underlying the fast genome evolution that occurs during animal domestication are poorly understood. Here, we present a genome-wide epigenetic dataset that quantifies DNA hydroxymethylation at single nucleotide resolution among full-sib Nile tilapia (Oreochromis niloticus) with distinct growth performance. In total, we obtained 355 million, 75 bp reads from 5 large- and 5 small-sized fish on an Illumina NextSeq500 platform. We identified several growth-related genes to be differentially hydroxymethylated, especially within gene bodies and promoters. Previously, we proposed that DNA hydroxymethylation greatly affects the earliest responses to adaptation and potentially drives genome evolution through its targeted enrichment and elevated nucleotide transversion rates. This dataset can be analysed in various contexts (e.g., epigenetics, evolution and growth) and compared to other epigenomic datasets in the future, namely DNA methylation and histone modifications. With forthcoming advancements in genome research, this hydroxymethylation dataset will also contribute to better understand the epigenetic regulation of key genomic features, such as cis-regulatory and transposable elements.
Similar content being viewed by others
Background & Summary
Animal domestication has been associated with early changes in gene expression1 and phenotypic divergence2, in addition to transcriptionally relevant genetic mutations that often require longer periods of time to emerge. Previous studies have shown that epigenetic modifications are likely involved in the process of adaptation under captivity2,3, overlap with genetic mutations after several generations and potentially drive genome evolution4. These recent findings have broadened our knowledge and understanding of how epigenetic modifications alter rapidly gene expression and contribute towards normal development and phenotypic divergence, including potentially pathological conditions such as diabetes5 and cancer6.
Growth is one of the main traits of interest in farmed animals intended for food production. One of the primary tissues involved in somatic growth is the liver, which is the main binding target of growth hormone and essential in insulin-growth factor I production. In teleosts, the liver is also an important organ for the synthesis of several yolk proteins (e.g., vitellogenin) which are transported to the oocytes for uptake7. Therefore, epigenetic modifications capable of altering hepatic gene expression in females have impact on egg size and quality, which are linked to the growth potential of the offspring8.
DNA 5-hydroxymethylcytosine (5hmC) is an oxidized derivative of 5-methylcytosine (5mC) that has received very little attention in the field of epigenetics despite its profound stability and association with critical molecular functions, namely gene transcription and histone accessibility9,10,11,12,13,14,15. It forms slowly right after DNA replication by ten-eleven translocation (TET) enzymes as opposed to 5mCs which are formed during DNA replication by DNA methyltransferases16. Although DNA hydroxymethylation has been characterized as a unique epigenetic modification with distinct roles in gene regulation17, methods such as reduced representation and whole genome bisulfite sequencing (RRBS and WGBS, respectively) produce data that make 5mCs and 5hmCs indistinguishable. To date, liver DNA hydroxymethylation and the role of TET enzymes have only been described in human diseases including hepatocellular carcinoma18 and non-alcoholic fatty liver disease (NAFLD)19. In our previous work, we have shown that DNA hydroxymethylation is an abundant epigenetic modification within the somatotropic axis, including the liver20. Interestingly, genes such as ppargc1a, which is associated with metabolic reprogramming and the coordination of glucose and fatty acid metabolism in the liver, are involved in the function of TET enzymes21.
Recently, we performed a genome-wide mapping of 5hmCs at single nucleotide resolution using reduced representation hydroxymethylation profiling (RRHP) and we captured 68% (1,096,820) of the total CCGG sites (1,613,446) across the Nile tilapia genome. Based on our data filtering methodology, we identified 138,000 significantly hydroxymethylated cytosines in two groups of Nile tilapia with distinct growth rates. Statistical analysis revealed 2,677 DhmCs (q < 0.05), out of which, 2,237 had significantly higher levels of DNA hydroxymethylation in large fish compared to their smaller counterparts (Supplementary file 1a-c; please see section “Data Records”).
