Abstract
BRCA1 deficient breast cancers are aggressive and chemoresistant due, in part, to their enrichment of cancer stem cells that can be generated from carcinoma cells by an epithelial-mesenchymal transition (EMT). We previously discovered that BRCA1 deficiency activates EMT in mammary tumorigenesis. How BRCA1 controls EMT and how to effectively target BRCA1-deficient cancers remain elusive. We analyzed murine and human tumors and identified a role for Tgfβr2 in governing the molecular aspects of EMT that occur with Brca1 loss. We utilized CRISPR to delete Tgfβr2 and specific inhibitors to block Tgfβr2 activity and followed up with the molecular analysis of assays for tumor growth and metastasis. We discovered that heterozygous germline deletion, or epithelia-specific deletion of Brca1 in mice, activates Tgfβr2 signaling pathways in mammary tumors. BRCA1 depletion promotes TGFβ-mediated EMT activation in cancer cells. BRCA1 binds to the TGFβR2 locus to repress its transcription. Targeted deletion or pharmaceutical inhibition of Tgfβr2 in Brca1-deficient tumor cells reduces EMT and suppresses tumorigenesis and metastasis. BRCA1 and TGFβR2 expression levels are inversely related in human breast cancers. This study reveals for the first time that a targetable TGFβR signaling pathway is directly activated by BRCA1-deficiency in the induction of EMT in breast cancer progression.
Similar content being viewed by others
Introduction
Clinically, breast cancer comprises three main subtypes: human epidermal growth factor receptor 2 (HER2) positive breast cancer, hormone receptor [estrogen receptor (ER) and/or progesterone receptor (PgR)]-positive breast cancer, and triple-negative breast cancer (TNBC), the latter of which lacks expression of ER, PgR, and HER2 [1, 2]. Gene-expression analyses have categorized human breast tumor into six intrinsic subtypes: basal-like (BL), claudin-low (CL), Her2-enriched, luminal A, luminal B, and normal breast-like, each of which has unique biological and prognostic features [3]. Of these subtypes of breast cancer, the CL subtype is a TNBC characterized by the high enrichment for epithelial to mesenchymal transition (EMT) markers and cancer stem cell (CSC)-like features [3, 4]. Basal-like breast cancer (BLBC) accounts for approximately 70% of TNBCs and is a leading cause of cancer deaths worldwide. The high mortality rate of BLBCs can be attributed to the aggressive and metastatic capacity of these tumors and the limited number of effective therapeutic options. BLBCs are highly aggressive and metastatic in part due to their enrichment of CSCs, which are thought to drive clinical relapse and metastasis [5, 6]. CSCs are more resistant to radio- and chemotherapy and can be generated from carcinoma cells by an EMT program [7,8,9], which is a process whereby epithelial cells lose many of their epithelial characteristics and acquire mesenchymal features [9]. While some molecular regulators of EMT have been identified [10], therapeutically targetable mechanisms controlling EMT in BLBCs remain elusive.
TGFβ signaling plays a key role in inducing and maintaining the mesenchymal and stem cell states of multiple tissues, including breast tissues [11]. TGFβ ligands bind to their receptors, TGFβR1/2, which then activate downstream effectors through SMAD-dependent and SMAD-independent pathways, to activate EMT and drive CSC function [12, 13]. Further, the expression of TGFβR2 is enhanced in ER-negative tumor cells, and is reduced in ER-positive tumor cells [14, 15]. Overexpression of TGFβR2 or upregulation of TGFβ signature genes is associated with lung metastasis and lower survival rates in BLBCs [16, 17]. Together, these prior studies demonstrate an association between TGFβR2 and aggressive tumor phenotypes. Yet, the role of TGFβR2 and its status as a putative therapy target in ER-negative BRCA1 deficient tumors is currently unknown.
We and others have reported that BRCA1 deficiency activates p16, p18, and RB, key proteins in controlling cell cycle progression [18,19,20,21]. Loss of Brca1 in p16 or p18 deficient mice activates EMT and induces BLBCs, whereas, loss of p18 induces luminal type mammary tumors [18,19,20, 22]. In this study, we follow up on these results to identify the molecular mechanisms by which Brca1 loss promotes EMT.
Results
Germline deletion of Brca1 activates Tgfβ signaling and EMT with enhanced expression of Tgfβr2 in mammary tumor cells
We previously demonstrated that heterozygous germline deletion of Brca1 in p18 null mice activated EMT and led to BLBCs with enhanced CSC populations along with the increased potential for metastasis {Figs. 1A, S1A–C, and details in [18, 19, 22]}. To identify the molecular determinants for activation of EMT in Brca1 deficient mammary tumors in an unbiased manner, we performed microarray analysis. GSEA revealed enrichment of a signature for active Tgfβ signaling in p18−/−; Brca1+/− tumors when compared to p18−/− and p18+/− tumors (Fig. 1B). Consistent with activation of known downstream pathways by BRCA1 loss [23, 24], signatures for NF-КB pathway activity were also enriched in p18−/−; Brca1+/− tumors (Fig. S1D). Most (9/10) p18−/−; Brca1+/− tumors expressed high levels of Tgfβr2, whereas most (8/10) p18−/− and p18+/− tumors expressed a low level of Tgfβr2. Analysis of tumors after removal of an outlier p18−/−;Brca1+/− tumor, a p18+/− tumor and an outlier p18−/− sample, revealed that Tgfβr2 mRNA levels in p18−/−;Brca1+/− tumors were significantly higher than those in p18−/− tumors (Fig. S2). Interestingly, we found that Tgfβr2 mRNA did not elevate in K14-Cre;p53f/f;Brca1f/f mammary tumors, suggesting that upregulation of Tgfβr2 may be unique to Brca1 germline mutant tumors also deficient in p18. We also noted that within individual intrinsic subtypes in genetically engineered mouse models existed a range of expression values for Tgfβr2, with statistically significant association of elevated mRNA in CL subtype (Fig. S2).
