Abstract
Anion exchanger 1 (AE1), a member of the solute carrier (SLC) family, is the primary bicarbonate transporter in erythrocytes, regulating pH levels and CO2 transport between lungs and tissues. Previous studies characterized its role in erythrocyte structure and provided insight into transport regulation. However, key questions remain regarding substrate binding and transport, mechanisms of drug inhibition and modulation by membrane components. Here we present seven cryo-EM structures in apo, bicarbonate-bound and inhibitor-bound states. These, combined with uptake and computational studies, reveal important molecular features of substrate recognition and transport, and illuminate sterol binding sites, to elucidate distinct inhibitory mechanisms of research chemicals and prescription drugs. We further probe the substrate binding site via structure-based ligand screening, identifying an AE1 inhibitor. Together, our findings provide insight into mechanisms of solute carrier transport and inhibition.





Similar content being viewed by others
Data availability
Density maps and structure coordinates have been deposited in the Electron Microscopy Data Bank (EMDB) and the Protein Data Bank (PDB): AE1-apo (EMD-26165 and PDB 7TY4), AE1–bicarbonate (EMD-26168 and PDB 7TY7), AE1–DIDS (EMD-41082 and PDB 8T6V), AE1–H2DIDS (EMD-26167 and PDB 7TY6), AE1–DEPC (EMD-26171 and PDB 7TYA), AE1–dipyridamole (EMD-41081 and PDB 8T6U), AE1–NIF (EMD-26169 and PDB 7TY8). Source data are provided with this paper.
References
Elgsaeter, A., Stokke, B. T., Mikkelsen, A. & Branton, D. The molecular basis of erythrocyte shape. Science 234, 1217–1223 (1986).
Poole, J. The Diego blood group system–an update. Immunohematology 15, 135–143 (1999).
Levine, P., Robinson, E. A., Layrisse, M., Arends, T. & Domingues Sisco, R. The Diego blood factor. Nature 177, 40–41 (1956).
Reithmeier, R. A. et al. Band 3, the human red cell chloride/bicarbonate anion exchanger (AE1, SLC4A1), in a structural context. Biochim. Biophys. Acta 1858, 1507–1532 (2016).
Jennings, M. L. Cell physiology and molecular mechanism of anion transport by erythrocyte band 3/AE1. Am. J. Physiol. Cell Physiol. 321, C1028–C1059 (2021).
Kodippili, G. C. et al. Analysis of the mobilities of band 3 populations associated with ankyrin protein and junctional complexes in intact murine erythrocytes. J. Biol. Chem. 287, 4129–4138 (2012).
Thornell, I. M. & Bevensee, M. O. Regulators of Slc4 bicarbonate transporter activity. Front. Physiol. 6, 166 (2015).
Vallese, F. et al. Architecture of the human erythrocyte ankyrin-1 complex. Nat. Struct. Mol. Biol. 29, 706–718 (2022).
Xia, X., Liu, S. & Zhou, Z. H. Structure, dynamics and assembly of the ankyrin complex on human red blood cell membrane. Nat. Struct. Mol. Biol. 29, 698–705 (2022).
Falke, J. J. & Chan, S. I. Molecular mechanisms of band 3 inhibitors. 1. Transport site inhibitors. Biochemistry 25, 7888–7894 (1986).
Falke, J. J. & Chan, S. I. Molecular mechanisms of band 3 inhibitors. 2. Channel blockers. Biochemistry 25, 7895–7898 (1986).
Falke, J. J. & Chan, S. I. Molecular mechanisms of band 3 inhibitors. 3. Translocation inhibitors. Biochemistry 25, 7899–7906 (1986).
Galanter, W. L. & Labotka, R. J. The binding of nitrate to the human anion exchange protein (AE1) studied with 14N nuclear magnetic resonance. Biochim. Biophys. Acta 1079, 146–151 (1991).
Arakawa, T. et al. Crystal structure of the anion exchanger domain of human erythrocyte band 3. Science 350, 680–684 (2015).
Huynh, K. W. et al. CryoEM structure of the human SLC4A4 sodium-coupled acid-base transporter NBCe1. Nat. Commun. 9, 900 (2018).
Wang, W. et al. Cryo-EM structure of the sodium-driven chloride/bicarbonate exchanger NDCBE. Nat. Commun. 12, 5690 (2021).
