Abstract
The neurotensin receptor 1 (NTS1) is a G protein-coupled receptor (GPCR) with promise as a drug target for the treatment of pain, schizophrenia, obesity, addiction, and various cancers. A detailed picture of the NTS1 structural landscape has been established by X-ray crystallography and cryo-EM and yet, the molecular determinants for why a receptor couples to G protein versus arrestin transducers remain poorly defined. We used 13CεH3-methionine NMR spectroscopy to show that binding of phosphatidylinositol-4,5-bisphosphate (PIP2) to the receptor’s intracellular surface allosterically tunes the timescale of motions at the orthosteric pocket and conserved activation motifs – without dramatically altering the structural ensemble. β-arrestin-1 further remodels the receptor ensemble by reducing conformational exchange kinetics for a subset of resonances, whereas G protein coupling has little to no effect on exchange rates. A β-arrestin biased allosteric modulator transforms the NTS1:G protein complex into a concatenation of substates, without triggering transducer dissociation, suggesting that it may function by stabilizing signaling incompetent G protein conformations such as the non-canonical state. Together, our work demonstrates the importance of kinetic information to a complete picture of the GPCR activation landscape.
Similar content being viewed by others
Introduction
Cells rely on membrane-embedded receptors to maintain awareness of the extracellular environment without compromising membrane integrity. The G protein-coupled receptor (GPCR) superfamily is the largest group among such eukaryotic cell surface receptors, comprising more than 800 proteins1,2. They are ubiquitously expressed throughout the human body and are pivotal in a broad range of physiological processes including vision, taste, sense of smell, nervous functions, immune regulation, reproduction, and cancer3,4. Ligand binding at the extracellular, orthosteric site allosterically induces conformational changes across the GPCRs’ signature seven transmembrane (7TM) helices that prime the intracellular face for interaction with transducer proteins such as G proteins, β-arrestins (βArr), and G protein-coupled receptor kinases (GRKs)5. The neurotensin receptor 1 (NTS1) is a high-affinity target for the endogenous 13-residue peptide agonist neurotensin (NT)6. NT functions as a neuromodulator of the central nervous system (CNS) as well as a paracrine and endocrine modulator of the digestive tract and cardiovascular system7.
The strong expression overlaps of the dopamine system with both NT and NTS1 has led to considerable evidence for functional synergy in psychostimulant and opioid drug addiction8,9,10,11,12. Despite the long-standing interest in NTS1 as a potential therapeutic target for substance use disorders (SUDs), the handful of small-molecule NTS1 agonists and antagonists that have been developed all suffer from on-target side effects such as hypothermia13,14, hypotension15, and impaired motor control15,16. The classical model of GPCR activation implies that ligand-bound receptors signal equally (aka balanced) through G protein and β-arrestin (βArr) transducer pathways. The recent recognition of biased signaling, in which ligands preferentially activate one transducer pathway over the other, offers a new treatment avenue that may reduce on-target side effects17,18,19. A high-throughput functional screen and ligand optimization campaign targeting NTS1 led to the development of ML314, an allosteric ligand that selectively activates βArr2 pathways without stimulating the Gq pathway (i.e. βArr2-biased allosteric modulator; BAM), which reduces addictive behaviors toward methamphetamine and cocaine in several mouse models20,21. ML314 also functions as a βArr1-BAM22.
Yet, the molecular determinants for why a ligand promotes G protein:receptor versus arrestin:receptor complexation remain poorly defined. The predominant hypothesis is that agonists stabilize distinct receptor conformations to preferentially activate one pathway over another. Recent cryo-EM structures of NTS1:βArr1 and NTS1:G protein, however, reveal a remarkably conserved receptor architecture with a 0.67 Å all-atom RMSD23,24,25,26. Solution NMR has proven indispensable for identifying receptor conformers that are beyond the resolution of static structural methods27, but few studies have investigated G protein28,29,30 or βArr31,32,33 ternary complexes; to date, the only GPCRs characterized by NMR in complex with mimetics of both transducers are NTS133, M2 muscarinic34, and the β2-adrenergic receptor31,32,35,36. Here, we uniformly-label 13CεH3-methionine residues located within the NTS1 transmembrane bundle and near the ligand-binding site to demonstrate how ligands and PIP2 dynamically prepare the receptor for transducer interaction. The differential conformational kinetics upon coupling to βArr and G protein transducer molecules suggest a role for dynamics in functional selectivity.
Results
PIP2 strengthens correlated motions of the orthosteric pocket and PIF motif
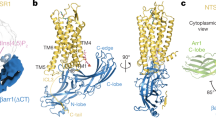
Phosphatidylinositol-4,5-bisphosphate (PIP2), and its analog (C8-PIP2; here termed PIP2), enhance both G protein activation37,38 and arrestin recruitment23. A NTS1:βArr1 cryo-EM complex includes one PIP2 molecule bound in a pocket formed by the intracellular portions of TMs 1/2/423; comparison with both canonical and non-canonical NTS1:G protein cryo-EM models suggests this PIP2 position could also coordinate the interaction of NTS1 with the Gαβ subunits24,25. Here, we used a previously characterized minimal methionine enNTS1 variant (herein enNTS1ΔM4)22 to test if PIP2 directly affects the receptor’s conformational ensemble in the absence of transducers. enNTS1ΔM4 was derived from a thermostabilized rat NTS1 (rNTS1) variant (enNTS1)39,40 by removing four of the ten endogenous methionines to eliminate signal overlap in NMR spectra. The six remaining residues (M2044.60, M2084.64, M2445.45, M2505.51, M3306.57, and M3527.36; superscript refers to Ballesteros-Weinstein numbering41) are located at the extracellular region, the base of the orthosteric pocket, and the PIF motif (Fig. 1a). Mutation of the four methionine residues did not adversely affect enNTS1’s structural integrity or function22. Two-dimensional (2D) 1H-13C heteronuclear multiple quantum correlation (HMQC) spectra were collected for apo, NT8-13 bound, ML314 bound, and NT8-13:ML314 bound [13CεH3-methionine]-enNTS1ΔM4 in the presence and absence of PIP2 (Fig. 1b–e). NT8-13 is an orthosteric ligand comprised of NT’s six C-terminal amino acids and is sufficient to produce full agonist activity42. ML314 is classified as a biased allosteric modulator (BAM) because it binds an alternative unknown site to potentiate NT8-13-mediated βArr1 recruitment while simultaneously attenuating agonist-induced G protein activation22. ML314 was reported to selectively stimulate βArr2 recruitment independent of NT8-1320,21, a pharmacological classification known as an agonist (ago)-BAM, although this was not observed in enNTS1ΔM4 or wildtype human NTS1 functional assays22.
a Cartoon representation of thermostabilized rNTS1 (PDB 4BWB) with labelled methionine methyl groups shown as yellow spheres (superscript - Ballesteros-Weinstein nomenclature41) and NT8-13 shown as purple sticks. Overlays of Apo-state (b), NT8-13 (c), ML314 (d), and NT8-13 & ML314 (e) bound 1H-13C HMQC spectra in the absence and presence of 130 μM (2x molecular equivalents over enNTS1) PIP2. The corresponding peak intensities are plotted in Supplemental Fig. S1. All spectra were recorded at 600 MHz, in 3 mm thin wall precision NMR tubes (Wilmad), with enNTS1ΔM4 concentrations of 66 μM.
All NMR spectra were collected at 66 μM [13CεH3-methionine]-enNTS1ΔM4 with identical acquisition, processing, and display parameters; thus, we can directly compare both the chemical shift values (i.e. structure) and signal intensities (i.e. dynamics) for each liganded state. Surprisingly, except for the NT8-13:ML314 bound state, PIP2 induced only subtle chemical shift perturbations indicating minimal effect on conformer populations (Figs. 1 and 2). However, when both agonist and BAM are present, PIP2 perturbs the M2044.60, M2445.45, and M2505.51 chemical shifts (i.e. pushes the structural equilibrium) towards the NT8-13 bound state (Figs. 1e and 2a, d, e). These chemical shift changes could reflect population changes towards more balanced NTS1:transducer complexes23,37,38, negative cooperativity between PIP2 and ML314 sites, or even direct binding competition.
Expanded views of individual resonances shown in Fig. 1. M2044.60 (a), M3306.57 (b), M3527.36 (c), M2445.54 (d), and M2505.51 (e) in the absence/presence of 130 μM (2× molecular equivalents over enNTS1) PIP2 for Apo-state (black/yellow), ML314-bound (orange/cyan), NT8-13-bound (magenta/dark green), and NT8-13:ML314-bound (blue/brown) enNTS1. The resonances of other residues within the extracted region are drawn at 50% transparency. The 1H 1D traces given above each spectrum correspond to the dashed lines indicated in each 2D spectrum below. All spectra were recorded at 600 MHz, in 3 mm thin wall precision NMR tubes (Wilmad), with enNTS1ΔM4 concentrations of 66 μM.
For the other liganded states, PIP2 primarily alters peak shapes and/or intensities, which reflect changes in receptor dynamics on the slow (ms-s) and fast-intermediate (ps-ms) timescales, respectively. In several instances, such as apo or ML314 bound M2044.60, a single uniform resonance develops multiplet character without adopting a new chemical shift value (Fig. 2a). The simplest explanation for this behavior is that the methyl group was rapidly exchanging between two or more distinct environments, and PIP2 slowed the interconversion rate without modifying the underlying nature of each conformer. Multiple NT8-13:ML314 bound resonances exhibit the opposite phenomena with peaks coalescing as PIP2 accelerates the rate of conformational exchange (Fig. 2b, d, e). In other cases, such as ML314-bound M3306.57 or apo M2445.45, PIP2 amends the relative intensity (i.e. equilibrium populations) of multiplet components (Figs. 2b, d and S1). Interestingly, PIP2 remodels the apo M2445.45 doublet intensity to match that of the M314-bound state, yet addition of PIP2 to ML314:enNTS1ΔM4 has no additional effect (Fig. 2d). Given that PIP2 may bind NTS1 at multiple locations, this may suggest partially overlapping interfaces or negative cooperatively between sites. A loss in peak intensity is typically caused by line broadening that may signify either changes in the intrinsic transverse relaxation rate (R2) and/or exchange broadening (Rex). R2 relaxation results from physical properties of the methyl group on the ps-ns timescale, while Rex reflects conformational interconversion in the μs-ms regime43. In the apo state, PIP2 sharpens M3306.57 adjacent to the ligand-binding pocket even as it broadens M2044.60 at the base of the pocket and M2505.51 of the connector region (Figs. 1b, 2, and Supplementary Fig. 1). The NT8-13 bound receptor resonances are universally broadened by PIP2 (Figs. 1c and 2 and Supplementary Fig. 1) whereas the ML314:enNTS1ΔM4 complex appears rigidified in the extracellular region (M3306.57), near the base of the orthosteric pocket (M2044.60), and surrounding the PIF motif (M2505.51; Figs. 1d and 2 and Supplementary Fig. 1).