A Representative IHC analysis of mammary tumors with antibodies against Vim. Vim positive tumor cells (Black arrows) and stromal cells (Yellow arrows) are indicated. B Microarray analysis of mammary tumors. GSEA enrichment plot for a signature for Tgfβ signaling activity. NES: Normalized Enrichment Score (NES), Nominal p value (p), and False Discovery Rate q-value (FDR q) were detected comparing p18−/−;Brca1+/− tumors (n = 10) to p18−/− and p18+/− tumors including nine p18−/− and one p18+/− tumors. C Representative IHC analysis of mammary tumors from a p18−/− and p18−/−; Brca1+/− mice. To quantify the Tgfβr2 positive tumor cells, the intensity of Tgfβr2 antibody-specific staining by IHC in tumor cells were categorized into -, 1 + , 2 + , and 3 + . The representative images in the boxed area for each category were shown. D Summary of mammary tumors in mice with Balb/c background. EMT + tumors are tumors that are positive for at least two EMT markers (decreased E-Cad, increased Vim, Fn1, or CD29), and two EMT-TFs (Twist, Snail, Slug, Foxc2 or p-Fra1) in >2% tumor cells. Tgfβr2 + tumors are tumors that are positive for Tgfβr2 with 2+ or 3+ intensity in >20% tumor cells. The asterisk (*) denotes a significance from p18−/−; Brca1+/− and p18−/− tumors by two-tailed Fisher’s exact test. E Representative immunofluorescent staining of mammary tumors from a p18−/− and p18−/−; Brca1+/− mice. Note that the majority of Tgfβr2 positive p18−/−; Brca1+/− tumor cells were co-stained with Ck14 (blue arrows). Ck14 singly positive basal epithelial cells (white arrows) in the normal gland are indicated.
We performed IHC and found a significantly enhanced frequency of Tgfβr2-positive tumors when p18−/−; Brca1+/− tumors were compared with p18−/− tumors. Furthermore, most (11/12) p18−/−; Brca1+/− tumors with EMT features expressed high levels of Tgfβr2 (Fig. 1C, D). Interestingly, we observed an insignificantly increased frequency of p18−/−; Brca1+/− tumors that were Tgfβr1-positive when compared with p18−/− tumors (7/14 positive in p18−/−; Brca1+/− tumors versus 6 /14 positive in p18−/− tumors) (data not shown). Since TGFβR2 is also expressed in mammary stromal cells [12, 14], we determined the expression of Tgfβr2 and Ck14, an epithelial marker, in tumor cells. We noticed that the majority of Tgfβr2 positive p18−/−;Brca1+/− tumor cells were co-stained with Ck14, indicative of the epithelium origin of the Tgfβr2 positive tumor cells (Fig. 1E). These results imply a possible negative correlation between Brca1 and Tgfβr2 in mouse mammary tumors.
We examined the expression of Tgfβr1/2 in mouse embryos in which Tgfβ signaling and EMT play a critical role in development [9]. The expression of Tgfβr2, along with a few EMT-inducing transcription factors (EMT-TFs), was significantly increased in p18−/−; Brca1+/− embryos relative to their p18−/− counterparts. We noted that the expression of Tgfβr1 was not different in this same comparison (Fig. S3). Together, our data depicts a clear relationship between Brca1, Tgfβr2, and EMT in our murine system.
Specific deletion of Brca1 in mammary epithelia enhances Tgfβr2 expression and activates EMT in mammary tumors
We generated p18−/−; Brca1f/f; MMTV-Cre (p18−/−; Brca1MGKO) mice, in which Brca1 was specifically deleted in mammary epithelial cells (MECs). 47% (7/15) of p18−/− and 73% (11/15) of p18−/−; Brca1MGKO mice developed mammary tumors [19]. Interestingly, we observed lung metastasis in 36% (4/11) of p18−/−; Brca1MGKO tumor-bearing mice, but did not observe metastasis in the lungs of mice with p18−/− tumors. Consistent with prior reports of a linkage between EMT and metastatic potential [25], significantly more p18−/−; Brca1MGKO tumors than p18−/− tumors were positive for EMT markers including vimentin (Vim), fibronectin (Fn), Twist, and p-Fra1 {details in [19]}. These data demonstrate that specific deletion of Brca1 in p18 null mammary epithelia induces a malignant mammary tumor phenotype with EMT features.
We compared p18−/−; Brca1MGKO tumors to tumor-free mammary tissues, and to p18−/− tumors. Both Tgfβr2 mRNA and protein levels, as well as the level of phosphorylated-Smad2 (p-Smad2), were significantly increased in p18−/−; Brca1MGKO tumors relative to those in tumor-free mammary tissues of the same mice. In comparison, Tgfβr2 mRNA was moderately enhanced in p18−/− tumors relative to that in tumor-free mammary tissues of the same mice (Fig. 2A, B). Furthermore, when compared with p18−/− tumors, most p18−/−; Brca1MGKO tumors expressed an increased level of Tgfβr2 and p-Smad2, as well as p-Akt, p-Erk, and p-Jnk, all of which are Smad-independent downstream effectors activated by Tgfβ-Tgfβr2 signaling (Fig. 2C). Immunostaining revealed that Tgfβr2 and p-Smad2 were highly expressed in p18−/−; Brca1MGKO tumor cells, and that the expression pattern of Tgfβr2 in p18−/−; Brca1MGKO tumors was similar with the pattern in p18−/−; Brca1+/− tumors (Figs. 1C, E, 2D). When the positive cells were quantified, we found that H scores for both Tgfβr2 and p-Smad2 in p18−/−; Brca1MGKO tumor cells were significantly more than the H scores in p18−/− counterparts (Fig. 2E). These data confirm that during mammary tumorigenesis, loss of Brca1 activates Tgfβr2 signaling pathways in an epithelium-autonomous manner.
A, B Tumors (T) from p18−/− and p18−/−; Brca1MGKO mice were analyzed by qRT-PCR (A) and western blot (B). Tumor-free mammary glands (TF) from the same mouse were used as controls. Data in (A) represent the mean ± SD of three tumors in each group. The asterisk (*) denotes a significance from tumors and tumor-free tissues of the same genotype by a two-tailed, unpaired T test. C Four representative tumors from p18−/− and p18−/−; Brca1MGKO mice were analyzed by western blot. D Representative immunostaining of mammary tumors from a p18−/− and p18−/−; Brca1MGKO mice. E p18−/− and p18−/−; Brca1MGKO mammary tumors were analyzed by IHC, and H-scores for Tgfβr2 and p-Smad2 were calculated. The results represent the mean ± SD of five individual tumors per group for Tgfβr2 and four individual tumors per group for p-Smad2. F p18−/− and p18−/−; Brca1MGKO mammary tumor cells were transplanted into MFPs of NSG mice. Four weeks later, recipient mice with tumors generated (>0.5 cm3 in size) were analyzed. #five tumor-bearing mice displayed lung metastasis by H.E. analysis. G 3 ×104 p18−/− and p18−/−; Brca1MGKO mammary tumor cells were cultured to generate primary tumorspheres in 10 days. The number of spheres large than 50 μm was quantified from triplicate experiments. The results represent the mean ± SD of three individual tumors per group. Statistical significance in (E) and (G) was determined by a two-tailed, unpaired T test.