Casey, J. R., Pirraglia, C. A. & Reithmeier, R. A. Enzymatic deglycosylation of human Band 3, the anion transport protein of the erythrocyte membrane. Effect on protein structure and transport properties. J. Biol. Chem. 267, 11940–11948 (1992).
Zhang, D., Kiyatkin, A., Bolin, J. T. & Low, P. S. Crystallographic structure and functional interpretation of the cytoplasmic domain of erythrocyte membrane band 3. Blood 96, 2925–2933 (2000).
Figueroa, D. The Diego blood group system: a review. Immunohematology 29, 73–81 (2013).
Daniels, G. L. et al. Blood group terminology 2004: from the International Society of Blood Transfusion committee on terminology for red cell surface antigens. Vox Sang. 87, 304–316 (2004).
Schubert, D. & Boss, K. Band 3 protein–cholesterol interactions in erythrocyte membranes. Possible role in anion transport and dependency on membrane phospholipid. FEBS Lett. 150, 4–8 (1982).
Kalli, A. C. & Reithmeier, R. A. F. Interaction of the human erythrocyte Band 3 anion exchanger 1 (AE1, SLC4A1) with lipids and glycophorin A: molecular organization of the Wright (Wr) blood group antigen. PLoS Comput. Biol. 14, e1006284 (2018).
Gregg, V. A. & Reithmeier, R. A. Effect of cholesterol on phosphate uptake by human red blood cells. FEBS Lett. 157, 159–164 (1983).
Zhu, Q. & Casey, J. R. The substrate anion selectivity filter in the human erythrocyte Cl−/HCO3− exchange protein, AE1. J. Biol. Chem. 279, 23565–23573 (2004).
Lu, F. et al. Structure and mechanism of the uracil transporter UraA. Nature 472, 243–246 (2011).
Case, D. A. et al. AMBER 2020 (Univ. California, San Francisco, 2020).
Guarnieri, F. & Mezei, M. Simulated annealing of chemical potential: a general procedure for locating bound waters. Application to the study of the differential hydration propensities of the major and minor grooves of DNA. J. Am. Chem. Soc. 118, 8493–8494 (1996).
Knauf, P. A., Law, F. Y., Leung, T. W., Gehret, A. U. & Perez, M. L. Substrate-dependent reversal of anion transport site orientation in the human red blood cell anion-exchange protein, AE1. Proc. Natl Acad. Sci. USA 99, 10861–10864 (2002).
Okubo, K., Kang, D., Hamasaki, N. & Jennings, M. L. Red blood cell band 3. Lysine 539 and lysine 851 react with the same H2DIDS (4,4′-diisothiocyanodihydrostilbene-2,2′-disulfonic acid) molecule. J. Biol. Chem. 269, 1918–1926 (1994).
Lu, J. & Boron, W. F. Reversible and irreversible interactions of DIDS with the human electrogenic Na/HCO3 cotransporter NBCe1-A: role of lysines in the KKMIK motif of TM5. Am. J. Physiol. Cell Physiol. 292, C1787–C1798 (2007).
Jin, X. R., Abe, Y., Li, C. Y. & Hamasaki, N. Histidine-834 of human erythrocyte band 3 has an essential role in the conformational changes that occur during the band 3-mediated anion exchange. Biochemistry 42, 12927–12932 (2003).
Izuhara, K., Okubo, K. & Hamasaki, N. Conformational change of band 3 protein induced by diethyl pyrocarbonate modification in human erythrocyte ghosts. Biochemistry 28, 4725–4728 (1989).
Liantonio, A. et al. Niflumic acid inhibits chloride conductance of rat skeletal muscle by directly inhibiting the CLC-1 channel and by increasing intracellular calcium. Br. J. Pharmacol. 150, 235–247 (2007).
Cousin, J. L. & Motais, R. Inhibition of anion permeability by amphiphilic compounds in human red cell: evidence for an interaction of niflumic acid with the band 3 protein. J. Membr. Biol. 46, 125–153 (1979).
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004).
Carlsson, J. et al. Ligand discovery from a dopamine D3 receptor homology model and crystal structure. Nat. Chem. Biol. 7, 769–778 (2011).