The M3527.36 peak pattern in both NT8-13:enNTS1ΔM4 (Figs. 1c and 2c) and NT8-13:ML314:enNTS1ΔM4 (Figs. 1e and 2c) complexes reflects a multi-state equilibrium with mixed exchange regimes44. At first approximation, the presence of three peaks (states A, B, and C) indicates that M3527.36 exchanges between three chemical environments qualitatively on the slow (i.e. ms-s) timescale22,45. Density functional theory (DFT)-guided NMR analysis22 suggests that state A represents tight packing of TM1/6/7 and lid-like engagement of the N-terminus against the bound NT8-13 as observed in the NTSR1-H4X:NT8-13 X-ray structure (PDB 6YVR46), whereas M3527.36B may reflect detachment of the receptor N-terminus and local stabilization of extracellular TM1/6/7 as observed in the NTSR1-H4X:SR142948A structure (PDB 6Z4Q46). In the agonist bound state, PIP2 modestly reduced the total (sum of states A and B) M3527.36 peak intensity by 22.6%, which suggests a subtle adjustment towards a faster state A to state B interconversion rate although still within the ms-s timescale45. At the same time, the relative M3527.36B population increased by 138.8% while the M3527.36A state was effectively unchanged (Figs. 1c and 2c, and Supplementary Fig. 1). One potential explanation for this behavior is that state B is a composite of two microstates exchanging on the intermediate-fast (μs-ms) timescale. When both agonist and BAM are present, PIP2 increases the total (sum of states A and B) M3527.36 peak intensity by 164% without changing the relative state A and B populations (66 and 34%, respectively), which would correspond to a reduction of the A to B interconversion rate on the slow (ms-s) timescale45. The M3527.36B resonance is simultaneously perturbed upfield in the 1H dimension and downfield in the 13C dimension (Figs. 1e and 2c). There are several possible explanations for this behavior: (i) a change in the relative populations of fast-exchanging (μs-ms) microstates that comprise state B, (ii) structural changes of the M3527.36B chemical environment itself, and (iii) remodeling of both the thermodynamic and kinetic properties of states A and B. Taken together, PIP2 clearly remodels the receptor’s structural and kinetic ensemble. To better understand the nature of PIP2-mediated spectra changes, we investigated complexes of enNTS1ΔM4 with arrestin and G protein mimetics. For the remainder of this study, unless otherwise stated, all samples include PIP2.
βArr1 alters the kinetic landscape of the NTS1 conformational ensemble
Recent cryo-EM structures of the NTS1:βArr1 complex required either protein fusion26 or intermolecular cross-linking23 to stabilize intrinsic dynamics, suggesting that NMR could provide additional information on the nature of these underlying motions. We utilized the pre-activated βArr1-3A variant to maximize the affinity for unphosphorylated enNTS1ΔM447,48. Microscale thermophoresis (MST) measured the affinity of βArr1-3A for the NT8-13:enNTS1ΔM4 complex as 0.99 ± 0.1 μM, which corresponds to >98% receptor occupancy at a 2:1 βArr1-3A:enNTS1ΔM4 when assuming a single-site ligand depletion binding model (Fig. 3a). 2D transverse relaxation optimized spectroscopy (TROSY)-based 1H-15N heteronuclear single quantum correlation (HSQC) spectra of [U-2H,13C,15N]-βArr1-3A at increasing enNTS1ΔM4 concentrations confirm the formation of a specific complex (Supplementary Fig. 2a). The 2D 1H-13C HMQC of [13CεH3-methionine]-enNTS1ΔM4:βArr1-3A ternary complex formation resulted in the appearance of two additional methionine peaks (Fig. 3b; asterisks). Collecting another 1H-13C HMQC spectrum using both unlabeled enNTS1ΔM4 and βArr1-3A revealed that both peaks belong to βArr1-3A; while the major resonance is always detectable, the minor peak is only visible in the presence of the receptor (Supplementary Fig. 2b). Neither resonance is observable when the experiment is repeated using βArr1 C-terminally truncated after N382 (βArr1-ΔCT48). As arrestin recruitment requires displacement of its self-associated C-tail26,49, we conclude that the major and minor resonances correspond to βArr1’s C-terminal M411 in the bound-basal and dissociated receptor-bound (or post-receptor-bound) state, respectively. In the absence of PIP2, βArr1-3A promotes limited [13CεH3-methionine]-enNTS1ΔM4 spectral changes further supporting the lipid’s role in high affinity transducer complexation (Supplementary Fig. 2c).
a Microscale thermophoresis (MST) measured the affinity of βArr1-3A for NT8-13:enNTS1ΔM4 as 0.99 ± 0.1 μM ( ± SEM) using a single-site quadratic binding model. Data was collected as n = 3 biologically independent experiments with 5 or 6 technical repeats. Source data are provided as a Source Data file. Overlays of (b) NT8-13:enNTS1ΔM4 (forest green) and NT8-13:enNTS1ΔM4:βArr1-3A (purple), (c) ML314:enNTS1ΔM4 (cyan) and ML314:enNTS1ΔM4:βArr1-3A (tan), and (d) NT8-13:ML314:enNTS1ΔM4 (maroon) and NT8-13:ML314:enNTS1ΔM4:βArr1-3A (royal blue) 1H-13C HMQC spectra. Transducer spectra included 2.3x molar equivalents βArr1-3A. Peaks marked with an asterisk represent natural abundance βArr1-3A M411 (also see Supplementary Fig. 2b). Extracted spectral region of (e) M2445.45 and (f) M2505.51 from NT8-13:enNTS1ΔM4:βArr1-3A (purple), ML314:enNTS1ΔM4:βArr1-3A (tan), and NT8-13:ML314:enNTS1ΔM4:βArr1-3A (royal blue) 1H-13C HMQC spectra. One dimensional 1H cross-sectional slices (corresponding to dotted line) shown on top. Dots denote the residue’s chemical shift position in spectra of the corresponding colour with additional dots shown for ligand-only spectra (NT8-13:enNTS1ΔM4, forest green; ML314:enNTS1ΔM4, cyan; NT8-13:ML314:enNTS1ΔM4, maroon). All spectra were recorded at 600 MHz with receptor concentrations of 66 μM.
βArr1-3A leads to exchange broadening of M2044.60, M3527.36 state A and presumably M3527.36 state B, although the latter is overlapped with the major βArr1-3A M411 resonance (Fig. 3b; asterisks). To better understand the chemical exchange kinetics, we performed a titration series with increasing βArr1-3A concentrations added to separate, otherwise identical, NT8-13:enNTS1ΔM4 samples (Supplementary Fig. 3). The intensity of every resonance decreased in a concentration-dependent manner reflecting the ternary complex’s longer rotational correlation time. The especially rapid broadening of M3527.36A suggests that arrestin back-coupling enhances μs-ms exchange kinetics at the periphery of the orthosteric pocket (Supplementary Fig. 3); this is consistent with the lack of density for the homologous hNTS1 M3467.36 sidechain in both NT8-13:hNTS1:βArr1 cryo-EM models23,26. The three methionines nearest to the transducer interface (M2044.60, M2445.45, and M2505.51) all split into at least two distinct conformational states exchanging on the ms-s timescale (Supplementary Fig. 3g–i). The major M2445.45 peak settled at a chemical shift position linearly between the apo- and NT8-13 bound states (Supplementary Fig. 3h) with a second peak produced at a similar chemical shift observed for the apo- and ML314 bound states (Fig. 1b, d). βArr1-3A splits M2505.51 into two peaks centered at the NT8-13-bound chemical shift (Figs. 1c, 3b, and Supplementary Fig. 3I). Taken together, this indicates βArr1-3A is modulating the exchange kinetics of pre-existing agonist-bound conformations of the PIF motif – consistent with the proposed β2-adrenergic receptor activation mechanism50.
BAM potentiates the exchange dynamics of the βArr-ternary complex
To test if exchange kinetics of the receptor’s conformational ensemble plays a general role in arrestin activation, we investigated ML314 ternary and ML314:NT8-13 quaternary complexes. The ML314:enNTS1ΔM4:βArr1-3A complex spectrum was qualitatively quite similar to NT8-13:enNTS1ΔM4:βArr1-3A with enhanced μs-ms exchange peripheral to the orthosteric pocket and slower ms-s motions near to the transducer interface (Fig. 3b, c). Yet, there were several key differences. M3527.36 state B remained visible at 2.3 molar equivalents βArr1-3A and even increased in intensity relative to ML314 alone (Fig. 3c); thus βArr1-3A stabilizes the ML314 bound enNTS1ΔM4 conformer, which we hypothesize reflects a detached N-terminus and tightly packed TM1/TM2/TM7 interface22. M2445.45 splits into at least five peaks including substates populated in the presence of ML314 and NT8-13:βArr1-3A (Fig. 3b–e). ML314 maintains M2505.51 in the furthest downfield position of any ligand with βArr1-3A perturbing it even further and simultaneously splitting it into an ensemble of at least three substates (Fig. 3c, f). As βArr1-3A also pushes M2505.51 downfield relative to NT8-13:enNTS1ΔM4, we hypothesize this chemical environment signifies a transducer-competent conformer (Fig. 3b). The simultaneous addition of ML314 and NT8-13 to enNTS1ΔM4:βArr1-3A collapses both M2445.45 and M2505.51 to a subset of resonances that more generally reflect a concatenation of the ML314:βArr1-3A and NT8-13:βArr1-3A ternary complexes (Fig. 3b–f).
NTS1:Gαiq conformational and kinetic ensemble is distinct from NTS1:βArr1-3A
NTS1 can uniquely couple to all major Gα protein subtypes (Gαq/11, Gαi/o, Gαs, and Gα12/13)51 with the strongest preference towards Gq activation. This reflects a higher affinity and nucleotide exchange rate for Gαq, at least compared to Gαi, primarily driven by the six C-terminal residues of the α5 helix52. Since Gαq is inherently unstable53,54, we took advantage of the Gαiq chimera originally used to demonstrate that coupling specificity can largely be reduced to the G protein C-terminus52. The enNTS1ΔM4:Gαiq chimera complex affinity was 1.12 ± 0.1 μM, which corresponds to >98% receptor occupancy at a 2:1 transducer:receptor ratio and assuming a single-site ligand depletion binding model (Fig. 4a). NT8-13:enNTS1ΔM4:Gαiq complex formation was further supported by concentration-dependent changes in 1D 1H and 2D 1H-13C HMQC spectra of [13CεH3-methionine]-enNTS1ΔM4:Gαiq (Fig. 4b and Supplementary Fig. 4) as well as a 1H-15N TROSY-HSQC spectrum of [U-15N,13C,2H]-[Gαiq:enNTS1ΔM4 (Supplementary Fig. 5a). All samples included apyrase to maximize complex affinity24,55, and tris-(2-carboxyehtyl)-phosphine (TCEP) to limit Gαiq chimera self-association, which both had little to no observable effect on receptor resonances (Supplementary Fig. 5b). Two additional resonances, originating from unlabeled Gαiq protein, are observed in the 2D 1H-13C HMQC spectrum but cannot be assigned to specific residues (Supplementary Fig. 6a).
a Microscale thermophoresis (MST) measured the affinity of Gαiq for NT8-13:enNTS1ΔM4 as 1.12 ± 0.1 μM (±SEM) using a single-site quadratic binding model. Data was collected as n = 3 biologically independent experiments in technical triplicates. Source data are provided as a Source Data file. b 1H 1D spectra of three unassigned Gαiq resonances exhibiting enNTS1ΔM4-dependent chemical shift perturbations. Spectra were recorded at 600 MHz with receptor concentrations of 64 μM and variable Gαiq concentrations: 0 μM (0×), 96 μM (1.5×), and 192 μM (3×).