BRCA1 depletion enhances tumor initiation potential and promotes TGFβ-mediated EMT activation in breast cancer cells
We previously reported that CSC-enriched cells were expanded in p18−/−; Brca1MGKO mammary tumors relative to their p18−/− counterparts [19], and that p18−/−; Brca1MGKO tumor cells displayed features that had undergone EMT [26]. We tested the tumor-initiating capacity of tumor cells and found that as low as 1 ×105 of p18−/−; Brca1MGKO cells were able to regenerate tumors, whereas as high as 6 × 106 of p18−/− cells did not yield tumors. Notably, half of the mammary tumors generated by 6 × 106 p18−/−; Brca1MGKO tumor cells metastasized to the lung (Fig. 2F). Furthermore, we detected significantly more primary tumorspheres formed by p18−/−; Brca1MGKO tumor cells than those done by p18−/− tumor cells (Fig. 2G). Together, these illustrate that loss of Brca1 in breast cancer cells enhances the CSC population and its property in tumor initiation.
To test if Tgfβr2 activation promotes EMT in Brca1 deficient tumor cells, we used a TGFβ challenge assay. We confirmed the increase of Tgfβr2 expression in p18−/−; Brca1MGKO tumor cells relative to that in p18−/− cells (Fig. 3A). In p18−/− cells, TGFβ treatment resulted in modest changes in EMT marker genes (increase of mesenchymal and decrease of epithelial genes), which was likely due to the diminished Tgfβr2 expression observed in these tumor cells. However, in p18−/−; Brca1MGKO cells, the dramatic elevation of EMT markers was noted (Figs. 3B, S4A). We then knocked down BRCA1 in MCF7, T47D, and MDA-MB-231 cell lines, all of which are BRCA1 wild type (Wt) [19]. We discovered that the depletion of BRCA1 stimulated the expression of TGFβR2 mRNA. In response to TGFβ treatment, BRCA1-depleted cells, but not control cells, expressed a distinctly lower level of epithelial markers (CDH1, CK8) and higher level of mesenchymal markers (PDGFRβ, VIM), as well as EMT-TFs including FOSL1 (encoding FRA1), TWIST, and SNAIL (Figs. 3C–F, S4B, C). These results indicate that BRCA1 deficiency in cancer cells stimulates TGFβR2 expression and promotes TGFβ-mediated EMT activation.
A, B Representative p18−/− and p18−/−; Brca1MGKO mammary tumor cells were analyzed by western blot (A) or treated with vehicle or TGFβ for different time periods, and then analyzed by qRT-PCR (B). Data in (B) represent the mean ± SD from duplicates of two independent experiments from two different pairs of primary tumor cell lines. C, E MCF7 (C) and MDA-MB-231 (E) cells were infected with either pGIPZ-empty (sh-Ctrl.) or pGIPZ-shBRCA1 targeting different sequences of human BRCA1 (sh-BRCA1-B7 and sh-BRCA1-G6). Cells stably expressing sh-Ctrl or shBRCA1 were analyzed by qRT-PCR. Data represent the mean ± SD from triplicates of each of the two independent experiments. D, F mRNA levels in MCF7-sh-Ctrl and MCF7-sh-BRCA1-G6 (D), or MDA-MB-231-sh-Ctrl and MDA-MB-231-sh-BRCA1-G6 (F) cells treated with TGFβ for 10 (D) or 24 (F) hours were analyzed. Data represent the mean ± SD from duplicates of two independent experiments. The asterisk (*) in (B) (D) and (F) denotes a statistical significance from vehicle- and TGFβ-treated samples determined by a two-tailed, paired T test. The asterisk (*) in (C) and (E) denotes a statistical significance from sh-BRCA1 and sh-Ctrl samples determined by a two-tailed, paired T test.
BRCA1 suppresses transcription of TGFβR2
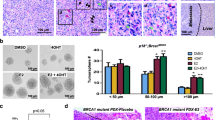
We transfected BRCA1 into BRCA1-mutant cell lines, HCC1937 and SUM149, respectively. The introduction of functional BRCA1 led to lower mRNA and protein levels of TGFβR2. Consistent with our previous finding [19], overexpression of BRCA1 resulted in a decrease of mesenchymal markers and EMT-TFs (Figs. 4A, S5). These data confirm that BRCA1 represses the expression of TGFβR2 and inhibits EMT.
A SUM149 cells were transfected with pBabe-empty (Empty) or pBabe-HA-BRCA1 (BRCA1). The expression of genes indicated was determined by western blot and/or qRT-PCR 72 h after transfection. Data represent the mean ± SD from triplicates of two independent experiments. The asterisk (*) denotes a statistical significance from empty- and BRCA1-expressing samples determined by a two-tailed, paired T test. B Diagram showing the location of putative BRCA1 binding sites (dark black bars) in the human TGFβR2 gene, and that of primers (P1 - P8) used for ChIP analysis. +1, transcription start site. C ChIP analysis of endogenous BRCA1 binding to putative BRCA1 sites on the TGFβR2 locus in T47D cells. Normal IgG was used as a negative control. The ratio of binding signal to input was compared. Data represent the mean ± SD from triplicates of two independent experiments. The asterisk (*) denotes a statistical significance from IgG and anti-BRCA1 immunoprecipitated samples by a two-tailed, unpaired T test. D ChIP analysis of exogenous BRCA1 binding to the TGFβR2 locus in HCC1937 cells transfected with pBabe-empty (Empty) or pBabe-HA-BRCA1 (BRCA1). Normal IgG was used as a negative control. The ratio of binding signal to input was compared. Data represent the mean ± SD from triplicates of two independent experiments. The asterisk (*) denotes a statistical significance from Empty- and BRCA1-expressing samples immunoprecipitated by anti-BRCA1 by a two-tailed, unpaired T test. E A schematic representation of WT and Mut TGFβR2 promoter-reporter constructs. Numbers represent a position relative to the transcription start site and a letter (X) denotes a mutated GATA3 binding site. The location of primers used for ChIP analysis is shown. F T47D-sh-Ctrl and T47D-sh-BRCA1 cells were transfected with Renilla and pGL3-TGFβR2-WT or pGL3-TGFβR2-Mut (right panel), and SUM149 cells were transfected with Renilla and pGL3-TGFβR2-WT or pGL3-TGFβR2-Mut, as well as pBabe-empty or pBabe-HA-BRCA1 (left panel), which were then collected after 48 h and assayed for luciferase activity. Data represent the mean ± SD from triplicates of two independent experiments. The asterisk (*) denotes a statistical significance from BRCA1 and empty (left panel), or sh-BRCA1 and sh-Ctrl (right panel) samples determined by a two-tailed, paired T test. SUM149, BRCA1 mutant (Mut); T47D, BRCA1 wild type (Wt).