Overall, we showed that DNA hydroxymethylation levels differ significantly in the liver of full-sib Nile tilapia with distinct growth rates during their early domestication. Our annotation revealed several differentially hydroxymethylated cytosines (DhmCs) that are particularly enriched within gene bodies and promoters, supporting their functional significance in transcriptional regulation. Furthermore, genes involved in growth and metabolic functions had higher DNA hydroxymethylation levels in large- than in small-sized fish (Supplementary file 2; please see section “Data Records”). Finally, we proposed that DNA hydroxymethylation is an epigenetic marker that likely regulates gene transcription and contributes to phenotypic divergence and adaptation in animals undergoing domestication. We are confident that this dataset has a great future potential because it addresses for the first time the changes of the DNA hydroxymethylome in phenotypically divergent full-sib animals during their early domestication. Forthcoming developments in genome annotation will likely provide valuable clues towards the understanding of how these epigenetic marks regulate gene expression, phenotypic divergence and genome evolution.
Methods
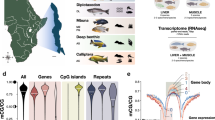
A more detailed description of the experimental and computational methodology is described in our sister publication2. Briefly, wild Nile tilapia were captured from Nile River in Luxor, Egypt. As a mouthbrooding fish species, female Nile tilapia with eggs in their mouths were captured and their eggs were collected and transported to our breeding facility in Bodø, Norway. After a successful reproduction cycle using our founder population (F0), we observed phenotypic divergence among full-siblings in the first generation born in captivity (F1).
Sample size (n) was calculated using two methods. First, because RRHP produces a count-based data set similar to RNA-Seq, 5 individuals with a sequencing depth of approximately 20 million reads are able to distinguish 10-fold transcriptional difference22. Furthermore, RRHP using human samples can detect low 5hmC signals (20%, q < 0.05) with only 20–30 million reads23. In this study, the average depth among libraries was over 36 million reads, even if the Nile tilapia genome contains only 1.6 million CCGGs compared to 4.6 million in the human genome. Second, sample size was also calculated by taking into consideration the weight and length of the fish. Since the endpoint of this analysis was quantitative data (5hmC counts), we used the formula [n = 2·(SD·(Z0 + Z≠)/d)2], where SD stands for standard deviation among all individuals, Z0 from Z table and a = 0.05, Z≠ from Z table (power at 80%) and d-difference between mean weights which resulted in n = 4.04.
Ethics approval
Animal handling and procedures were performed in accordance to the EU Directive 2010/63 on the use of animals for scientific purposes, and approved by the Norwegian Animal Research Authority (FOTS license no. 1042) and Nord University’s ethics committee.
Sampling procedure
In total, 10 full-sib Nile tilapia females (large- and small-size individuals, n = 5) were reared for 5 months in a 1500 L tank which was connected to the recirculating aquaculture system at Nord University’s research station (Fig. 1). All fish were caught by net and euthanized by clove oil overdose for 3 minutes at 28 °C. Pure clove oil (Sigma Aldrich, USA) was diluted in 1:10 (v/v) in 96% ethanol, and 20 ml of the mix was added for every 10 L of water in a 20 L oxygenated tank. Liver tissue was carefully dissected from the left lobe near the entry point of the portal vein. All samples were placed in 2 ml sterile cryogenic tubes, frozen instantly in liquid nitrogen and stored in −80 °C. The fish of this experiment represent our first generation (F1) of Nile tilapia born in captivity, as shown in Fig. 1. Detailed information on the animals used in this experiment can be found in Table 1.
Extraction of DNA for RRHP
DNA was extracted using the Quick-DNA miniprep plus kit (Zymo Research, USA), following the protocol recommended for solid tissues without any modifications. Briefly, 10 mg of liver was added in a solution of 95 μl DNAse and RNAse free water, 95 μl solid tissue buffer and 10 μl Proteinase K (20 mg/ml). The mixture was vortexed and incubated for 3 hours at 55 °C. The tissue was solubilized in 400 μl of genomic binding buffer. DNA integrity was measured using TapeStation 2200 (Agilent Technologies, USA) and DNA purity was determined on a NanoDrop Spectrophotometer 1000 (ThermoFisher Scientific, USA). Prior to library preparation, DNA concentration and subsequent dilutions were measured using the dsDNA broad range assay on a Qubit 3.0 fluorometer (ThermoFisher Scientific).