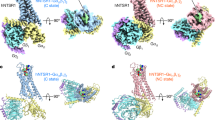
A comparison of the Gαiq and βArr1-3A titration series is particularly revealing (Fig. 5a–d and Supplementary Figs. 3 and 4). M2505.51 and M3306.57 concomitantly split into multiple, overlapping resonances in both ternary complexes, but βArr1-3A qualitatively promotes these changes at a slightly lower relative concentration. It is reasonable to anticipate a subset of similarly behaving resonances considering the highly conserved receptor architecture observed across all transducer complex structures and presumably partially-overlapped allosteric coupling networks23,24,25,26. There are three striking differences between Gαiq and βArr1-3A ternary complex spectra. First, M2445.45 remains a single unperturbed resonance in the presence of up to 3x molar excess of Gαiq. We do not observe any changes, apart from a subtle intensity reduction, suggesting that (i) NT8-13 alone induces a fully-active, G protein-competent M2445.45 conformation; (ii) the isolated Gαiq subunit is insufficient to stabilize the fully-active state; or (iii) that M2445.45 does not play a role in Gαiq coupling. Secondly, both transducers split M3527.36 state A into at least two resonances but βArr1-3A leads to exchange broadening at lower concentrations (Fig. 5a–d and Supplementary Figs. 3k and 4k). Finally, Gαiq stabilizes three M2044.60 resonances while βArr1-3A broadens all peaks before selecting a single state at 3x molar equivalents (Fig. 5a–d and Supplementary Figs. 3g and 4g). A detailed comparison of M2044.60 is challenging due to the overall weak intensities and similar resonance patterns observed at sub-stoichiometric transducer concentrations (Fig. 5a–d and Supplementary Figs. 3g and 4g). Yet, taken together, these chemical shift and intensity changes suggest that Gαiq and βArr1-3A differentially modulate the kinetics of enNTS1ΔM4 conformational ensembles near the connector region and orthosteric binding pocket.
a Overlay of NT8-13:enNTS1ΔM4:βArr1-3A (purple) and NT8-13:enNTS1ΔM4:Gαiq (tomato) 1H-13C HMQC spectra. Peaks marked with an asterisk represent natural abundance βArr1-3A and Gαiq resonances (Supplementary Figs. 2b and 6a). b Overlay of NT8-13:ML314:enNTS1ΔM4:βArr1-3A (royal blue) and NT8-13:ML314:enNTS1ΔM4:Gαiq (medium purple) 1H-13C HMQC spectra. Peaks marked with an asterisk represent natural abundance βArr1-3A and Gαiq resonances. Extracted spectra regions of enNTS1ΔM4:Gαiq and enNTS1ΔM4:βArr1-3A methionine resonances in the presence of (c) NT8-13 and (d) NT8-13:ML314. One dimensional 1H cross-sectional slices (corresponding to dashed lines in 2D spectra) shown on top. Dots denote the residue’s chemical shift position in spectra of the corresponding color with additional dots shown for ligand-only spectra (NT8-13:enNTS1ΔM4, forest green; NT8-13:ML314:enNTS1ΔM4, maroon). Gαiq containing spectra were recorded at 600 MHz with receptor concentrations of 64 μM and βArr1-3A containing spectra with receptor concentration of 66 μM.
BAM stabilizes a distinct Gαiq quaternary complex
Although multiple studies confirm that ML314 attenuates G protein activation and downstream signaling20,21,22, it’s unclear if this results from a reduction in NTS1-G protein complex affinity, nucleotide exchange rate, or unproductive complex conformation. Surprisingly, MST measurements reveal that ML314 only modestly reduces the affinity of Gαiq for NT8-13:enNTS1ΔM4 to 1.76 ± 0.2 μM (Supplementary Fig. 6b). We collected a ML314:NT8-13:enNTS1ΔM4:Gαiq spectrum for insight into the mechanism by which ML314 attenuates G protein activation. We observe differential effects across the receptor with residues adjacent to the orthosteric pocket appearing to adopt more βArr1-competent-like conformers and those near the transducer-binding interface continuing to populate Gαiq-competent conformers. For example, ML314 reduces the M3527.36 state A intensity as observed in βArr1 ternary and quaternary complexes (Fig. 5a–d and Supplementary Figs. 3 and 6). At the bottom of the orthosteric pocket, ML314 again pushes M2044.60 along a linear trajectory towards a state that is only observed in βArr1-3A ternary and quaternary complexes, we hypothesize this may affect the hydrogen-bond network22 that governs receptor activation (Supplementary Fig. 3g). M2445.45 shows no substantial difference, apart from a subtle reduction in peak intensity, between NT8-13:enNTS1ΔM4, ML314:NT8-13:enNTS1ΔM4 and NT8-13:enNTS1ΔM4:Gαiq (Supplementary Fig. 6). Whereas residue M2505.51, the closest probe to the transducer interface, begins to reflect a concatenation of Gαiq and βArr1-competent states (Fig. 5a–d and Supplementary Fig. 6). Perhaps the most dramatic change is observed in the M3306.57 multiplet pattern. While titration of either transducer initially splits M3306.57 in the 1H dimension, βArr1 ultimately stabilizes the downfield resonance (Supplementary Figs. 3 and 4).
Discussion
Over the last two decades, extensive crystallographic and cryo-electron microscopy studies have laid a structural foundation for GPCR activation in terms of inactive, intermediate, and active-state models. Sophisticated spectroscopic and computational studies expanded the conformational landscape to include high energy intermediate states from the fs-ns fluctuation of bond angles and side chain rotamers56 to the ns-μs toggling of microswitches36,50,57,58, the μs-ms conformational exchange of secondary structure35,59, and the ms-s activation of transducers59,60. Site-selective NMR labeling strategies, which are sensitive to molecular motions over the picosecond to second timescale, have proven especially powerful for describing how ligands and transducers remodel the GPCR conformational landscape27,61. Here, we employed endogenous 13CεH3-methionine probes located around the extracellular vestibule and near the connector region to expand our understanding of how ligands, allosteric modulators and transducers regulate NTS1 motions.
X-ray crystallography and MD studies suggest that ligand binding is communicated to the transducer interface through correlated motions near the connector or transmission region50,62. To further explore PIP2-mediated cooperativity between the orthosteric pocket (M3306.57) and connector region (M2445.45 and M2505.51), we measured the pairwise correlation of normalized peak intensities for each liganded state (Supplementary Fig. 7). As all spectra were collected using the same acquisition parameters on identically-prepared samples, the differential peak volumes should reflect individual or cumulative dynamics across the fast (ps-ns), intermediate (μs-ms), and slow (ms-s) NMR timescales. For example, ps-ns motions suggested by DFT analysis22,63 could alter peak intensities through transverse (T2) or longitudinal (T1) relaxation mechanisms64, although we hypothesize that the SOFAST-HMQC experiments employed dramatically reduce the possibility of differential T1 relaxation65. Slow exchange dynamics, such as peak splitting, were controlled for by summing the intensities across all resonances of a given residue; thus, we attribute differences to motions on the sub-microsecond timescale. Nonetheless, future experiments will be required to quantitate the T1, T2, and generalized order parameters for each methionine methyl group.
This analysis relies on the assumption that residue pairs involved in the same allosteric network will exhibit a concerted response – reminiscent of chemical shift covariance analysis (CHESCA)66 and methionine chemical shift-based order parameter analysis22,63. Residues M2445.45 and M2505.51 which are located before and after P2495.50, respectively, inform on the dynamics across the TM5 kink; the effect is relatively consistent regardless of PIP2, although the better linear correlation suggests an improved dynamic scaling between those residues. Similar results are observed for the pairwise correlation of M2445.45 and M3306.57, hinting at a subtle allosteric effect throughout the extracellular vestibule. Lastly, we looked at the correlation between M2505.51 and M3306.57. In the absence of PIP2, the two residues are effectively uncoupled (R2 = 0.14) but lipid addition increases the R2 to 0.69 and the slope to 0.99. The modest pairwise peak intensity correlations (Supplementary Fig. 7) between the orthosteric pocket and connector region alludes to a long-range allosteric coupling (Fig. 6a) and potential mechanism for PIP2’s ability to stabilize active states by strengthening the pairwise peak intensity correlations.
a The weak pair-wise correlation of peak intensities near the orthosteric pocket and connector region are strengthened by PIP2 to reveal several long-range allosteric communication pipelines. b β-arrestin-1 association slows the timescale of M2444.45 and M2505.51 conformational exchange whereas G protein coupling has little to no effect. Our NMR spectra qualitatively suggest the pre-existence of transducer-competent conformations in the agonist-bound state and that ML314, a β-arrestin biased allosteric modulator (BAM), fine-tunes exchange between those states.
Linking our observation to a specific enNTS1ΔM4:PIP2 interface is challenging. A hNTS1:βArr1-ΔCT cryo-EM structure23 and a native mass spectrometry (MS) approach38 both indicate PIP2 binds at the TM1/2/4 groove; however, the same MS study proposes an additional interface formed by TM1/738. Binding is dominated by electrostatic interactions between the polyanionic phosphorylated inositol head group of PIP2 and basic residues (Arg and Lys) of the receptor23,38,67,68. In the case of enNTS1, directed evolution resulted in three mutations (N2625.63R, K2635.64R and H3056.32R) at the intracellular tips of TMs 5 and 6, which could elevate the relevance of this second site39,40. NT8-13:ML314:enNTS1ΔM4 was the only ligand complex to exhibit substantial chemical shift changes upon addition of PIP2 (Fig. 1e). Perhaps this reflects counteracting allosteric networks, or perhaps even direct binding competition, of a balanced (PIP2) and biased (ML314) ligand in the absence of transducer. The latter may also be inferred from the Apo-M2445.45 peak splitting observed on addition of PIP2 (Fig. 2d) resulting in a similar peak shape and intensity compared to ML314:enNTS1ΔM4 and ML314:enNTS1ΔM4:PIP2. However, we cannot exclude the possibility that spectral changes are unrelated to transducer-competent conformations.