We and others have shown that GATA3 recruits BRCA1 to its binding sites in the promoters of FOXC1/2 and TWIST genes to repress their transcription [19, 27]. Analysis of the TGFβR2 locus revealed at least twelve putative BRCA1/GATA3 binding sites (Fig. 4B). We performed ChIP assays using 8 pairs of primers that cover all twelve putative BRCA1/GATA3 binding sites. Six out of eight amplicons (P1, P2, P4, P5, P7, and P8) in T47D cells and three out of eight amplicons (P1, P2, and P5) in HCC1937 cells transfected with BRCA1 were specifically enriched in the immunoprecipitation of BRCA1 (Fig. 4C, D). Together with the experimental evidence that BRCA1 represses TGFβR2, we attribute these observations to direct repression by the binding of BRCA1 to the TGFβR2 locus.
We generated TGFβR2 promoter-luciferase fusion plasmids, pGL3-TGFβR2-WT containing a promoter region that covers P5 amplicon, and pGL3-TGFβR2-Mut in which a BRCA1/GATA3 binding site of pGL3-TGFβR2-WT was mutated (Fig. 4E). We transfected the plasmids into SUM149 cells and found that relative to an empty control; ectopic BRCA1 drastically reduced the activity of pGL3-TGFβR2-WT promoter by ~60%. On the other hand, the transfection slightly reduced the activity of pGL3-TGFβR2-Mut promoter by ~18% (Fig. 4F, left). Furthermore, we detected that the activity of pGL3-TGFβR2-WT promoter was significantly increased in T47D-sh-BRCA1 cells relative to their T47D-sh-Ctrl cell counterparts. However, this was not the case when we examined the activity of pGL3-TGFβR2-Mut promoter (Fig. 4F, right). These data reaffirm our conclusion that BRCA1 binds to BRCA1/GATA3 sites in TGFβR2 promoter to suppress its activity.
Targeted deletion of Tgfβr2 in Brca1 deficient tumor cells suppresses EMT and inhibits tumor initiation and metastasis
We knocked out Tgfβr2 in p18−/−; Brca1MGKO tumor cells and found that Tgfβr2 KO led to lower expression of the EMT markers Vim, Fn, Snail, Zeb1, Slug, and higher expression of the epithelial markers including Cdh1 and Epcam (Fig. 5A, B). Morphologically, Tgfβr2-depleted tumor cells exhibited more epithelial-like phenotypes including islets of cells with close cell-cell contacts and cobblestone morphology, but less mesenchymal-like phenotypes including a reduction in spindle-shaped cells (Fig. 5C). We then treated cells with TGFβ, and found that in response to the treatment the elevation of Fn and Vim in control-depleted cells was drastically reduced in Tgfβr2-depleted cells (Figs. 5D, S6A). These results indicate that depletion of Tgfβr2 in p18−/−; Brca1MGKO tumor cells inhibits EMT in vitro.
A–C p18−/−; Brca1MGKO tumor cells were transfected with Tgfβr2 (Tgfβr2 CRISPR) and Control (Ctrl CRISPR) Double Nickase plasmids then selected with puromycin for 3 days. Tgfβr2- and control-knockout cells were then analyzed by western blot (A), Q-RT-PCR (B), and microscope for cell morphology (C). The areas with epithelial-like cells are indicated in (C). D Tgfβr2- and Ctrl-knockout p18−/−; Brca1MGKO mammary tumor cells were treated with vehicle or TGFβ for different time periods, and the expression of Fn was then determined. Data in (B) and (D) represent the mean ± SD from duplicates of two independent experiments. The asterisk (*) in (B) denotes a statistical significance from Tgfβr2- and Ctrl-knockout samples, and in (D) denotes a statistical significance from TGFβ treated and vehicle treated samples determined by a two-tailed, paired T test. E, F 6 ×105 Tgfβr2- and Ctrl-knockout p18−/−; Brca1MGKO mammary tumor cells were inoculated into the left and right inguinal MFPs of NSG mice, respectively, in a pairwise manner. Two weeks after transplantation, mice were dissected. The volume (E) and expression of Vim (F) of the regenerated tumors were determined by measurement and IHC. Data in (E) represent the mean ± SD of four tumors in each group. The asterisks (*) denote a statistical significance from Tgfβr2- and Ctrl-knockout tumors determined by a two-tailed, paired T test. G–I 6 ×105 Tgfβr2- or Ctrl-knockout p18−/−; Brca1MGKO mammary tumor cells were inoculated into the MFPs of NSG mice. When newly generated tumors reached the maximum size allowed by IACUC in 4–7 weeks, or the mice became moribund, lungs were examined for gross appearance (G), H.E. staining (H), and quantification of the number of metastatic nodules (I). M, metastatic nodules. Data in (I) represent the mean ± SD for the numbers of metastatic nodules detected in all lobes of the lungs in each group (n = 4). Statistical significance was determined by a two-tailed, unpaired T test.
We transplanted tumor cells into mice. Though all four mice received 6 × 105 Tgfβr2- and control-depleted p18−/−; Brca1MGKO tumor cells developed tumors, tumors generated by Tgfβr2-knockout cells were significantly smaller than tumors generated by control cells (Fig. 5E). Tumors generated by Tgfβr2-depleted cells expressed noticeably reduced levels of Vim and p-Fra1 when compared with tumors generated by control cells (Figs. 5F, S6B). These results demonstrate that the deletion of Tgfβr2 in Brca1-deficient tumor cells inhibits EMT and suppresses tumorigenesis.
We also discovered that mammary tumors generated by Tgfbr2 KO cells produced significantly less metastatic nodules in the lung when compared with the tumors initiated by control cells (Fig. 5G-I). These results indicate that the deletion of Tgfβr2 inhibits the metastatic potential of Brca1 deficient mammary tumor cells.