RRHP library preparation and sequencing
Library construction for reduced representation 5hmC profiling was performed according to the manufacturer’s protocol (RRHP – Zymo Research) as described in our sister paper2. For the present dataset, we used 200 ng of genomic DNA from liver as input for the preparation of every library including a positive 5hmC-irrelevant control. Genomic DNA was digested for 8 hours at 37 °C using the restriction enzyme MspI (Fig. 1). The DNA fragments were adapter ligated and extended with RRHP-specific adapters. Here, the P5 adapter reconstitutes the restriction site CCGG at the junction while the P7 destroys it. After the glucosylation step, fragments with glucosylated 5hmC modifications at the CCGG junctions were resistant to the second MspI digestion, and thus retained their P5 adapters as opposed to fragments with unmodified or methylated cytosines which were cleaved. For our control library, the second MspI digestion was omitted resulting in the amplification of all adapterized fragments (unrelated to 5hmC content). All libraries were size selected and amplified for a total of 10 cycles according to the protocol. Samples with higher levels of DNA hydroxymethylation produce a higher number of adapter ligated fragments; therefore, the libraries were sequenced with equal volume rather than equal molarity. Library integrity was assessed using DNA high-sensitivity Tapestation 2200 (Fig. 2a). The libraries were sequenced using two high output, 75-cycle sequencing kits in single-end reading mode, using Nord University’s NextSeq500 Illumina platform. For the control libraries, half of the volume was sequenced in each flow cell to test for flow cell effects and save sequencing depth. After verifying that the two controls had no flow cell effects, the two controls were merged into a single fastq file. For both flow cells, final library pools were diluted to 1.5 nM prior to sequencing and 34% PhiX was included to account for the low complexity (CCGG) in the first 4 sequencing cycles.
RRHP library integrity and quantification after size selection and purification using DNA high-sensitivity tape on a Tapestation 2200, and processing of sequenced reads. (a) From left to right, L is the DNA high-sensitivity ladder, BL1-BL5 represent the large-sized individuals, respectively. SL1-SL5 wells represent the small-sized individuals, respectively. C is the positive (5hmC-irrelevant) control library. Dark areas correspond to the size and concentration of the RRHP libraries after final purification. (b) A representative example of the per base sequence content for RRHP libraries during quality control using the FastQC program. (c) Visualization of the mean quality scores across all RRHP libraries using the MultiQC software.
Processing and alignment of RRHP sequenced reads
Base call files were demultiplexed and converted to fastq files using the bcl2fastq v2.20.0.422 software (Illumina). Samples were identified based on their index sequence and concatenated accordingly in LINUX environment. Quality control of the sequenced reads was performed using the FastQC program24. The per base sequence content revealed the low complexity (CCGG) at the start of most reads which is typical for RRHP libraries (Fig. 2b). The mean quality scores of all samples were plotted using the MultiQC program25 (Fig. 2c). Adaptor sequences were removed using trim_galore v0.4.426 (Supplementary File 3; please see section “Data Records”) and reads were aligned to the latest Nile tilapia genome27 using bowtie v0.12.828 (Supplementary File 4; please see section “Data Records”) with the parameters–chunkmbs 1000 -S -v 1 -n 1 -m 3–strata–best -p 32.