Recent cryo-EM structures of hNTS1:βArr1 and hNTS1:Gαiβγ ternary complexes possess very high receptor structural similarity, which raises questions as to the role of conformational dynamics in functional selectivity. Although technically challenging, other studies have also begun to integrate kinetic information for a more complete description of conformational landscapes69,70,71. Both M2445.45 and, to a lesser extent, M2505.51 resonances split in response to βArr1 but not Gαiq (Figs. 5c and 6b). This peak pattern is indicative of a chemical exchange process on the slow NMR timescale suggesting a reasonably high activation barrier between substates. Interestingly, the βArr-biased allosteric modulator ML314 alone (or in combination with NT8-13) also induces splitting of those resonances while PIP2 splits Apo-state M2445.45 analogous to M314, suggesting pre-selection of βArr-binding-competent states and offering a mechanistic basis for transducer-bias. Although speculative, we hypothesize that peak splitting in the 1H dimension may originate from local fluctuations of one or several aromatic rings22,63. Finally, it is interesting to note the continued presence of Gαiq competent chemical shifts in the ML314:NT8-13:enNTS1ΔM4:Gαiq spectrum (Fig. 4c–f). Similar studies using enNTS1 19F-labeled at the cytoplasmic tip of TM6 (Q301C6.28)33 reveal that a peptide corresponding to the Gq α5-helix binds ML314:enNTS1ΔM4 complexes with high-affinity by stabilizing unique TM6 conformers. Taken together, we hypothesize that ML314, and its successor SBI-55372,73, may bias NTS1 pharmacology by stabilizing signaling incompetent G protein conformations, such as a non-canonical Gα subunit24,25 and/or α5-helix pose74,75. Our observation that βArr1 but not G protein selects a pre-existing enNTS1 conformation near the highly conserved PIF motif is consistent with a previous NMR study33 that utilized a 19F probe at the intracellular tip of TM6 of enNTS1, suggesting that this conformational selection mechanism extends from the PIF motif within the TM domain across the intracellular transducer binding region. The observed peak splitting of M2445.45 and M2505.51 is suggestive of conformational exchange at the ms-s timescale which agrees with the slow-exchange regime established for the TM6 movements of enNTS1 by Dixon et al. However, they observed enNTS1 TM6 adopts a new conformational state upon Gαq-peptide binding (induced-fit) which we can neither confirm nor exclude based on our current data.
Our model system employed several strategies to minimize the challenges to solution NMR studies of GPCR complexes such as multiple simultaneous binding partners, heterogenous phosphorylation patterns, high molecular weights, and inherent instability. We employed the pre-activated βArr-3A variant to control for uncertainty related to the number and position of phosphorylated residues in the receptor C-terminus. To minimize line-broadening side effects of slowly tumbling systems, we employed DDM detergent micelles and focused on only the Gα subunit that comprises nearly the entire complex interface. The Gαiq chimera provides a robust scaffold to explore an otherwise unstable cognate receptor:G protein pair53,54. Thermostabilized enNTS1 permits extended data acquisition times that would otherwise be impossible; it binds ligands with similar affinity to rNTS176, and couples directly to G protein and β-arrestin, although with reduced affinity. Future studies will explore reversion of thermostabilizing mutations to further recover wildtype signaling capabilities. Nonetheless, loss-of-function mutations are more common than gain-of-function phenotypes suggesting that the molecular mechanisms of enNTS1/transducer coupling represent native allosteric pipelines. A long-term aim is to couple quantitative sidechain motions with all atom molecular dynamics to map allosteric connection pathways. Accurate quantitation requires highly deuterated systems28,77 that can be achieved by elegant means78,79,80, but are most easily afforded by E. coli expression systems.
Methods
E. coli expression and purification of enNTS1ΔM4
13CεH3-methionine labelled enNTS1ΔM4 (M2044.60/M2084.64/M2445.45/M2505.51/M3306.57/M3527.36) was expressed as a MBP-enNTS1ΔM4-muGFP fusion protein using the following protocol. 5 mL of a LB day-pre-culture containing 100 mg/L ampicillin and 1% (w/v) glucose were inoculated with a single colony of E. coli OverExpress C43(DE3) cells (Sigma-Aldrich) freshly transformed with pDS170-enNTS1. After 9 h (37 °C, 225 rpm) this pre-culture was centrifuged (1700×g, at RT, 5 min) and the pellet used to inoculate 250 mL of a defined medium pre-culture and grown overnight (1 L flask, 37 °C, 225 rpm). The defined medium consisted of an autoclaved basal salts solution (30 mM KH2PO4, 23 mM K2HPO4, 16 mM Na2HPO4, 17 mM NaCl, 37 mM NH4Cl, adjusted to pH 7.4 with NaOH) supplemented with sterile filtered trace metal stock solution (1% v/v)81, 2 mM MgSO4, 0.4% w/v glucose, 50 mg/L thiamine and 100 mg/L ampicillin. After 13 h the defined medium pre-culture reached an OD600 of ~2, was centrifuged and its pellet resuspended into 250 mL of fresh medium of which 20 mL was used to inoculate 1 L of the same defined medium per 2.8 L flask. Expression cultures were grown for 6.5 h (37 °C, 225 rpm) to an OD600 of 0.4. The flasks were then cooled on ice for 2 min at which point 50 mg/L 13CH3-methionine (Cambridge Stable Isotopes) was added along with 100 mg/L each of lysine, threonine, phenylalanine, and 50 mg/L each of leucine, isoleucine and valine. The expression cultures were placed into a 16 °C incubator for 15 min, then induced with 250 mM isopropyl β-D-1-thiogalactopyranoside (IPTG) and protein expression was carried out at 16 °C and 225 rpm for 12–16 h. For unlabeled expression of MBP-enNTS1ΔM4-muGFP, 5 mL of LB pre-culture was used to inoculate 500 mL 2YT medium containing 100 mg/L ampicillin and 0.2% (w/v) glucose in 2 L flasks. The flasks were incubated at 37 °C and 225 rpm until an OD600 of 0.4 was reached. The flasks were cooled on ice for 2 min, then induced with 250 μM IPTG. Protein expression was carried out at 16 °C and 225 rpm for 12–16 h. All expression cultures were harvested by centrifugation (5000×g, 4 °C, 15 min) and the combined pellets resuspended with 50 mL of wash buffer (25 mM HEPES, 100 mM NaCl, pH 8) per 1 L of expression culture and centrifuged again (3000×g, 4 °C, 15 min). The cell pellets were kept frozen at −80 °C until further use.
Cell pellets equivalent to 3–4 L culture were thawed on ice, resuspended in 50 mL of 100 mM HEPES, 400 mM NaCl, 20% glycerol, pH 8 with an EDTA free protease inhibitor tablet (Roche), 100 mg lysozyme and 10 mg DNAse. After rocking for 30 min at 4 °C the cells were sonicated on ice; mixed with 15 mL of DM solution (1.6 g n-decyl-β-D-maltopyranoside, Anatrace dissolved in 15 mL MilliQ water) and 15 mL of CHS/CHAPS solution (0.018 g cholesterol hemi succinate (CHS, Sigma) and 0.09 g 3-((3-cholamidopropyl) dimethylammonio)−1-propanesulfonate hydrate (CHAPS-hydrate, Sigma) dissolved in 15 mL MilliQ water). The solubilization mix was rocked for 2 h at 4 °C; centrifuged (12,000×g, 4 °C, 30 min) and the supernatant was filtered using a 45 μm syringe filter (Millipore), adjusted to 10 mM imidazole and mixed with 3 mL of Talon resin equilibrated in 25 mM HEPES, 300 mM NaCl, 10% glycerol, 0.15% DM, pH 8 and rocked for 1.5 h at 4 °C. The resin retaining the receptor was washed twice with 25 mL of 25 mM HEPES, 500 mM NaCl, 10% glycerol, 0.15% DM, 10 mM imidazole, 0.2 mM PMSF (phenylmethylsulfonyl fluoride), 8 mM ATP, 10 mM MgCl2, pH 8. Detergent exchange to DDM (n-dodecyl-β-D-maltopyranoside, Anagrade, Anatrace) was initiated by washing the resin twice with 25 mL of 25 mM HEPES, 100 mM NaCl, 10% Glycerol, 0.05% DDM, 0.2 mM PMSF, pH 8. The fusion protein was eluted with 15 mL of 25 mM HEPES, 100 mM NaCl, 10% glycerol, 0.05% DDM, 350 mM imidazole, 0.2 mM PMSF, pH 8. IMAC elutions were cleaved with HRV 3 C protease (produced in-house) prior to concentrating using an Amicon 30 kDa MWCO concentrator (Millipore) and dilution with ion exchange chromatography (IEX) loading buffer (20 mM HEPES pH 8.0, 10% Glycerol, 0.02% DDM) to obtain a combined NaCl/Imidazole/Na2SO4 concentration of less than 50 mM. The cleaved receptor solution was then loaded onto a 5 mL HiTrap SP HP column (GE Healthcare) using an Akta Start system (GE Healthcare) and washed with the same buffer until the signal remained stable. The column was then washed with four column volumes of IEX wash buffer (20 mM HEPES pH 7.4, 10% Glycerol, 63 mM NaCl, 0.02% DDM) after which a 1 mL Ni-NTA HisTrap column (GE Healthcare) was inserted after the HiTrap SP HP column and the system was washed with another 10 mL of IEX wash buffer containing 10 mM Imidazole. The cleaved receptor was then eluted with IEX elution buffer (20 mM HEPES pH 7.4, 10% Glycerol, 1 M NaCl, 0.03% DDM, 20 mM Imidazole) and the receptor containing fractions concentrated to ~400 μL for injection onto a S200 Increase SEC column (GE Healthcare) using a 500 μL loop and an Akta Pure System (GE Healthcare). The receptor containing fractions from SEC purification using SEC buffer (50 mM Potassium phosphate pH 7.4, 100 mM NaCl, 0.02% DDM) were then concentrated and buffer exchanged (for NMR experiments) using NMR buffer (50 mM Potassium phosphate pH 7.4, 100 mM NaCl in 100% D2O) to reduce the residual H2O concentration to <1%. Receptor samples were then aliquoted and stored at −80 °C until further use. enNTS1ΔM4 used in NMR experiments retains a C-terminal Avi-tag (which was used previously for capture in ligand-binding and thermostability assays) and the amino acid sequence is:
GPGSTSESDTAGPNSDLDVNTDIYSKVLVTAIYLALFVVGTVGNGVTLFTLARKKSLQSLQSRVDYYLGSLALSSLLILLFALPVDVYNFIWVHHPWAFGDAGCKGYYFLREACTYATALNVVSLSVERYLAICHPFKAKTLLSRSRTKKFISAIWLASALLSLPMLFTMGLQNLSGDGTHPGGLVCTPIVDTATLRVVIQLNTFMSFLFPMLVASILNTVIARRLTVLVHQAAEQARVSTVGTHNGLEHSTFNVTIEPGRVQALRRGVLVLRAVVIAFVVCWLPYHVRRLMFVYISDEQWTTALFDFYHYFYMLSNALVYVSAAINPILYNLVSANFRQVFLSTLASLSPGWRHRRKKRPTFSRKPNSVSSNHAFSTASGLNDIFEAQKIEWHEGSGLEVLFQ
Expression and purification of βArr1-3A
The pET15 expression plasmid harboring the cysteine-free hβArr1 gene (where 6 cysteine residues were mutated to other amino acid types: C59V, C125S, C140L, C242V, C251V and C269S) was a kind gift from Ashish Manglik. In this plasmid the hβArr1-3A sequence was modified by mutating I386A, V387A, F388A (termed 3A mutant)82 via site-directed mutagenesis (using the forward and reverse primer 5′-GAG TTA GAT ACT AAT GAT GAT GAT GCG GCC GCG GAG GAC TTT GCA CGC CAG CG-3′ and 5′-CGC TGG CGT GCA AAG TCC TCC GCG GCC GCA TCA TCA TCA TTA GTA TCT AAC TC-3′, respectively). hβArr1-3A gene was preceded by a 6x His tag, HRV 3 C protease cleavage site and Protein C tag. This sequence was modified by inserting an additional HRV 3C protease cleavage site between the Protein C sequence and the hβArr1-3A gene to allow complete removal of N-terminal tags. Insertion of the additional HRV 3C protease cleavage site was done by site directed mutagenesis (using the forward and reverse primer 5′-CCT GAT TGA TGG CAA AGG CGG AAG TGG AGG ACT GGA AGT GCT GTT CCA GGG CCC GTC GGG CGA CAA GGG AAC ACG TGT CTT CAA G-3′ and 5′-CTT GAA GAC ACG TGT TCC CTT GTC GCC CGA CGG GCC CTG GAA CAG CAC TTC CAG TCC TCC ACT TCC GCC TTT GCC ATC AAT CAG G-3′, respectively). 5 mL of a LB day pre-culture containing 100 mg/L carbenicillin and 1% (w/v) glucose were inoculated with a single colony of E. coli BL21(DE3) cells (Lucigen, Middleton, WI) freshly transformed with the βArr1-3A expression plasmid. After 9 h (37 °C, 225 rpm) 10 μL of LB pre-culture were added to 50 mL of a Terrific Broth (TB) pre-culture containing 100 mg/L carbenicillin and 1% (w/v) glucose, and incubated overnight at (30 °C, 225 rpm). The next morning the 50 mL of TB pre-culture were added to shaker flasks containing 950 mL of the same medium and incubated (37 °C, 225 rpm) to reach an OD600 of 0.6 at which point the temperature was reduced to 20 °C and the culture flasks were incubated further until an OD600 of 1.0 was reached. The flasks were then cooled on ice for 5 min prior to induction with 0.4 mM IPTG. Protein expression was carried out at 20 °C and 225 rpm for 19 h. The cells were harvested by centrifugation (5000 × g, 4 °C, 15 min) and the combined pellets resuspended with wash buffer (25 mM HEPES, 100 mM NaCl, pH 8) and the washed cells were then pelleted by centrifugation (3000×g, 4 °C, 15 min) and stored at −80 °C. Thawed cells were resuspended in solubilization buffer (20 mM HEPES pH 8, 500 mM NaCl, 2 mM MgCl2, 15% glycerol, 1 Roche EDTA free Protease inhibitor tablet, 0.4 mM PMSF, 1 mg/mL Lysozyme, 1 μL/mL DNAse) and left stirring at 4 °C for 30 min prior to sonication on ice. Cell debris was removed by centrifugation (24,000×g, 4 °C, 45 min) and the supernatant filtered using a 45 μm syringe filter (Millipore). The filtrate was then incubated for 1 h rotating at 4 °C with 2 mL Ni-NTA resin (Thermo Fisher) per 1.5 L of expression culture. The resin was then washed with 15 mL wash buffer 1 (20 mM HEPES pH 8, 300 mM NaCl, 10% glycerol, 10 mM imidazole) followed by 12 mL wash buffer 2 (same as wash buffer 1 but containing 20 mM imidazole) and 12 mL wash buffer 3 (same as wash buffer 1 but containing 25 mM imidazole) per 1 mL of resin. βArr1-3A was eluted with ~10 mL of elution buffer (20 mM HEPES pH 7.5, 150 mM NaCl, 10% glycerol, 200 mM imidazole) per 1 mL of resin. The 6× His-tag was removed via His-tagged HRV 3 C protease (produced in-house) cleavage overnight rotating at 4 °C. The cleavage reaction was then concentrated using an Amicon 30 kDa MWCO centrifugal concentrator (Millipore) and diluted with 20 mM HEPES pH 7.5 to obtain a combined NaCl/Imidazole/Na2SO4 concentration of less than 50 mM. The solution was again filtered using a 45 μm syringe filter (Millipore) prior to loading onto a 5 mL HiTrap Q IEX column (GE Healthcare) equilibrated with 20 mM HEPES pH 7.5 followed by a wash step with the same buffer until a conductivity of 5 mS/cm was reached. The column was then further washed with IEX wash buffer (20 mM HEPES pH 7.5, 50 mM NaCl) until the A280 signal stabilized. The column was eluted using a 25 min gradient stretching from 50 mM to 500 mM NaCl. The βArr1-3A containing fractions were then pooled and concentrated to ~750 μL using an Amicon 30 kDa MWCO centrifugal concentrator (Millipore) prior to injection onto a HiLoad 16/600 S200pg SEC column (GE Healthcare) equilibrated with SEC buffer (20 mM HEPES pH 6.8, 150 mM NaCl) using a 1 mL loop. The βArr1-3A containing SEC fractions were pooled, concentrated to 296 μM and aliquots stored at −80˚C until further use. The amino acid sequence of βArr1-3A used in NMR experiments is:
GPSGDKGTRVFKKASPNGKLTVYLGKRDFVDHIDLVDPVDGVVLVDPEYLKERRVYVTLTVAFRYGREDLDVLGLTFRKDLFVANVQSFPPAPEDKKPLTRLQERLIKKLGEHAYPFTFEIPPNLPSSVTLQPGPEDTGKALGVDYEVKAFVAENLEEKIHKRNSVRLVIRKVQYAPERPGPQPTAETTRQFLMSDKPLHLEASLDKEIYYHGEPISVNVHVTNNTNKTVKKIKISVRQYADIVLFNTAQYKVPVAMEEADDTVAPSSTFSKVYTLTPFLANNREKRGLALDGKLKHEDTNLASSTLLREGANREILGIIVSYKVKVKLVVSRGGLLGDLASSDVAVELPFTLMHPKPKEEPPHREVPENETPVDTNLIELDTNDDDAAAEDFARQRLKGMKDDKEEEEDGTGSPQLNNR
[U-15N,13C,2H]-βArr1-3A was produced using the above construct with a modified expression protocol. 5 mL of a LB day pre-culture containing 100 mg/L carbenicillin and 1% (w/v) glucose were inoculated with a single colony of E. coli BL21(DE3) cells (Lucigen, Middleton, WI) freshly transformed with the βArr1-3A expression plasmid. After 9 h (37 °C, 225 rpm), 250 μL of the LB pre-culture was centrifuged at 5000×g, resuspended in 5 mL LB/D2O containing 100 mg/L carbenicillin and 1% (w/v) glucose, and incubated overnight at (30 °C, 225 rpm). The next morning, 250 μL of the LB/D2O pre-culture was centrifuged at 5000×g, resuspended in 5 mL M9/D2O, and incubated (37 °C, 225 rpm). 1 L M9/D2O was prepared from: 1 L (99.9% 2H) D2O, 13.6 g Na2HPO4, 6 g KH2PO4, 1 g NaCl, 0.07 g CaCl2, 0.5 g MgSO4, 0.05 g carbenicillin, 1.5 g 15N4Cl, 3 g d7,13C-glucose, trace metal stock, and vitamin stock. After 9 h, the entire 5 mL M9/D2O pre-culture was added to 90 mL of a M9/D2O pre-culture and incubated overnight (30 °C, 225 rpm). The next morning, the 100 mL M9/D2O pre-culture was added to shaker flasks containing 900 mL of the same medium and incubated (37 °C, 225 rpm) to reach an OD600 of 0.8 at which point the temperature was reduced to 20 °C and the culture was incubated further until an OD600 of 1.0 was reached. The flasks were then cooled on ice for 5 min prior to induction with 0.4 mM IPTG. Protein expression was carried out at 20 °C and 225 rpm for 19 h. The cells were harvested by centrifugation (5000×g, 4 °C, 15 min) and the combined pellets resuspended with wash buffer (25 mM HEPES, 100 mM NaCl, pH 8) and the washed cells were then pelleted by centrifugation (3000×g, 4 °C, 15 min) and stored at −80 °C. Purification followed the same strategy as outlined above for unlabeled βArr1-3A.
For MST experiments, unlabeled βArr1-3A was expressed and purified as above, but the N-terminal 6xHis tag was left intact yielding the amino acid sequence:
MGSSHHHHHHLEVLFQGPGGEDQVDPRLIDGKGGSGGLEVLFQGPSGDKGTRVFKKASPNGKLTVYLGKRDFVDHIDLVDPVDGVVLVDPEYLKERRVYVTLTVAFRYGREDLDVLGLTFRKDLFVANVQSFPPAPEDKKPLTRLQERLIKKLGEHAYPFTFEIPPNLPSSVTLQPGPEDTGKALGVDYEVKAFVAENLEEKIHKRNSVRLVIRKVQYAPERPGPQPTAETTRQFLMSDKPLHLEASLDKEIYYHGEPISVNVHVTNNTNKTVKKIKISVRQYADIVLFNTAQYKVPVAMEEADDTVAPSSTFSKVYTLTPFLANNREKRGLALDGKLKHEDTNLASSTLLREGANREILGIIVSYKVKVKLVVSRGGLLGDLASSDVAVELPFTLMHPKPKEEPPHREVPENETPVDTNLIELDTNDDDAAAEDFARQRLKGMKDDKEEEEDGTGSPQLNNR
Expression and purification of Gαiq
The codon optimized gene for the Gαiq chimera52 was purchased from GenScript (Piscataway, NJ) in a puc57 vector and subcloned into a pIQ expression vector with an open reading frame encoding an N-terminal 6xHis tag followed by a NNNNNNNNNNG linker, a MBP sequence and a HRV 3 C protease cleavage site (LEVLFQGP). The Gαiq gene was amplified via PCR using the forward and reverse primer 5′-CAT CAT GGA TCC GGT TGC ACC CTG TCT GCG GAA GAC-3′ and 5′-CAG CTA TGA CCA TGA TTA CGC-3′, respectively. Subcloning was performed via the BamHI and HindIII restriction sites. 