Inhibition of Tgfβr2 in Brca1 deficient tumor cells suppresses EMT and tumor initiation
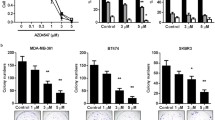
We treated p18−/−; Brca1MGKO tumor cells with a TGFβR2 inhibitor ITD1, which specifically enhances degradation of Tgfβr2 [28]. We confirmed that ITD1 treatment reduced the level of Tgfβr2 and its downstream effector protein p-Smad2 (Fig. 6A). ITD1 treatment of the cells from primary p18−/−; Brca1MGKO tumorspheres significantly reduced secondary tumorsphere formation (Fig. 6B). We transplanted tumorsphere-dissociated cells that were pretreated with ITD1 into mice. Pretreatment of p18−/−; Brca1MGKO tumorsphere-dissociated cells with ITD1 led to significantly smaller tumors than those with vehicle pretreatment (Fig. 6C). Analysis of the newly generated tumors revealed that relative to control tumors, ITD1 tumors exhibited less Tgfβr2, p-Smad2, and EMT-TFs including Twist and Snail, but more E-Cad (Fig. 6D, E). These data suggest that pharmaceutical inhibition of Tgfβr2 in Brca1-deficient tumor cells suppresses EMT and their potential for tumor initiation.
A 18−/−; Brca1MGKO tumor cells treated with ITD1 at 4 μm for 24 h were analyzed by western blot. B 104 cells dissociated from18−/−; Brca1MGKO primary tumorspheres were treated with ITD1. Secondary spheres formed after 6 days of treatment were counted from quadruplicate experiments. Data represent the mean ± SD from two independent primary tumorspheres. C 1000 p18−/−; Brca1MGKO tumorsphere-dissociated cells pretreated with DMSO or ITD1 for 6 days were transplanted into MFPs of NSG mice. Four weeks later, mice were dissected and tumor volumes were measured. Values represent the average tumor volumes ± SD of four tumors. The asterisk (*) in (B) and (C) denotes a statistical significance from ITD1 treated and DMSO treated samples determined by a two-tailed, unpaired T test. D, E Representative tumors generated by DMSO- or ITD1-pretreated p18−/−; Brca1MGKO cells were analyzed by Western blot (D) and IHC (E). The H-scores for p-Smad2, Twist, and Tgfβr2 in IHC were calculated (E, right panel). The results represent the mean ± SD of four individual tumors per group. Statistical significance was determined by a two-tailed, unpaired T test.
BRCA1 and TGFβR2 expression levels are inversely related in human breast cancers
Prompted by these observations in mouse models and prior studies showing enhanced expression of TGFβR2 in CSC-enriched BLBC cells [14, 15], we examined the relationship between BRCA1 with TGFβR2 mRNA levels in human breast cancer sample sets [3, 29, 30]. Just as in our mouse model, we observed a significantly inverse relationship between BRCA1 and TGFβR2, and a less significantly inverse relationship between BRCA1 and TGFβR1 (Figs. 7A, S7B). Consistent with our observations in mouse models, human CL tumors show low BRCA1 mRNA and higher TGFβR2 mRNA levels (Figs. 7B, S7A).
A Correlation analysis of the expression of BRCA1 and TGFβR2 for MetaBric breast cancer patients. B Analysis of gene expression in NKI295 breast cancer patients according to tumor subtype. BL basal-like, CL claudin-low, H2 Her2-enriched, LA luminal A, LB luminal B, NBL normal breast-like. C Representative immunostaining of ER positive and negative invasive human breast cancers with antibodies against TGFβR2. D, E Summary of expression of TGFβR2 by IHC and BRCA1 by Q-RT-PCR (D). The expression levels of TGFβR2 were quantified by H-scores. The expression of BRCA1 was determined by Q-RT-PCR. BRCA1 mRNA levels relative to that of T47D cells were determined as we previously reported [19]. Analysis of the expression of TGFβR2 by IHC and BRCA1 by Q-RT-PCR in ER + and ER-invasive breast cancers (E). Statistical significance was determined by a two-tailed, unpaired T test. F Correlation analysis of BRCA1 mRNA levels and H scores of TGFβR2 expression for breast cancer samples.
To test if these observations are consistent at the protein level we turned to our previously published resource of 43 invasive breast cancers [19]. We found that TGFβR2 H scores were significantly higher in ER-negative than ER-positive tumors, while BRCA1 mRNA was lower in ER-negative tumors. Importantly, across both ER-negative and ER-positive tumors, we observed a significant inverse relationship between TGFβR2 protein and BRCA1 mRNA levels (Figs. 7C–F, S7C). Together, these clinical findings are consistent with our results in mice, thereby suggesting an opportunity to use murine systems to further explore how BRCA1 status, TGFβR2 signaling, and EMT intersect to control the biology of human tumors.
We examined EMT, basal-like, and CL signatures in the TCGA breast cancer dataset [3, 31, 32]. Not accounting for intrinsic subtype, BRCA1 mutant tumors showed an elevation of EMT/CL-like features as compared to BRCA1 Wt tumors (Fig. S8A), although not to the same extent as CL tumors. Since none of the five CL tumors were BRCA1 mutant, we were unable to analyze the correlation between BRCA1 mutation status and CL features in this subtype. Thus, we next limited our analysis to basal-like breast cancers, and again found a tendency for enhanced EMT/CL-like features in BRCA1 basal-like tumors, however, these differences were not statistically significant when compared to BRCA1 Wt tumors. We speculate that these observations might indicate that loss of BRCA1 leads to more EMT-like features in a proportion of cells within a tumor, but not the extent that the intrinsic subtype of the tumor bulk is significantly changed. In agreement, we observed that the basal subtype expression signature in BRCA1 mutant basal-like tumors also trended lower than BRCA1 Wt basal-like tumors (Fig. S8B).
Discussion
A number of studies have reported a relationship between BRCA1 and the basal-like TNBC subtype [2, 33, 34]. Here we expand on these studies by showing Brca1 alters the expression of Tgfβr2 to elevate Tgfβ signaling and EMT in breast cancer cells. Targeted deletion of, or pharmaceutical inhibition of, Tgfβr2 in Brca1-deficient tumor cells reduced EMT and suppressed tumorigenesis and metastasis. Moreover, we found BRCA1 mRNA was low and TGFβR2 mRNA was high in the claudin-low TNBC subtype. Thus, BRCA1 function may not only be implicated in the basal-like molecular subtype but also in claudin-low as well. Importantly, this also builds upon the body of work showing a correlation between EMT and the CL subtype [4, 35, 36] by showing a mechanistic connection between BRCA1 and TGFβ signaling in an epithelium-autonomous manner.