Maneri, L. R. & Low, P. S. Structural stability of the erythrocyte anion transporter, band 3, in different lipid environments. A differential scanning calorimetric study. J. Biol. Chem. 263, 16170–16178 (1988).
Maneri, L. R. & Low, P. S. Fatty acid composition of lipids which copurify with band 3. Biochem. Biophys. Res. Commun. 159, 1012–1019 (1989).
Stewart, A. K. et al. Functional characterization and modified rescue of novel AE1 mutation R730C associated with overhydrated cation leak stomatocytosis. Am. J. Physiol. Cell Physiol. 300, C1034–C1046 (2011).
Jennings, M. L. & Smith, J. S. Anion–proton cotransport through the human red blood cell band 3 protein. Role of glutamate 681. J. Biol. Chem. 267, 13964–13971 (1992).
Yang, E. et al. A Ser725Arg mutation in Band 3 abolishes transport function and leads to anemia and renal tubular acidosis. Blood 131, 1759–1763 (2018).
Zhekova, H. R. et al. Identification of multiple substrate binding sites in SLC4 transporters in the outward-facing conformation: insights into the transport mechanism. J. Biol. Chem. 296, 100724 (2021).
Thurtle-Schmidt, B. H. & Stroud, R. M. Structure of Bor1 supports an elevator transport mechanism for SLC4 anion exchangers. Proc. Natl Acad. Sci. USA 113, 10542–10546 (2016).
Ficici, E., Faraldo-Gomez, J. D., Jennings, M. L. & Forrest, L. R. Asymmetry of inverted-topology repeats in the AE1 anion exchanger suggests an elevator-like mechanism. J. Gen. Physiol. 149, 1149–1164 (2017).
Kalli, A. C. & Reithmeier, R. A. F. Organization and dynamics of the red blood cell band 3 anion exchanger SLC4A1: insights from molecular dynamics simulations. Front. Physiol. 13, 817945 (2022).
Colas, C., Ung, P. M. & Schlessinger, A. SLC transporters: structure, function, and drug discovery. MedChemComm 7, 1069–1081 (2016).
Garaeva, A. A. & Slotboom, D. J. Elevator-type mechanisms of membrane transport. Biochem. Soc. Trans. 48, 1227–1241 (2020).
Gasbjerg, P. K. & Brahm, J. Kinetics of bicarbonate and chloride transport in human red cell membranes. J. Gen. Physiol. 97, 321–349 (1991).
Garaeva, A. A., Guskov, A., Slotboom, D. J. & Paulino, C. A one-gate elevator mechanism for the human neutral amino acid transporter ASCT2. Nat. Commun. 10, 3427 (2019).
Gunn, R. B. & Frohlich, O. Asymmetry in the mechanism for anion exchange in human red blood cell membranes. Evidence for reciprocating sites that react with one transported anion at a time. J. Gen. Physiol. 74, 351–374 (1979).
Bjornsson, T. D. & Mahony, C. Clinical pharmacokinetics of dipyridamole. Thromb. Res. Suppl. 4, 93–104 (1983).
Legrum, B. & Passow, H. Inhibition of inorganic anion transport across the human red blood cell membrane by chloride-dependent association of dipyridamole with a stilbene disulfonate binding site on the band 3 protein. Biochim. Biophys. Acta 979, 193–207 (1989).
Offman, E. et al. Pharmacokinetics and pharmacodynamics of the antiplatelet combination aspirin (acetylsalicylic acid) plus extended-release dipyridamole are not altered by coadministration with the potent CYP2C19 inhibitor omeprazole. Am. J. Cardiovasc. Drugs 13, 113–120 (2013).
Saniabadi, A. R. et al. Dipyridamole increases human red blood cell deformability. Eur. J. Clin. Pharmacol. 42, 651–654 (1992).
Gimsa, J. & Ried, C. Do band 3 protein conformational changes mediate shape changes of human erythrocytes? Mol. Membr. Biol. 12, 247–254 (1995).
Wagner, J. R. et al. POVME 3.0: software for mapping binding pocket flexibility. J. Chem. Theory Comput. 13, 4584–4592 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Yamashita, K., Palmer, C. M., Burnley, T. & Murshudov, G. N. Cryo-EM single-particle structure refinement and map calculation using Servalcat. Acta Crystallogr. D Struct. Biol. 77, 1282–1291 (2021).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
The PyMOL Molecular Graphics System, Version 2.0 (Schrödinger, LLC).