50 mL of a LB day pre-culture containing 100 mg/L carbenicillin and 1% (w/v) glucose were inoculated with a single colony of E. coli BL21(DE3) cells (Lucigen, Middleton, WI) freshly transformed with the Gαiq expression plasmid. After 9 h (37 °C, 225 rpm) 20 mL of LB pre-culture were centrifuged (3000×g, RT, 5 min) and the resuspended pellet was used to inoculate 1 L of 2xYT medium containing 100 mg/L carbenicillin and 0.2% (w/v) glucose. The cultures were incubated (37 °C, 225 rpm) to reach an OD600 of 0.7. The flasks were then cooled on ice for 5 min prior to induction with 1 mM IPTG. Protein expression was carried out at 25 °C and 225 rpm for 16 h. The cells were harvested by centrifugation (5000×g, 4 °C, 15 min) and the combined pellets resuspended with wash buffer (25 mM HEPES, 100 mM NaCl, pH 8) and the washed cells were then pelleted by centrifugation (3000×g, 4 °C, 15 min) and stored at −80 °C. Gαiq was purified following a protocol for purification of miniG proteins83. A cell pellet from 2 L of E. coli culture was thawed on ice and resuspended in buffer A (40 mM HEPES pH 7.5, 100 mM NaCl, 10 mM imidazole, 10% v/v glycerol, 5 mM MgCl2, 50 μM GDP) to a volume of 50 mL. A protease inhibitor tablet (Roche), 1 mM PMSF, 5 U DNAse I, 25 mg lysozyme, and 100 μM DTT were added, and the cell suspension was stirred at medium speed at 4 °C for 30 min. The cells were then lysed by sonication on ice for 7 min, with pulses of 2 s on/4 s off, at an amplitude of 70% and the lysate centrifuged at 20,000×g for 45 min at 4 °C. The supernatant was filtered using a 45 μm syringe filter (Millipore) prior to loading onto a 5 mL HisTrap Fast Flow column (GE Healthcare), pre-equilibrated with ice-cold buffer A. The column was washed with 10 column volumes of ice-cold buffer B (20 mM HEPES pH 7.5, 500 mM NaCl, 40 mM imidazole, 10% v/v glycerol, 1 mM MgCl2, 50 μM GDP) and eluted with ~5 column volumes of ice-cold buffer C (20 mM HEPES pH 7.5, 100 mM NaCl, 500 mM imidazole, 10% v/v glycerol, 1 mM MgCl2, 50 μM GDP). The elute was complemented with 1 mM DTT, 100 mM Na2SO4, and His-tagged HRV 3 C protease (produced in-house) at a 3 C:Gαiq ratio of 1:20 w/w, transferred to a 10 kDa cut-off dialysis tube (Thermo Fisher), and dialyzed overnight at 4 °C against 1 L of buffer D (20 mM HEPES pH 7.5, 100 mM NaCl, 10% v/v glycerol, 1 mM MgCl2, 10 μM GDP). The next day imidazole was added to a concentration of 20 mM and the solution was incubated with 4 mL of Ni-NTA resin (Thermo Fisher) pre-equilibrated with buffer D on a turning wheel for 30 min at 4 °C. The suspension was then poured onto 1 mL of fresh Ni-NTA resin (pre-equilibrated with buffer D) in a disposable plastic column and the flow-through was eluted by gravity. The column was washed with 5 mL of buffer D. The wash was combined with the flow-through and concentrated to ~1.6 mL using an Amicon 10 kDa MWCO centrifugal concentrator (Millipore) prior to loading onto a HiLoad 16/600 S200pg SEC column (GE Healthcare) pre-equilibrated with buffer E (10 mM HEPES pH 7.5, 100 mM NaCl, 10% v/v glycerol, 1 mM MgCl2, 1 μM GDP, 100 μM TCEP). The Gαiq peak fractions were pooled and concentrated to ~0.5 mL followed by centrifugation at 16000×g for 5 min to remove aggregates. The solution containing 721 μM Gαiq at a purity of >95% (as determined by SDS-Page) was flash-frozen in aliquots and stored at stored at −80 °C until further use. The amino acid sequence of Gαiq used in NMR experiments is:
GPGSGCTLSAEDKAAVERSKMIDRNLREDGEKAAREVKLLLLGAGESGKSTIVKQMKIIHEAGYSEEECKQYKAVVYSNTIQSIIAIIRAMGRLKIDFGDSARADDARQLFVLAGAAEEGFMTAELAGVIKRLWKDSGVQACFNRSREYQLNDSAAYYLNDLDRIAQPNYIPTQQDVLRTRVKTTGIVETHFTFKDLHFKMFDVGGQRSERKKWIHCFEGVTAIIFCVALSDYDLVLAEDEEMNRMHESMKLFDSICNNKWFTDTSIILFLNKKDLFEEKIKKSPLTICYPEYAGSNTYEEAAAYIQCQFEDLNKRKDTKEIYTHFTCATDTKNVQFVFDAVTDVIIKNNLKEYNLV
[U-15N,13C,2H]-Gαiq was produced using the above construct with a modified expression protocol. 5 mL of a LB day pre-culture containing 100 mg/L carbenicillin and 1% (w/v) glucose were inoculated with a single colony of E. coli BL21(DE3) cells (Lucigen, Middleton, WI) freshly transformed with the Gαiq expression plasmid. After 9 h (37 °C, 225 rpm), 250 μL of the LB pre-culture was centrifuged at 5000×g, resuspended in 5 mL LB/D2O containing 100 mg/L carbenicillin and 1% (w/v) glucose, and incubated overnight at (30 °C, 225 rpm). The next morning, 250 μL of the LB/D2O pre-culture was centrifuged at 5000×g, resuspended in 5 mL M9/D2O, and incubated (37 °C, 225 rpm). After 9 h, the entire 5 mL M9/D2O pre-culture was added to 90 mL of a M9/D2O pre-culture and incubated overnight (30 °C, 225 rpm). The next morning, the 100 mL M9/D2O pre-culture was added to shaker flasks containing 900 mL of the same medium and incubated (37 °C, 225 rpm) to reach an OD600 of 0.8 at which point the temperature was reduced to 25 °C and the culture was incubated further until an OD600 of 1.0 was reached. The flasks were then cooled on ice for 5 min prior to induction with 1 mM IPTG. Protein expression was carried out at 25 °C and 225 rpm for 20 h. The cells were harvested by centrifugation (5000×g, 4 °C, 15 min) and stored at −80 °C. Purification followed the same strategy as outlined above for unlabeled Gαiq.
A second Gαiq construct was cloned for MST experiments from the above template. The MBP and 3 C cleavage site were removed via restriction free cloning (using the forward and reverse primers 5′-GAG AGG ATC GCA TCA CCA TCA CCA TCA CGG ATC TGG ATC CGG TTG CAC CCT GTC TGC GGA AGA C-3′ and 5′-GTC TTC CGC AGA CAG GGT GCA ACC GGA TCC AGA TCC GTG ATG GTG ATG GTG ATG CGA TCC TCT C-3′, respectively) to place the 6xHis tag in closer proximity to the protein core. This construct was purified as above except for the 3C cleavage step to yield the amino acid sequence:
MRGSHHHHHHGSGSGCTLSAEDKAAVERSKMIDRNLREDGEKAAREVKLLLLGAGESGKSTIVKQMKIIHEAGYSEEECKQYKAVVYSNTIQSIIAIIRAMGRLKIDFGDSARADDARQLFVLAGAAEEGFMTAELAGVIKRLWKDSGVQACFNRSREYQLNDSAAYYLNDLDRIAQPNYIPTQQDVLRTRVKTTGIVETHFTFKDLHFKMFDVGGQRSERKKWIHCFEGVTAIIFCVALSDYDLVLAEDEEMNRMHESMKLFDSICNNKWFTDTSIILFLNKKDLFEEKIKKSPLTICYPEYAGSNTYEEAAAYIQCQFEDLNKRKDTKEIYTHFTCATDTKNVQFVFDAVTDVIIKNNLKEYNLV
Microscale thermophoresis
βArr1-3A and Gαiq chimera constructs containing N-terminal 6x His-tags were labeled with the His-Tag Labeling Kit RED-tris-NTA 2nd Generation (NanoTemper). A constant concentration of either RED-βArr1-3A or RED-Gαiq chimera was incubated with varying concentrations of NT8-13-bound enNTS1ΔM4 before being loaded into NanoTemper premium capillaries and assayed via a NanoTemper Monolith NT MST instrument. βArr1-3A experiments consisted of 16 samples in 50 mM Potassium phosphate pH 7.4, 100 mM NaCl containing 25 nM RED-βArr1-3A, 40 µM NT8-13, 40 µM C8-PIP2, 0.03% DDM and a 1:2 dilution series of enNTS1ΔM4 from 20 μM to 0.61 nM. Gαiq chimera experiments consisted of 16 samples in 50 mM Potassium phosphate pH 7.4, 100 mM NaCl containing 25 nM RED-Gαiq chimera, 100 µM NT8-13, 100 µM C8-PIP2, 0.03% DDM, 2 mM MgCl2, 10 μM GDP, 0.2 units of apyrase and a 1:2 dilution series of enNTS1ΔM4 from 50 μM to 1.53 nM. Data was collected using 80% LED and 80% MST power in experimental triplicate and at least three technical repeats. Raw data was fit to a Hill function to establish upper and lower asymptote values. These values were then used to normalize the data and subsequently fit to a quadratic binding function using Sigma Plot v15 (Systat Software Inc.).
NMR spectroscopy
NMR spectra were collected on 600 MHz Bruker Avance Neo spectrometers equipped with triple resonance cryoprobes and operated with Topspin v3.6.2. 2D 1H-13C SOFAST-HMQC spectra65 were recorded with 25% non-uniform sampling (NUS) at 298 K with a 1H spectral width of 12 ppm (1024 data points in t2) and a 13C spectral width of 25 ppm (128 data points in t1), relaxation delays of 450 ms, and 2048 scans per t1 data point resulting in acquisition times of 10 h per spectrum. A 2.25 ms PC9 120 degree 1H pulse84 was applied for excitation and a 1 ms r-SNOB shaped 180 degree 1H pulse85 was used for refocusing. The 13C carrier frequency was positioned at 17 ppm, and the 1H at 4.7 ppm, while band selective 1H pulses were centered at 1.8 ppm. 1D 1H spectra were recorded at 298 K with a spectral width of 13.7 ppm (2048 data points), a relaxation delay of 1 s, and 128 scans. 15N-TROSY-HSQC spectra were recorded at 298 K with 50% NUS using a Poisson gap schedule86, a 1H spectral width of 12 ppm (2048 data points in t2), a 15N spectral width of 40 ppm (128 data points in t1), relaxation delays of 1 s, and 128 scans per t1 data point resulting in acquisition times of ~9 h per spectrum.
The [U-15N,13C,2H]-βArr1-3A sample was prepared to a volume of 160 μL NMR buffer in 3 mm tubes (Wilmad), containing 20 μM DSS and 0.05% Na2N3. NT8-13 was added to a final concentration of 500 μM. PIP2 was added to a final concentration of 130 μM (~1.5x molar equivalents of the final receptor concentration). Unlabeled enNTS1ΔM4 was buffer exchanged three times with NMR buffer (to >99%) prior to combining with the transducer. Following each addition of unlabeled enNTS1ΔM4, the samples were incubated for 1 h at room temperature prior to starting experiments.
The [U-15N,13C,2H]-Gαiq sample was prepared to a volume of 160 μL NMR buffer in 3 mm tubes (Wilmad) containing 20 μM DSS, 0.05% Na2N3, 2 mM MgCl2, 100 μM TCEP and 0.25 units apyrase. NT8-13 was added to a final concentration of 500 μM. PIP2 was added to a final concentration of 130 μM (~1.5x molar equivalents of final receptor concentration). Unlabeled enNTS1ΔM4 was buffer exchanged three times with NMR buffer (to >99%) prior to combining with the transducer. Following each addition of unlabeled enNTS1ΔM4, the samples were incubated for 1 h at room temperature prior to starting experiments.
[13CεH3-methionine]-enNTS1ΔM4 samples were prepared to volumes of 160 μL in 3 mm tubes (Wilmad), containing 20 μM DSS and 0.05% Na2N3. Ligands were added to a final concentration of 500 μM. NT8-13 (5–10 mM) stock solutions were prepared in 100% D2O and ML314 (20 mM) in 100% DMSO-d6. PIP2 was added to a final concentration of 130 μM (~2× molar equivalents of receptor). βArr1-3A and Gαiq aliquots of 0.3–3x molar equivalents of receptor were buffer exchanged three times with NMR buffer (to >99%) prior to combining with the receptor. βArr1-3A containing samples were incubated for 1 h at room temperature prior to starting experiments. Gαiq containing samples were supplemented with 2 mM MgCl2, 100 μM TCEP, and 10 μM GDP and incubated for 1 h at room temperature prior to adding 0.25 units of Apyrase (NEB) and incubation for a further 1 h at room temperature.