While our study demonstrates TGFβR2 induces malignant phenotypes in p18/BRCA1-deficient tumors, other studies show opposing roles in tumor suppression or tumor promotion [12, 14]. In favor of the malignant role for TGFβR2, overexpression of TGFβR2 or upregulation of TGFβ signature genes is associated with lung metastasis as well as lower survival in human BLBCs [16, 17], and higher stromal TGFβR2 is associated with poorer prognosis breast cancers [37]. Inhibition of TGFβR2 in human or mouse breast cancer cell lines reduces metastasis and blocks chemotherapy-induced CSC expansion [14, 38]. In support of the tumor suppressor function of TGFβR2, it has been found that in humans, reduction of or loss of TGFβR2 is associated with an increased risk of developing high-grade carcinoma in situ and invasive breast cancer [39, 40]. In addition, loss of TGFβR2 in stromal cancer-associated fibroblasts is linked to shortened patient survival [41]. In mouse models, the deletion of Tgfβr2, or expression of dominant-negative type Tgfβr2, results in or accelerates mammary tumor development and progression [14]. Collectively, these prior data and our study suggest that the role of Tgfβr2 is context-dependent and possibly related to BRCA1 function/regulation.
Understanding the genomic context contributing to Tgfβr2 expression and function could be important for guiding therapies. In particular, our study suggests that a targetable TGFβR signaling pathway exists in many BRCA1-deficient TNBC. Very few therapeutic options are available for ER-negative BLBCs. More than half of BLBCs have a dysfunctional BRCA1 pathway and harbor defects in DNA damage repair which make these patients initially respond well to DNA-damaging agents [2, 34, 42]. However, tumor recurrence and acquired resistance to DNA-damaging agents combine to decrease the survival of such patients [43, 44]. In addition, most BRCA1-deficient BLBCs carry a dysfunctional INK4-RB pathway [2, 45,46,47]. Specifically, our combination of p18 and Brca1 loss model those tumors with INK4-RB pathway deficits. Therefore, our results might suggest a context where Tgfβr2 might contribute to malignancy in human TNBC and reveal, for the first time, that a targetable TGFβR signaling pathway is directly activated by BRCA1-deficiency in the induction of EMT in breast cancers.
While this is all very straightforward in our controlled in vitro and mouse model systems, the manifestation in human tumors is more complex. In the limited number of CL tumors examined, which have molecular similarities to EMT [35], patterns of BRCA1 and TGFβR2 matched our experimental data in mouse models and human cell lines. Given that none of these tumors was BRCA1 mutant, these processes might also be epigenetically influenced. Further, our analysis of human basal-like tumors suggests that BRCA1 mutations might impact the proportion of cells with EMT-like features as there appears to be a trade-off between “basal-ness” and “claudin-low-ness” tracking with BRCA1 status. This may imply that additional cues are required to drive EMT in these cells, which we anticipate might include local production of TGFβ; additional experimental studies will be needed to confirm this.
The above observations highlight our incomplete understanding of the mechanisms leading to EMT and claudin-low characteristics in TNBC. Yet, with the parallels between our mouse models and human tumors (and cell lines), we anticipate continued progress using this murine system to understand TNBC tumor heterogeneity. In addition to carrying a dysfunctional INK4-RB pathway, most BRCA1-deficient BLBCs harbor p53 mutations [2, 45,46,47]. Most studies of Brca1 in mice co-mutate Brca1 with one of the genes in the p53 pathway [48]. However, mutation of p53 alone stimulates TGFβR2 and induces basal-like and CL subtype mammary tumors [4, 49, 50]. Our results demonstrate that Brca1 suppresses TGFβR2-mediated EMT in mammary tumorigenesis in the context of INK-RB inactivation under a genetically intact p53 background. Further, we find that some of the Brca1 deficient mammary tumors are CL [3, 33, 51] and that Brca1 deficiency induces basal-like mammary tumors with activation of EMT [18, 19], which is recently confirmed by an independent group [52, 53]. Thus, we provide an important mouse model for specific subsets of TNBC and uncover key molecular alterations that govern tumor heterogeneity and the malignant potential of these tumor types that might be effectively managed by targeting the TGFβ pathway.
Materials and methods
Mice, histopathology, and immunostaining
The generation of p18−/−, p18+/−, and p18−/−:Brca1+/− mice in Balb/c background, and p18−/− and p18−/−;Brca1f/f; MMTV-Cre (p18−/−; Brca1MGKO) mice in Balb/c-B6 mixed background has been previously described [18, 19, 54]. The Institutional Animal Care and Use Committee at the University of Miami and Shenzhen University approved all animal procedures. Animals were housed in a specific pathogen-free environment with a 12/12 light cycle. Animals were euthanized by exposure to isoflurane followed by cervical dislocation. At least four female mice were analyzed for each genotype, or were transplanted with each type of tumor cells. Histopathology and immunohistochemistry (IHC) were performed as previously described [18, 19, 54]. The primary antibodies used were: TGFβR1, TGFβR2 (Santa Cruz), p-Smad2, p-FRA1 (Cell signaling), CK5 (Covance), and Vim (Abcam). Immunocomplexes were detected using the Vectastain ABC DAB kit according to the manufacturer’s instructions (Vector Laboratories). The positive results of IHC were quantified by H-score, as previously described [55].
Mammary tumor cell preparation, cell culture, tumorsphere formation assay, overexpression and knockdown of BRCA1, and TGFβ treatment
Primary mammary tumors were dissected and cell suspensions were prepared as previously described [18, 19, 54]. T47D, HCC1937, SUM149, MDA-MB-231, and MCF7 cells were cultured per ATCC recommendations. For primary tumorsphere formation assay, mammary tumor cells were plated onto ultra-low attachment plates, in serum-free DMEM-F12, as previously described [19, 54]. Primary tumorspheres formed were collected and counted after 10 days of culture. For the secondary tumorsphere formation assay, primary tumorspheres formed were collected and dissociated. 104 dissociated cells were plated in triplicates with or without ITD1 (BioVision) treatment. Secondary tumorspheres that formed after 6 days of culture were counted. For ectopic expression of BRCA1, HCC1937 and SUM149 cells were transfected with pBabe-empty or pBabe-HA-BRCA1 as previously described [19]. For BRCA1 knockdown, T47D, MCF7, and MDA-MB-231 cells were infected with pGIPZ-empty (Sh-Ctrl) and pGIPZ-shBRCA1 (Sh-BRCA1) lentiviral vectors that were purchased from Open Biosystems, as previously described [19]. For TGFβ treatment, cells were starved in a serum-free medium for 24 h and then exposed to DMSO or TGFβ with an indicated period before analysis.