Sanematsu, K. et al. Intracellular acidification is required for full activation of the sweet taste receptor by miraculin. Sci. Rep. 6, 22807 (2016).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Mezei, M. Simulaid: a simulation facilitator and analysis program. J. Comput. Chem. 31, 2658–2668 (2010).
Sastry, G. M., Adzhigirey, M., Day, T., Annabhimoju, R. & Sherman, W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 27, 221–234 (2013).
Lyne, P. D., Lamb, M. L. & Saeh, J. C. Accurate prediction of the relative potencies of members of a series of kinase inhibitors using molecular docking and MM-GBSA scoring. J. Med. Chem. 49, 4805–4808 (2006).
Sterling, T. & Irwin, J. J. ZINC 15—ligand discovery for everyone. J. Chem. Inf. Model 55, 2324–2337 (2015).
Jorgensen, W. L., Maxwell, D. S. & Tirado-Rives, J. Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J. Am. Chem. Soc. 118, 11225–11236 (1996).
Friesner, R. A. et al. Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J. Med. Chem. 49, 6177–6196 (2006).
Schlessinger, A. et al. Structure-based discovery of prescription drugs that interact with the norepinephrine transporter, NET. Proc. Natl Acad. Sci. USA 108, 15810–15815 (2011).
Adams, D. J. Grand canonical ensemble Monte Carlo for a Lennard–Jones Fluid. Mol. Phys. 29, 307–311 (1975).
Acknowledgements
This work was supported by NIH grant GM133504, a Sloan Research Fellowship in Neuroscience, an Edward Mallinckrodt, Jr. Foundation Grant, a McKnight Foundation Scholars Award (all to D.W.), NIH T32 Training Grant GM062754 and DA053558 (G.Z.), R01 CA277794 (A.S. and K.H.), NIH grant U01AG046170 (B.Z.), NIH grant RF1AG057440 (B.Z.), NIH grant R01AG068030 (B.Z.) and NIH grants R01DK073681, R01DK067555 and R01DK061659 (R.O.). Some of this work was performed at the National Center for cryo-EM Access and Training (NCCAT) and the Simons Electron Microscopy Center located at the New York Structural Biology Center, supported by the NIH Common Fund Transformative High Resolution Cryo-Electron Microscopy program (U24 GM129539,) and by grants from the Simons Foundation (SF349247) and NY State Assembly. We further acknowledge cryo-EM resources at the National Resource for Automated Molecular Microscopy located at the New York Structural Biology Center, supported by grants from the Simons Foundation (SF349247), NYSTAR and the NIH National Institute of General Medical Sciences (GM103310) with additional support from Agouron Institute (F00316) and NIH (OD019994). For additional data collection we are also grateful to staff at the Laboratory for BioMolecular Structure (LBMS), which is supported by the DOE Office of Biological and Environmental Research (KP160711). This work was supported in part through the computational resources and staff expertise provided by Scientific Computing at the Icahn School of Medicine at Mount Sinai. We also thank J. F. Fay for help with initial data processing.
Author information
Authors and Affiliations
Contributions
M.J.C. designed experiments, expressed and purified protein for grid freezing, collected data, refined structures and edited the manuscript. S.Y. and A.C.S. purified protein, prepared samples for grid freezing, established and performed functional assays and edited the manuscript. S.V. performed molecular docking calculations with help from Y.Z., and helped analyze the structures. G.Z. prepared grids for structure determination and assisted with data collection, processing and refinement. Y.K.M. helped with data processing and structure refinement. R.H. helped establish protein expression and purification. K.H. performed volume calculations. A.S. supervised docking and volume calculation experiments and helped write the paper. R.O. performed molecular simulations and SACP analysis of substrate binding with help from M.M. B.Z. contributed to the study design and supervised computational studies. D.W. designed experiments, analyzed the data, supervised the overall project and management and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Reinhart A. F. Reithmeier and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Katarzyna Ciazynska, in collaboration with the Nature Structural & Molecular Biology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cryo-EM structure determination of AE1 complexes as exemplified by AE1-DIDS complex.