All [13CεH3-methionine]-enNTS1ΔM4 spectra were referenced against internal DSS, reconstructed with compressed sensing using qMDD87, and processed using NMRPipe88 where data were multiplied by cosinebells and zero-filled once in each dimension. All 2D 1H-13C SOFAST-HMQC spectra used in this study are reproduced together in Fig. S8.
Both [U-15N,13C,2H]-βArr1-3A and [U-15N,13C,2H]-Gαiq spectra were referenced against internal DSS, reconstructed with iterative soft thresholding (IST)86, and processed using NMRPipe88 where data were multiplied by cosinebells and zero-filled once in each dimension.
All spectra were analyzed in NMRFAM Sparky v1.47 (Goddard, T.D. and Kneller, D.G., University of California, San Francisco). Peak integrals were normalized and plotted using GraphPad Prism v9.5.1 (GraphPad Software).
13C-SOFAST-HMQC spectra of Apo-state, NT8-13-, ML314-, and NT8-13 & ML314-bound enNTS1ΔM4 without PIP2 were generated in a previous study22.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data used for graphs are provided with this paper. The chemical shift assignments of 13C-SOFAST-HMQC spectra generated in this study have been deposited in the Biological Magnetic Resonance Bank (BMRB) under accession codes 51908 (PIP2:Apo-state-enNTS1ΔM4), 51909 (PIP2:NT8-13:enNTS1ΔM), 51910 (PIP2:ML314:enNTS1ΔM4), 51911 (PIP2:NT8-13:ML314:enNTS1ΔM4), 51914 (NT8-13:enNTS1ΔM4:βArr1-3A), 51915 (PIP2:NT8-13:enNTS1ΔM4:βArr1-3A), 51916 (PIP2:ML314:enNTS1ΔM4:βArr1-3A), 51917 (PIP2:NT8-13:ML314:enNTS1ΔM4:βArr1-3A), 51921 (PIP2:NT8-13:enNTS1ΔM4:Gαiq), and 51927 (PIP2:NT8-13:ML314:enNTS1ΔM4:Gαiq). 13C-SOFAST-HMQC spectra of Apo-state, NT8-13-, ML314-, and NT8-13 & ML314-bound enNTS1ΔM4 were generated in a previous study22 and the chemical shift assignments were deposited in the BMRB under accession codes 51728 (Apo-state enNTS1ΔM4), 51735 (NT8-13:enNTS1ΔM), 51737 (ML314:enNTS1ΔM4), 51738 (NT8-13:ML314:enNTS1ΔM4). The NMR spectra generated during and/or analyzed during the current study are available from the corresponding author on reasonable request. PDB files referenced in this manuscript are available from the RCSB Protein Data Bank: 4BWB (HTGH4-ΔIC3:NT8-13), 6YVR (NTSR1-H4X:SR142948A, and 6Z4Q (NTSR1-H4X:NT8-13). Source data are provided with this paper.
References
Hauser, A. S., Attwood, M. M., Rask-Andersen, M., Schioth, H. B. & Gloriam, D. E. Trends in GPCR drug discovery: new agents, targets and indications. Nat. Rev. Drug Discov. 16, 829–842 (2017).
Katritch, V., Cherezov, V. & Stevens, R. C. Diversity and modularity of G protein-coupled receptor structures. Trends Pharmacol. Sci. 33, 17–27 (2012).
Lappano, R. & Maggiolini, M. G protein-coupled receptors: novel targets for drug discovery in cancer. Nat. Rev. Drug Discov. 10, 47–60 (2011).
Pierce, K. L., Premont, R. T. & Lefkowitz, R. J. Seven-transmembrane receptors. Nat. Rev. Mol. Cell Biol. 3, 639–650 (2002).
Weis, W. I. & Kobilka, B. K. The molecular basis of G protein–coupled receptor activation. Annu. Rev. Biochem. 87, 897–919 (2018).
Vincent, J. P., Mazella, J. & Kitabgi, P. Neurotensin and neurotensin receptors. Trends Pharmacol. Sci. 20, 302–309 (1999).
Tyler-McMahon, B. M., Boules, M. & Richelson, E. Neurotensin: peptide for the next millennium. Regul. Pept. 93, 125–136 (2000).
Binder, E. B., Kinkead, B., Owens, M. J., Kilts, C. D. & Nemeroff, C. B. Enhanced neurotensin neurotransmission is involved in the clinically relevant behavioral effects of antipsychotic drugs: evidence from animal models of sensorimotor gating. J. Neurosci. 21, 601–608 (2001).
Ferraro, L. et al. Neurotensin: a role in substance use disorder? J. Psychopharmacol. 30, 112–127 (2016).
Kempadoo, K. A. et al. Hypothalamic neurotensin projections promote reward by enhancing glutamate transmission in the VTA. J. Neurosci. 33, 7618–7626 (2013).
Opland, D. et al. Loss of neurotensin receptor-1 disrupts the control of the mesolimbic dopamine system by leptin and promotes hedonic feeding and obesity. Mol. Metab. 2, 423–434 (2013).
Woodworth, H. L. et al. Neurotensin receptor-1 Identifies a subset of ventral tegmental dopamine neurons that coordinates energy balance. Cell Rep. 20, 1881–1892 (2017).
Bissette, G., Nemeroff, C. B., Loosen, P. T., Prange, A. J. Jr. & Lipton, M. A. Hypothermia and intolerance to cold induced by intracisternal administration of the hypothalamic peptide neurotensin. Nature 262, 607–609 (1976).
Kitabgi, P. et al. Functional and pharmacological aspects of central neuropeptidergic transmission mediated by neurotensin and neuromedin n. Clin. Neuropharmacol. 15, 313A–314A (1992).
Carraway, R. & Leeman, S. E. The isolation of a new hypotensive peptide, neurotensin, from bovine hypothalami. J. Biol. Chem. 248, 6854–6861 (1973).
Osbahr, A. J. 3rd, Nemeroff, C. B., Manberg, P. J. & Prange, A. J. Jr. Centrally administered neurotensin: activity in the Julou-Courvoisier muscle relaxation test in mice. Eur. J. Pharmacol. 54, 299–302 (1979).
Costa-Neto, C. M., Parreiras, E. S. L. T. & Bouvier, M. A pluridimensional view of biased agonism. Mol. Pharmacol. 90, 587–595 (2016).
Masuho, I. et al. Distinct profiles of functional discrimination among G proteins determine the actions of G protein-coupled receptors. Sci. Signal. 8, ra123 (2015).
Pupo, A. S. et al. Recent updates on GPCR biased agonism. Pharmacol. Res. 112, 49–57 (2016).
Barak, L. S. et al. ML314: a biased neurotensin receptor ligand for methamphetamine abuse. ACS Chem. Biol. 11, 1880–1890 (2016).
Peddibhotla, S. et al. Discovery of ML314, a brain penetrant nonpeptidic beta-arrestin biased agonist of the neurotensin NTR1 receptor. ACS Med. Chem. Lett. 4, 846–851 (2013).
Bumbak, F. et al. Ligands selectively tune the local and global motions of neurotensin receptor 1 (NTS1). Cell Rep. 42, 112015 (2023).
Huang, W. et al. Structure of the neurotensin receptor 1 in complex with β-arrestin 1. Nature 597, 303–308 (2020).
Kato, H. E. et al. Conformational transitions of a neurotensin receptor 1–Gi1 complex. Nature 572, 80–85 (2019).
Zhang, M. et al. Cryo-EM structure of an activated GPCR-G protein complex in lipid nanodiscs. Nat. Struct. Mol. Biol. 28, 258–267 (2021).
Yin, W. et al. A complex structure of arrestin-2 bound to a G protein-coupled receptor. Cell Res. 29, 971–983 (2019).
Shimada, I., Ueda, T., Kofuku, Y., Eddy, M. T. & Wüthrich, K. GPCR drug discovery: integrating solution NMR data with crystal and cryo-EM structures. Nat. Rev. Drug. Discov. 18, 59–82 (2018).
Clark, L. D. et al. Ligand modulation of sidechain dynamics in a wild-type human GPCR. eLife 6, e28505 (2017).
Huang, S. K. et al. Delineating the conformational landscape of the adenosine A2A receptor during G protein coupling. Cell 184, 1884–1894.e14 (2021).
Rößler, P. et al. GPCR activation states induced by nanobodies and mini-G proteins compared by NMR spectroscopy. Molecules 25, 5984 (2020).
Shiraishi, Y. et al. Biphasic activation of β-arrestin 1 upon interaction with a GPCR revealed by methyl-TROSY NMR. Nat. Commun. 12, 7158 (2021).
Shiraishi, Y. et al. Phosphorylation-induced conformation of β2-adrenoceptor related to arrestin recruitment revealed by NMR. Nat. Commun. 9, 194 (2018).
Dixon, A. D. et al. Effect of ligands and transducers on the neurotensin receptor 1 conformational ensemble. J. Am. Chem. Soc. 144, 10241–10250 (2022).
Xu, J. et al. Structural and dynamic insights into supra-physiological activation and allosteric modulation of a muscarinic acetylcholine receptor. Nat. Commun. 14, 376 (2023).
Manglik, A. et al. Structural Insights into the dynamic process of β2-adrenergic receptor signaling. Cell 21, 1101–1111 (2015).
Nygaard, R. et al. The dynamic process of beta(2)-adrenergic receptor activation. Cell 152, 532–542 (2013).
Damian, M. et al. Allosteric modulation of ghrelin receptor signaling by lipids. Nat. Commun. 12, 3938 (2021).
Yen, H.-Y. et al. PtdIns(4,5)P2 stabilizes active states of GPCRs and enhances selectivity of G-protein coupling. Nature 559, 423–427 (2018).
Bumbak, F. et al. Optimization and 13CH3 methionine labeling of a signaling competent neurotensin receptor 1 variant for NMR studies. Biochim. Biophys. Acta Biomembr. 1860, 1372–1383 (2018).
Scott, D. J. & Pluckthun, A. Direct molecular evolution of detergent-stable g protein-coupled receptors using polymer encapsulated cells. J. Mol. Biol. 425, 662–677 (2013).
Ballesteros, J. A. & Weinstein, H. Methods in Neurosciences, Vol. 25 (ed. Stuart, C. S.) 366−428 (Academic Press, 1995).
Kanba, K. S., Kanba, S., Nelson, A., Okazaki, H. & Richelson, E. [3H]neurotensin(8-13) binds in human brain to the same sites as does [3H]neurotensin but with higher affinity. J. Neurochem. 50, 131–137 (1988).
Abragam, A. The Principles Of Nuclear Magnetism, p. 599 (Clarendon Press, Oxford, 1961).
Kovrigin, E. L. NMR line shapes and multi-state binding equilibria. J. Biomol. NMR 53, 257–270 (2012).
Palmer, A. G. 3rd, Kroenke, C. D. & Loria, J. P. Nuclear magnetic resonance methods for quantifying microsecond-to-millisecond motions in biological macromolecules. Methods Enzymol. 339, 204–238 (2001).
Deluigi, M. et al. Complexes of the neurotensin receptor 1 with small-molecule ligands reveal structural determinants of full, partial, and inverse agonism. Sci. Adv. 7, eabe5504 (2021).