CRISPR-mediated Tgfβr2 knockout, transplantation, and analysis of metastasis
For CRISPR-mediated Tgfβr2 knockout in primary tumor cells, Tgfβr2 Double Nickase and control Double Nickase plasmids (Santa Cruz) were transfected into p18−/−; Brca1MGKO primary tumor cells, respectively, following the manufacturer’s protocol. After selection with puromycin for 3 days, GFP-positive cells were FACS sorted for further analysis. For in vivo transplantation, primary p18−/−; Brca1MGKO and p18−/− or Tgfβr2- and control-depleted p18−/−; Brca1MGKO tumor cells were suspended in a 50% solution of Matrigel (BD) and then inoculated into the left and right inguinal mammary fat pads (MFPs) of 4-week-old female NSG mice (Jackson Laboratory), respectively, in a pairwise manner. Four or two weeks after transplantation, animals were euthanized and mammary tumors were analyzed. For ex vivo transplantation, primary p18−/−; Brca1MGKO tumor cells were cultured to generate primary tumorspheres. 104 cells dissociated from primary tumorspheres were treated with DMSO or ITD1 for 6 days. 1,000 live cells were transplanted into MFPs of NSG mice. Four weeks after transplantation, animals were euthanized and mammary tumors were analyzed. The tumor size was measured daily with a caliper. Tumor volumes were calculated as V = a × b2/2, while “a” is the largest diameter and “b” is the smallest.
For analysis of lung metastasis from mammary tumors, p18−/−; Brca1MGKO tumor cells were inoculated into the MFPs of NSG mice. When newly generated tumors either reached the IACUC designated endpoint size (1.3 cm3; in 4–7 weeks) or the mice became moribund, the lungs were examined for detection of metastasis. For quantification of the number of metastatic nodules in the lungs, fixed lung tissues of all five lobes were sagittally sectioned at 200-μm intervals. At least three sections for each lobe were prepared and stained with H.E. The metastatic nodules in each lobe of lung tissue were confirmed by H.E. staining, counted under a microscope, and averaged. The number of nodules in all lobes was then calculated.
Microarray analysis, western blot, and chromatin-immunoprecipitation (ChIP) assay
RNA was extracted and purified from tumors using a RNeasy kit (Qiagen). Tumor RNA was reverse transcribed, amplified, and labeled with Cy5. Wt mammary tissue reference RNA was reverse transcribed, amplified, and labeled with Cy3. The amplified sample and reference were co-hybridized to Agilent 4x180k custom mouse microarrays and were analyzed as previously described [51]. Gene expression data for p18−/−, p18+/−, and p18−/−: Brca1+/− tumors was uploaded to the UNC microarray database. The Log(base2) of R/G Lowess Normalized Ratio (Mean) was taken for each array. For each channel, Lowess Normalized Net (Mean) greater than or equal to 10 was used to determine the presence of a signal. Genes present across 70% of samples were maintained and the remaining missing values were calculated and replaced using KNN imputation. Next genes were median centered and samples were standardized for the working gene matrix. Gene set enrichment analysis (GSEA) was performed using the GenePattern server [56, 57]. The gene signatures for Tgfβ and NF-КB signaling activities and EMT were published [35, 58, 59]. Gene expression data of mammary tumors coming from p18−/−, p18+/−, and p18−/−;Brca1+/− mice were deposited on the Gene Expression Omnibus under accession GSE155239. Data were obtained using Agilent microarrays as published [60]. Data were combined with a published dataset containing K14-Cre;p53f/f;Brca1f/f mouse mammary tumors as well as additional intrinsically credentialed genetically engineered mouse models (GEMMs) using the UNC Microarray database [33]. Intensity values were lowess normalized and relative counts established using median centering.
For the western blot, tissue and cell lysates were prepared as previously reported [19, 54]. The primary antibodies used were: BRCA1 and TGFβR2, (Santa Cruz), Vimentin, E-cad, Snail, Twist, p-Fra1, p-Smad2, p-Erk, p-Jnk, p-Akt (Cell signaling), and Gapdh (Ambion). All blots were derived from the same experiment and were processed in parallel. ChIP assays were carried out as previously described [19, 54]. Briefly, cells were treated with 1.5% formaldehyde and sonicated. Anti-BRCA1 antibody (D-9, Santa Cruz) or control mouse IgG was used to precipitate chromatin associated with BRCA1. Q-PCR was performed to determine the relative abundance of target DNA. Specific primers for the analysis of BRCA1 binding to TGFβR2 are listed in Table. S1.
Dual-luciferase reporter assay
The human TGFβR2 promoter region -2589 to -1613 that covers P5 and part of P4 primers used for ChIP analysis and contains a GATA3/BRCA1 binding site with consensus sequences, TGATTG, at -2115 was inserted into the pGL3-basic (Promega), as pGL3-TGFβR2-WT. With pGL3-TGFβR2-WT as a template, GATA3 binding site TGATTG was mutated into TACTTG to generate pGL3-TGFβR2-Mut by site-directed mutagenesis. Renilla plasmid was a gift from S. Y. Fuchs (University of Pennsylvania). For the TGFβR2 promoter-luciferase reporter assay, T47D-sh-Ctrl, T47D-sh-BRCA1, and SUM149 cells were seeded in a 24-well plate. T47D-sh-Ctrl and T47D-sh-BRCA1 cells were transfected with Renilla and either pGL3-TGFβR2-WT or pGL3-TGFβR2-Mut. SUM149 cells were transfected with Renilla plus pGL3-TGFβR2-WT or pGL3-TGFβR2-Mut, as well as pBabe-empty or pBabe-HA-BRCA1. The transfection efficiency was monitored using the Renilla vector as an internal control. Two days after transfection, cell lysates were collected and subjected to luciferase assay using the Dual-Luciferase Reporter Assay System (Promega). Two independent transfection experiments were conducted, and each luciferase assay was performed in triplicates. Normalized data was calculated as the ratio of the firefly/Renilla luciferase activities.
Human tumor samples and gene-expression data sets
Formalin-fixed paraffin-embedded (FFPE) human breast cancer samples lacking patient-identifying information were obtained from the Tissue Bank Core Facility at the University of Miami and the Department of Pathology at Shenzhen University. Samples used for this study consisted of non-treated invasive breast carcinomas with known ER status. Tumor samples were microdissected. Samples with tumor cell content >75% were used for RNA extraction. The expression of BRCA1 was determined by Q-RT-PCR, as previously reported [19]. Pearson correlation coefficient and P value (two-tailed) of BRCA1 mRNA levels and H scores of TGFβR2 expression for breast cancer samples were calculated by GraphPad Prism software. The correlation of expression of TGFβR1/2 mRNA with BRCA1 mRNA was analyzed with NKI295 [29], MetaBric [30], and UNC337 [3] human breast cancer datasets. The NKI295 dataset was analyzed to compare gene expression versus six breast cancer subtypes using a two-way analysis of variance (ANOVA).