Extraction, filtering and differential analysis of RRHP data
Aligned reads starting with CCGG in both strands were extracted in text files using a custom Python script (Supplementary File 5; please see code availability statement). RRHP produces count-based datasets and their analysis is similar to that of RNA-Seq count-based datasets for differential expression. Therefore, the output text files were used as input in R where they were filtered twice based on 5hmC counts. At first, 5hmC positions were eliminated when ≥ 6 of all samples from both groups had zero counts. This threshold was applied to remove sites with low 5hmC levels and at the same time ensure that eliminated sites had zero counts across both groups. Since thresholds can be subjective, for the second step of filtering we calculated the median of 5hmC counts across the entire dataset (Ymax = 19), and removed 5hmC sites when ≥ 6 samples had counts below the median (Supplementary File 6; please see code availability statement). Using limma software29 in R, we log2 transformed the count data using the function voom and a linear model was fitted (function lmFit) using weighted least squares which incorporate the variance of each observation into the regression. Variability estimates were then adjusted using the eBayes function. The log-transformed dataset including the control libraries was plotted in a multidimensional scaling plot (Fig. 3). Differentially hydroxymethylated sites between the two groups were extracted based on the adjusted p value (q < 0.05). The false discovery rate (FDR) was controlled using the Benjamini-Hochberg correction, which is the default in limma.
DhmC annotation using HOMER
For the annotation of DhmCs, the annotatePeaks.pl perl script was used. This script is available upon installation of the HOMER software on LINUX systems (http://homer.ucsd.edu/homer/introduction/install.html). Since the Nile tilapia genome is not included in HOMER’s database, we performed a custom gene annotation as shown in the manual (http://homer.ucsd.edu/homer/ngs/annotation.html) and explained in detail within Supplementary File 6 (please see code availability statement). In brief, DhmC information from toptable output was adjusted to fulfil HOMER requirements. The resulting BED file had 6 columns separated by TABs. The first column is the chromosome information, second column - start position, third column - end position, forth column - unique peak ID (which can be digits from 1–2,677 counting the total number of DhmCs), fifth column can be empty or include any desired information to be retained after the annotation is completed (i.e. paste columns from toptable such as adjusted.p.values) and the sixth column - strand information (+/− or 0/1, where 0 = “+”, 1 = “-”).
Interactive visualisation of the data
An Integrative Genomics Viewer30 (IGV) session (Related Manuscript Files 1–6) has been created for exploration of 5hmC positions across the Nile tilapia genome in individual samples and visualisation of DhmC results as bar plots or heatmaps. Detailed instructions are provided in Supplementary File 7 (to access the IGV session please see section “Data Records”).
Data Records
The raw fastq files produced in this work are deposited in SRA under the accession number: SRP28505031 and consist of 2 controls (C1 and C2) derived from a single library sequenced in both flow cells, 5 liver samples (BL1-BL5) that represent the large-sized individuals and 5 liver samples (SL1-SL5) which represent the small-sized individuals (Table 2). Supplementary Files 1–4, as well as the IGV session (Supplementary File 7) for visualization of the RRHP dataset between large and small female Nile tilapia were deposited on DataverseNO32. The files can be found at the permanent DOI link: https://doi.org/10.18710/6VWVMQ.
Technical Validation
To avoid technical biases during library preparation, the samples were given random numbers prior to measuring DNA concentrations. The libraries were prepared simultaneously and the initial DNA concentration was the same for all samples. After final amplification and purification of all RRHP libraries, the groups were traced back and assigned to each sample. Differences in DNA hydroxymethylation levels across samples and groups were evident prior to sequencing as shown in Fig. 2a.
Overall, the sequencing quality of every library was high (average Phred Score > 32 per base pair) and it was measured using FastQC and MultiQC as shown in Fig. 2. Sequencing lane bias was tested by plotting the log-transformed count data among samples and positive controls in a multidimensional scaling plot (Fig. 3). The two controls were overlapping showing no significant differentiation; therefore, no action was needed regarding normalization of counts between flow cells.
Code availability
Supplementary files 5 and 6 were deposited in GitHub on 2022/12/2033. They can be found at the URL: https://github.com/IoannisKonstantinidis/RRHP_Code.
References
Christie, M. R., Marine, M. L., Fox, S. E., French, R. A. & Blouin, M. S. A single generation of domestication heritably alters the expression of hundreds of genes. Nat Commun 7, 10676, https://doi.org/10.1038/ncomms10676 (2016).