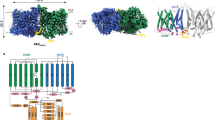
a, Analytical size exclusion chromatography and SDS-PAGE show monodisperse and pure protein of AE1-DIDS. Data were collected on 300 keV Krios, a representative micrograph is shown, and processed in cryoSPARC: particles were picked from motion corrected micrographs, subjected to 2D classification (representative classes are shown), followed by ab initio model building and 3D classification. After multiple rounds of 3D classification, the final particle stack was subjected to local CTF refinement followed by local refinement of the masked AE1 membrane domain with imposed C2 symmetry. Final map was obtained with GS-FSC indicating a resolution of 2.95 Å (AE1-DIDS) applying the 0.143 cutoff. Viewing direction distribution analysis (cryoSPARC) indicates sufficient coverage. An initial model was built in PHENIX, and then further refined in ServalCat for the generation of final maps and coordinates of mdAE1. Calculations in cryoSPARC indicate local resolutions of up to 2.5 Å around substrate and inhibitor binding sites. Viewing direction analysis indicates isotropic distribution of views in final particle stack. b, Superposition with other AE1 structures (AE1, PDB ID: 7TW2, dark red; AE1-Glycophorin complex, PDB ID: 7UZ3, lime green)7,8. Root mean square deviations of 0.997 Å (7TW2) and 0.372 Å (7UZ3) highlight similarity of protein conformation. Glycosylation modifications, cholesterols, lipids, waters, PIP2, and Glycophorin are shown in green, yellow, purple, red, grey, and teal, respectively. c, Cryo-EM structures allowed us to assign the complete extracellular surface including all Diego antigens (see Extended Data Table 2) with sidechains shown as spheres with yellow carbon atoms.
Extended Data Fig. 2 Cryo-EM Structure Determination of Apo AE1 and AE1-Bicarbonate complexes.
a, b, workflow of Apo AE1 (a) and AE1-Bicarbonate (b) data processing, showing representative 2D classes, followed by 2D classification, ab initio modeling and several rounds of 3D classification, with representative 3D classes shown. After multiple rounds of 3D classification, final particle stacks were subjected to CTF refinement followed by local refinement of the masked AE1 membrane domain with imposed C2 symmetry. GS-FSC calculations indicate global resolutions of 2.99 Å (Apo AE1) and 3.37 (AE1-Bicarbonate) applying the 0.143 cutoff. Viewing direction distribution analysis (cryoSPARC) indicates sufficient coverage. Initial models were built in PHENIX, and then further refined in ServalCat for the generation of final maps and coordinates. Calculations in cryoSPARC indicate local resolutions of lower than 3 Å around substrate and inhibitor binding sites.
Extended Data Fig. 3 Cryo-EM Structure Determination of AE1-H2DIDS and AE1-DEPC complexes.
a, b, workflow of AE1-H2DIDS (a) and AE1-DEPC (b) data processing, showing representative 2D classes, followed by 2D classification, ab initio modeling and several rounds of 3D classification, with representative 3D classes shown. After multiple rounds of 3D classification, final particle stacks were subjected to CTF refinement followed by local refinement of the masked AE1 membrane domain with imposed C2 symmetry. GS-FSC calculations indicate global resolutions of 2.98 Å (AE1-H2DIDS) and 3.07 (AE1-DEPC) applying the 0.143 cutoff. Viewing direction distribution analysis (cryoSPARC) indicates sufficient coverage. Initial models were built in PHENIX, and then further refined in ServalCat for the generation of final maps and coordinates. Calculations in cryoSPARC indicate local resolutions of lower than 3 Å around substrate and inhibitor binding sites.
Extended Data Fig. 4 Cryo-EM Structure Determination of AE1-Dipyridamole and AE1-NIF complexes.
a, b, workflow of AE1-Dipyridamole (a) and AE1-NIF (b) data processing, showing representative 2D classes, followed by 2D classification, ab initio modeling and several rounds of 3D classification, with representative 3D classes shown. After multiple rounds of 3D classification, final particle stacks were subjected to CTF refinement followed by local refinement of the masked AE1 membrane domain with imposed C2 symmetry. GS-FSC calculations indicate global resolutions of 3.13 Å (AE1-Dipyridamole) and 3.18 (AE1-NIF) applying the 0.143 cutoff. Viewing direction distribution analysis (cryoSPARC) indicates sufficient coverage. Initial models were built in PHENIX, and then further refined in ServalCat for the generation of final maps and coordinates. Calculations in cryoSPARC indicate local resolutions of lower than 3 Å around substrate and inhibitor binding sites.