Gurevich, V. V. The selectivity of visual arrestin for light-activated phosphorhodopsin is controlled by multiple nonredundant mechanisms. J. Biol. Chem. 273, 15501–15506 (1998).
Vishnivetskiy, S. A. et al. An additional phosphate-binding element in arrestin molecule. Implications for the mechanism of arrestin activation. J. Biol. Chem. 275, 41049–41057 (2000).
Palczewski, K., Buczylko, J., Imami, N. R., McDowell, J. H. & Hargrave, P. A. Role of the carboxyl-terminal region of arrestin in binding to phosphorylated rhodopsin. J. Biol. Chem. 266, 15334–15339 (1991).
Dror, R. O. et al. Activation mechanism of the beta2-adrenergic receptor. Proc. Natl Acad. Sci. USA 108, 18684–18689 (2011).
Müller, K. M. et al. Role of protein kinase C and epidermal growth factor receptor signalling in growth stimulation by neurotensin in colon carcinoma cells. BMC Cancer 11, 421 (2011).
Grisshammer, R. & Hermans, E. Functional coupling with Galpha(q) and Galpha(i1) protein subunits promotes high-affinity agonist binding to the neurotensin receptor NTS-1 expressed in Escherichia coli. FEBS Lett. 493, 101–105 (2001).
Kumar, A. & Pluckthun, A. In vivo assembly and large-scale purification of a GPCR - Galpha fusion with Gbetagamma, and characterization of the active complex. PLoS ONE 14, e0210131 (2019).
Nehme, R. et al. Mini-G proteins: novel tools for studying GPCRs in their active conformation. PLoS ONE 12, e0175642 (2017).
Rasmussen, S. G. et al. Crystal structure of the beta2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011).
Schoenlein, R. W., Peteanu, L. A., Mathies, R. A. & Shank, C. V. The first step in vision: femtosecond isomerization of rhodopsin. Science 254, 412–415 (1991).
Kofuku, Y. et al. Efficacy of the beta(2)-adrenergic receptor is determined by conformational equilibrium in the transmembrane region. Nat. Commun. 3, 1045 (2012).
Wu, F.-J. et al. Probing the correlation between ligand efficacy and conformational diversity at the α1A-adrenoreceptor reveals allosteric coupling of its microswitches. J. Biol. Chem. 295, 7404–7417 (2020).
Gregorio, G. G. et al. Single-molecule analysis of ligand efficacy in beta2AR-G-protein activation. Nature 547, 68–73 (2017).
Asher, W. B. et al. GPCR-mediated beta-arrestin activation deconvoluted with single-molecule precision. Cell 185, 1661–1675.e16 (2022).
Huang, S. K. & Prosser, R. S. Dynamics and mechanistic underpinnings to pharmacology of class A GPCRs: an NMR perspective. Am. J. Physiol. Cell Physiol. 322, C739–C753 (2022).
Huang, W. et al. Structural insights into micro-opioid receptor activation. Nature 524, 315–321 (2015).
Chashmniam, S., Teixeria, J. M. C., Paniagua, J. C. & Pons, M. A methionine chemical shift based order parameter characterizing global protein dynamics. ChemBioChem 22, 1001–1004 (2021).
Luginbühl, P. & Wüthrich, K. Semi-classical nuclear spin relaxation theory revisited for use with biological macromolecules. Prog. Nucl. Magn. Reson. Spectrosc. 40, 199–247 (2002).
Schanda, P. & Brutscher, B. Very fast two-dimensional NMR spectroscopy for real-time investigation of dynamic events in proteins on the time scale of seconds. J. Am. Chem. Soc. 127, 8014–8015 (2005).
Selvaratnam, R., Chowdhury, S., VanSchouwen, B. & Melacini, G. Mapping allostery through the covariance analysis of NMR chemical shifts. Proc. Natl Acad. Sci. USA 108, 6133–6138 (2011).
Ma, N., Lee, S. & Vaidehi, N. Activation microswitches in adenosine receptor A2A function as rheostats in the cell membrane. Biochemistry 59, 4059–4071 (2020).
Song, W., Yen, H.-Y., Robinson, C. V. & Sansom, M. S. P. State-dependent lipid interactions with the A2a receptor revealed by MD simulations using in vivo-mimetic membranes. Structure 27, 392–403.e3 (2018).
Hart, K. M., Ho, C. M., Dutta, S., Gross, M. L. & Bowman, G. R. Modelling proteins’ hidden conformations to predict antibiotic resistance. Nat. Commun. 7, 12965 (2016).
Pontiggia, F. et al. Free energy landscape of activation in a signalling protein at atomic resolution. Nat. Commun. 6, 7284 (2015).
Löhr, T., Kohlhoff, K., Heller, G. T., Camilloni, C. & Vendruscolo, M. A kinetic ensemble of the Alzheimer’s Aβ peptide. Nat. Comput. Sci. 1, 71–78 (2021).
Pinkerton, A. B. et al. Discovery of beta-arrestin biased, orally bioavailable, and CNS penetrant neurotensin receptor 1 (NTR1) allosteric modulators. J. Med. Chem. 62, 8357–8363 (2019).
Slosky, L. M. et al. beta-arrestin-biased allosteric modulator of NTSR1 selectively attenuates addictive behaviors. Cell 181, 1364–1379.e14 (2020).
Kim, K. et al. beta2-adrenoceptor ligand efficacy is tuned by a two-stage interaction with the Galphas C terminus. Proc. Natl Acad. Sci. USA 118, e2017201118 (2021).
Sandhu, M. et al. Conformational plasticity of the intracellular cavity of GPCR-G-protein complexes leads to G-protein promiscuity and selectivity. Proc. Natl Acad. Sci. USA 116, 11956–11965 (2019).
Bumbak, F., Bathgate, R. A. D., Scott, D. J. & Gooley, P. R. Expression and purification of a functional E. coli 13CH3-methionine-labeled thermostable neurotensin receptor 1 variant for solution NMR studies. Methods Mol. Biol. 1947, 31–55 (2019).
Sun, H., Kay, L. E. & Tugarinov, V. An optimized relaxation-based coherence transfer NMR experiment for the measurement of side-chain order in methyl-protonated, highly deuterated proteins. J. Phys. Chem. B 115, 14878–14884 (2011).
Linser, R. et al. Selective methyl labeling of eukaryotic membrane proteins using cell-free expression. J. Am. Chem. Soc. 136, 11308–11310 (2014).
Opitz, C., Isogai, S. & Grzesiek, S. An economic approach to efficient isotope labeling in insect cells using homemade 15N-, 13C- and 2H-labeled yeast extracts. J. Biomol. NMR 62, 373–385 (2015).
Sitarska, A. et al. Affordable uniform isotope labeling with (2)H, (13)C and (15)N in insect cells. J. Biomol. NMR 62, 191–197 (2015).
Cai, M. et al. An efficient and cost-effective isotope labeling protocol for proteins expressed in Escherichia coli. J. Biomol. NMR 11, 97–102 (1998).
Celver, J., Vishnivetskiy, S. A., Chavkin, C. & Gurevich, V. V. Conservation of the phosphate-sensitive elements in the arrestin family of proteins. J. Biol. Chem. 277, 9043–9048 (2002).
Carpenter, B. & Tate, C. G. Expression and purification of mini G proteins from Escherichia coli. Bio-Protoc. 7, e2235 (2017).
Kupce, E. & Freeman, R. Polychromatic selective pulses. J. Magn. Reson. Ser. A 102, 122–126 (1993).
Kupce, E., Boyd, J. & Campbell, I. D. Short selective pulses for biochemical applications. J. Magn. Reson. Ser. B 106, 300–303 (1995).
Hyberts, S. G., Milbradt, A. G., Wagner, A. B., Arthanari, H. & Wagner, G. Application of iterative soft thresholding for fast reconstruction of NMR data non-uniformly sampled with multidimensional Poisson Gap scheduling. J. Biomol. NMR 52, 315–327 (2012).
Kazimierczuk, K. & Orekhov, V. Y. Accelerated NMR spectroscopy by using compressed sensing. Angew. Chem. Int. Ed. Engl. 50, 5556–5559 (2011).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Acknowledgements
We are grateful to Prof. Aashish Manglik (UCSF) for providing the βArr1 construct used in this study. We would like to thank Prof. Andrew L. Lee (UNC) for fruitful discussions. The 14.1 T spectrometer at Indiana University used in this study was generously supported by the Indiana University Fund. Studies at the Florey Institute were supported by the Victorian Government’s Operational Infrastructure Support Program. The project was funded by: KAKENHI 21H04791 (A.I.), 21H051130 (A.I.), and JPJSBP120213501 (A.I.) from Japan Society for the Promotion of Science (JSPS); LEAP JP20gm0010004 (A.I.), and BINDS JP20am0101095 (A.I.) from the Japan Agency for Medical Research and Development (AMED); FOREST Program JPMJFR215T (A.I.) and JST Moonshot Research and Development Program JPMJMS2023 (A.I.) from Japan Science and Technology Agency (JST); Daiichi Sankyo Foundation of Life Science (A.I.); Takeda Science Foundation (A.I.); Ono Medical Research Foundation (A.I.); Uehara Memorial Foundation (A.I.); Agencia Estatal de Investigación, Spain grant PID2019-104914RB-I00 (J.C.P. and M.P.); Ministerio de Ciencia e Innovación, María de Maeztu grant CEX2021-001202-M (J.C.P.); Australian National Health and Medical Research Council (NHMRC) grants 1081844 and 1141034 (R.A.D.B., D.J.S., and P.R.G.); Indiana Precision Health Initiative (J.J.Z.).; and National Institutes of Health (NIH) grants R00GM115814 (J.J.Z.) and R35GM143054 (J.J.Z.).
Author information
Authors and Affiliations
Contributions
F.B. and F.Y. expressed and purified enNTS1 used for 2D 1H-13C SOFAST-HMQC experiments; F.B. expressed and purified unlabeled βArr1-3A and Gαiq used for 2D 1H-13C SOFAST-HMQC experiments; F.B. performed 2D 1H-13C SOFAST-HMQC experiments; J.B.B. performed MST experiments, and expressed and purified enNTS1, βArr1-3A, and Gαiq used therein; S.C.Z. expressed and purified [U-15N,13C,2H]-βArr1-3A; S.C.Z. and J.F. expressed and purified [U-15N,13C,2H]-Gαiq; S.C.Z. performed 15N-TROSY-HSQC experiments; M.P. and J.C.P. carried out DFT chemical shift calculations; F.B., J.B.B., S.C.Z., S.A.R., and J.J.Z. performed data analysis and data interpretation; A.I., M.P., J.C.P., H.W., and P.R.G. assisted with data interpretation; F.B., R.A.D.B., D.J.S., P.R.G., and J.J.Z. designed and supervised the project. F.B. and J.J.Z. wrote the manuscript with assistance from A.I., M.P., J.C.P., S.A.R., R.A.D.B., D.J.S., and P.R.G. All authors reviewed and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bumbak, F., Bower, J.B., Zemmer, S.C. et al. Stabilization of pre-existing neurotensin receptor conformational states by β-arrestin-1 and the biased allosteric modulator ML314. Nat Commun 14, 3328 (2023). https://doi.org/10.1038/s41467-023-38894-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-38894-8
- Springer Nature Limited