Konstantinidis, I. et al. Major gene expression changes and epigenetic remodelling in Nile tilapia muscle after just one generation of domestication. Epigenetics 15, 1052–1067, https://doi.org/10.1080/15592294.2020.1748914 (2020).
Le Luyer, J. et al. Parallel epigenetic modifications induced by hatchery rearing in a Pacific salmon. Proc Natl Acad Sci USA 114, 12964–12969, https://doi.org/10.1073/pnas.1711229114 (2017).
Anastasiadi, D. & Piferrer, F. Epimutations in developmental genes underlie the onset of domestication in farmed European sea bass. Mol Biol Evol 36, 2252–2264, https://doi.org/10.1093/molbev/msz153 (2019).
Hanson, M. A. & Godfrey, K. M. Genetics: Epigenetic mechanisms underlying type 2 diabetes mellitus. Nat Rev Endocrinol 11, 261–262, https://doi.org/10.1038/nrendo.2015.31 (2015).
Feinberg, A. P., Koldobskiy, M. A. & Gondor, A. Epigenetic modulators, modifiers and mediators in cancer aetiology and progression. Nat Rev Genet 17, 284–299, https://doi.org/10.1038/nrg.2016.13 (2016).
Arukwe, A. & Goksoyr, A. Eggshell and egg yolk proteins in fish: hepatic proteins for the next generation: oogenetic, population, and evolutionary implications of endocrine disruption. Comp Hepatol 2, 4, https://doi.org/10.1186/1476-5926-2-4 (2003).
Segers, F. H., Berishvili, G. & Taborsky, B. Egg size-dependent expression of growth hormone receptor accompanies compensatory growth in fish. Proc Biol Sci 279, 592–600, https://doi.org/10.1098/rspb.2011.1104 (2012).
Gao, D. et al. DNA methylation/hydroxymethylation regulate gene expression and alternative splicing during terminal granulopoiesis. Epigenomics 11, 95–109, https://doi.org/10.2217/epi-2018-0050 (2019).
Ponnaluri, V. K. et al. Association of 5-hydroxymethylation and 5-methylation of DNA cytosine with tissue-specific gene expression. Epigenetics 12, 123–138, https://doi.org/10.1080/15592294.2016.1265713 (2017).
Mellen, M., Ayata, P. & Heintz, N. 5-hydroxymethylcytosine accumulation in postmitotic neurons results in functional demethylation of expressed genes. Proc Natl Acad Sci USA 114, E7812–E7821, https://doi.org/10.1073/pnas.1708044114 (2017).
Gross, J. A. et al. Gene-body 5-hydroxymethylation is associated with gene expression changes in the prefrontal cortex of depressed individuals. Transl Psychiatry 7, e1119, https://doi.org/10.1038/tp.2017.93 (2017).
Bhattacharyya, S. et al. Altered hydroxymethylation is seen at regulatory regions in pancreatic cancer and regulates oncogenic pathways. Genome Res 27, 1830–1842, https://doi.org/10.1101/gr.222794.117 (2017).
Greco, C. M. et al. DNA hydroxymethylation controls cardiomyocyte gene expression in development and hypertrophy. Nat Commun 7, 12418, https://doi.org/10.1038/ncomms12418 (2016).
Wu, H. et al. Genome-wide analysis of 5-hydroxymethylcytosine distribution reveals its dual function in transcriptional regulation in mouse embryonic stem cells. Genes Dev 25, 679–684, https://doi.org/10.1101/gad.2036011 (2011).
Bachman, M. et al. 5-Hydroxymethylcytosine is a predominantly stable DNA modification. Nat Chem 6, 1049–1055, https://doi.org/10.1038/nchem.2064 (2014).
Kozlenkov, A. et al. A unique role for DNA (hydroxy)methylation in epigenetic regulation of human inhibitory neurons. Sci Adv 4, eaau6190, https://doi.org/10.1126/sciadv.aau6190 (2018).