Extended Data Fig. 5 Substrate-Binding in SLC4 and SLC26 Transporters.
a, Cryo-EM density of bicarbonate and binding site residues (marine mesh) using a contour level of 6σ. b, Binding site of uracil in the UraA-Uracil structure (PDB ID: 3QE7)24. c, d, Carbonate and sodium ions in the rabbit NDCBE (SLC4A8) cryo-EM structure (PDB ID: 7RTM, EMD-24683)15, with clashes between CO32− and T538, as well as Na+ and the backbone shown as red dotted lines. Ionic interactions and hydrogen bonds are shown as grey dotted lines. e, SACP simulations allow to calculate bicarbonate affinity to AE1 anion binding site (see methods). In some structures (~5%) we have identified a second bicarbonate binding site in the putative exit channel. f, Surface display shows putative anion exit channel leading from the bicarbonate binding site to the cytoplasmic site. Channel extends below the plane of section.
Extended Data Fig. 6 Structure and Density of AE1-Bound Inhibitors.
a, Chemical structures of different AE1 inhibitors and ligands used in this study. b, Cryo-EM density of DIDS (magenta) and key residues at a contour level of 6σ. c, H2DIDS (magenta) binding mode. d, Cryo-EM density of H2DIDS (magenta) and key residues at a contour level of 5σ. e, f, DEPC modifications of K539 and K851 (e) and corresponding cryo-EM densities (f) shown at a contour level of 8σ. g, h, Binding pocket of apo structure (g) and cryo-EM densities of residues (h) at a contour level of 6σ shown for comparison. i, Cryo-EM density of Dipyridamole (magenta) and key lysines at a contour level of 6σ. AE1’s gate and core domains are shown in tv blue and palecyan, respectively, and cryo-EM densities are shown as marine colored mesh.
Extended Data Fig. 7 Binding of NIF to AE1.
a-e, Close up of NIF binding site (a) and validation by molecular docking (b), with the best scoring docking pose shown (ΔG binding score: -52.33). c, Cryo-EM density of NIF and NIF binding site residues shown at a contour level of 3.5σ. d, Overlay of Apo (salmon) and NIF-bound AE1 (light blue) reveals subtle conformational rearrangements required for NIF binding. e, Calculation of NIF binding pocket volume and surface in POVME355. AE1’s gate and core domains are shown in tv blue and palecyan, respectively, and cryo-EM densities are shown as marine colored mesh.
Extended Data Fig. 8 Experimental testing of Compound 22 (Cmpd 22) analogs.
a, Inhibitory activity of 24 commercially available analogs of Cmpd 22 at 20 µM. Uptake experiments were performed with 2–6 technical repeats and are averaged from 3 independent experiments (n = 3). Data are represented as mean±s.e.m. b, Chemical structure of Cmpd 22 and selected analogs. c, d, Concentration response experiments show that Cmpd 22-J, Cmpd 22-R, and Cmpd 22-D have comparable inhibitory potencies as Cmpd 22 (c), while Cmpd 22-P and Cmpd 22-C appear to not inhibit AE1 (d). Concentration response uptake experiments were performed in triplicates and are averaged from 4 independent experiments (n = 4). Apparent potencies are calculated as apparent IC50 (mean) and error bars denote 95% confidence intervals. All uptake data have been normalized to Cmpd 22.
Supplementary information
Source data
Source Data Fig. 4 and Extended Data Figs. 1–5 and 8
AE1 uptake source data.
Source Data Extended Data Fig. 1
Size-exclusion chromatogram source data, FSC curve source data and uncropped SDS–PAGE image.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Capper, M.J., Yang, S., Stone, A.C. et al. Substrate binding and inhibition of the anion exchanger 1 transporter. Nat Struct Mol Biol 30, 1495–1504 (2023). https://doi.org/10.1038/s41594-023-01085-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-01085-6
- Springer Nature America, Inc.
This article is cited by
-
Mechanism of anion exchange and small-molecule inhibition of pendrin
Nature Communications (2024)