Udali, S. et al. Global DNA methylation and hydroxymethylation differ in hepatocellular carcinoma and cholangiocarcinoma and relate to survival rate. Hepatology 62, 496–504, https://doi.org/10.1002/hep.27823 (2015).
Lyall, M. J. et al. Non-alcoholic fatty liver disease (NAFLD) is associated with dynamic changes in DNA hydroxymethylation. Epigenetics 15, 61–71, https://doi.org/10.1080/15592294.2019.1649527 (2020).
Konstantinidis, I. et al. Epigenetic mapping of the somatotropic axis in Nile tilapia reveals differential DNA hydroxymethylation marks associated with growth. Genomics 113, 2953–2964, https://doi.org/10.1016/j.ygeno.2021.06.037 (2021).
Sinton, M. C., Hay, D. C. & Drake, A. J. Metabolic control of gene transcription in non-alcoholic fatty liver disease: the role of the epigenome. Clin Epigenetics 11, 104, https://doi.org/10.1186/s13148-019-0702-5 (2019).
Ching, T., Huang, S. & Garmire, L. X. Power analysis and sample size estimation for RNA-Seq differential expression. RNA 20, 1684–1696, https://doi.org/10.1261/rna.046011.114 (2014).
Petterson, A., Chung, T. H., Tan, D., Sun, X. & Jia, X. Y. RRHP: a tag-based approach for 5-hydroxymethylcytosine mapping at single-site resolution. Genome Biol 15, 456, https://doi.org/10.1186/s13059-014-0456-5 (2014).
Andrews, S. FastQC: a quality control tool for high thoughput sequence data https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Ewels, P., Magnusson, M., Lundin, S. & Kaller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048, https://doi.org/10.1093/bioinformatics/btw354 (2016).
Krueger, F. Trim Galore: a wrapper tool around Cutadapt and FastQC to consistently apply quality and adapter trimming to FastQ files, with some extra functionality for MspI-digested RRBS-type (Reduced Representation Bisufite-Seq) libraries https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/ (2012).
Conte, M. A., Gammerdinger, W. J., Bartie, K. L., Penman, D. J. & Kocher, T. D. A high quality assembly of the Nile Tilapia (Oreochromis niloticus) genome reveals the structure of two sex determination regions. BMC Genomics 18, 341, https://doi.org/10.1186/s12864-017-3723-5 (2017).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10, R25, https://doi.org/10.1186/gb-2009-10-3-r25 (2009).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43, e47, https://doi.org/10.1093/nar/gkv007 (2015).
Robinson, J. T. et al. Integrative Genomics Viewer. Nature Biotechnol 29, 24–26, https://doi.org/10.1038/nbt.1754 (2011).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP285050 (2021).
Konstantinidis, I., Sætrom, P., & Fernandes, J. Supporting Files for: Genome-wide hydroxymethylation profiles in liver of female Nile tilapia with distinct growth performance. DataverseNO https://doi.org/10.18710/6VWVMQ (2023).
GitHub https://github.com/IoannisKonstantinidis/RRHP_Code (2022).
Acknowledgements
We are thankful to Øivind Torslett, Steinar Johnsen and Kaspar Klaudiussen for their assistance in maintenance of the RAS, husbandry and welfare of the fish. This study has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant agreement no 683210) and from the Research Council of Norway under the Toppforsk programme (grant agreement no 250548/F20).
Author information
Authors and Affiliations
Contributions
I.K. carried out the sampling, laboratory work and bioinformatic analysis, interpreted the data and wrote the data descriptor. P.S. contributed significantly towards the establishment of the bioinformatic pipeline and revised the data descriptor. J.M.O.F. conceived the study, provided reagents and consumables, contributed significantly to the interpretation of the data and revised the data descriptor.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Konstantinidis, I., Sætrom, P. & Fernandes, J.M.O. Genome-wide hydroxymethylation profiles in liver of female Nile tilapia with distinct growth performance. Sci Data 10, 114 (2023). https://doi.org/10.1038/s41597-023-01996-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-01996-5
- Springer Nature Limited