Statistical analysis
All data are presented as the mean ± SD for at least three repeated individual experiments for each group. Quantitative results were analyzed by two tailed Fisher Exact test or two-tailed Student’s t-test. P < 0.05 was considered statistically significant.
Pearson correlation coefficient of BRCA1 and TGFβR2 expression was calculated by GraphPad Prism software. The NKI295 dataset was analyzed to compare gene expression versus six breast cancer subtypes using a two-way analysis of variance (ANOVA).
Data availability
Murine tumor gene expression data was deposited at the Gene Expression Omnibus under accession number GSE155239. All other datasets can be found as part of this manuscript.
References
Althuis MD, Fergenbaum JH, Garcia-Closas M, Brinton LA, Madigan MP, Sherman ME. Etiology of hormone receptor-defined breast cancer: a systematic review of the literature. Cancer Epidemiol Biomark Prev. 2004;13:1558–68.
Koboldt DC, Fulton RS, McLellan MD, Schmidt H, Kalicki-Veizer J, McMichael JF, et al. Comprehensive molecular portraits of human breast tumours. Nature 2012;487:330–7.
Prat A, Parker JS, Karginova O, Fan C, Livasy C, Herschkowitz JI, et al. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res. 2010;12:R68.
Herschkowitz JI, Zhao W, Zhang M, Usary J, Murrow G, Edwards D, et al. Comparative oncogenomics identifies breast tumors enriched in functional tumor-initiating cells. Proc Natl Acad Sci USA. 2012;109:2778–83.
Wright MH, Calcagno AM, Salcido CD, Carlson MD, Ambudkar SV, Varticovski L. Brca1 breast tumors contain distinct CD44+/CD24- and CD133+ cells with cancer stem cell characteristics. Breast Cancer Res. 2008;10:R10.
Wicha MS. Cancer stem cell heterogeneity in hereditary breast cancer. Breast Cancer Res. 2008;10:105.
Ye X, Weinberg RA, Epithelial-Mesenchymal. Plasticity: A Central Regulator of Cancer Progression. Trends cell Biol. 2015;25:675–86.
Mani SA, Guo W, Liao MJ, Eaton EN, Ayyanan A, Zhou AY, et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 2008;133:704–15.
Kalluri R, Weinberg RA. The basics of epithelial-mesenchymal transition. J Clin Invest. 2009;119:1420–8.
Ye X, Tam WL, Shibue T, Kaygusuz Y, Reinhardt F, Ng Eaton E, et al. Distinct EMT programs control normal mammary stem cells and tumour-initiating cells. Nature 2015;525:256–60.
Scheel C, Eaton EN, Li SH, Chaffer CL, Reinhardt F, Kah KJ, et al. Paracrine and autocrine signals induce and maintain mesenchymal and stem cell states in the breast. Cell 2011;145:926–40.
Moses H, Barcellos-Hoff MH. TGF-beta biology in mammary development and breast cancer. Cold Spring Harb Perspect Biol. 2011;3:a003277.
Derynck R, Muthusamy BP, Saeteurn KY. Signaling pathway cooperation in TGF-beta-induced epithelial-mesenchymal transition. Curr Opin Cell Biol. 2014;31:56–66.
Tan AR, Alexe G, Reiss M. Transforming growth factor-beta signaling: emerging stem cell target in metastatic breast cancer? Breast Cancer Res Treat. 2009;115:453–95.
Shipitsin M, Campbell LL, Argani P, Weremowicz S, Bloushtain-Qimron N, Yao J, et al. Molecular definition of breast tumor heterogeneity. Cancer Cell. 2007;11:259–73.
Buck MB, Fritz P, Dippon J, Zugmaier G, Knabbe C. Prognostic significance of transforming growth factor beta receptor II in estrogen receptor-negative breast cancer patients. Clin Cancer Res. 2004;10:491–8.
Padua D, Zhang XH, Wang Q, Nadal C, Gerald WL, Gomis RR, et al. TGFbeta primes breast tumors for lung metastasis seeding through angiopoietin-like 4. Cell 2008;133:66–77.
Bai F, Smith MD, Chan HL, Pei XH. Germline mutation of Brca1 alters the fate of mammary luminal cells and causes luminal-to-basal mammary tumor transformation. Oncogene 2013;32:2715–25.
Bai F, Chan HL, Scott A, Smith MD, Fan C, Herschkowitz JI, et al. BRCA1 Suppresses Epithelial-to-Mesenchymal Transition and Stem Cell Dedifferentiation during Mammary and Tumor Development. Cancer Res. 2014;74:6161–72.
Scott A, Bai F, Chan HL, Liu S, Ma J, Slingerland JM, et al. p16INK4a suppresses BRCA1-deficient mammary tumorigenesis. Oncotarget 2016;7:84496–507.
Sedic M, Skibinski A, Brown N, Gallardo M, Mulligan P, Martinez P, et al. Haploinsufficiency for BRCA1 leads to cell-type-specific genomic instability and premature senescence. Nat Commun. 2015;6:7505.
Bai F, Zhang LH, Liu X, Wang C, Zheng C, Sun J, et al. GATA3 functions downstream of BRCA1 to suppress EMT in breast cancer. Theranostics 2021;11:8218–33.
Mao R, Fan Y, Mou Y, Zhang H, Fu S, Yang J. TAK1 lysine 158 is required for TGF-beta-induced TRAF6-mediated Smad-independent IKK/NF-kappaB and JNK/AP-1 activation. Cell Signal. 2011;23:222–7.
Sakurai H, Shigemori N, Hasegawa K, Sugita T. TGF-beta-activated kinase 1 stimulates NF-kappa B activation by an NF-kappa B-inducing kinase-independent mechanism. Biochem Biophys Res Commun. 1998;243:545–9.
Andrechek ER, Cardiff RD, Chang JT, Gatza ML, Acharya CR, Potti A, et al. Genetic heterogeneity of Myc-induced mammary tumors reflecting diverse phenotypes including metastatic potential. Proc Natl Acad Sci USA. 2009;106:16387–92.
Wang C, Bai F, Zhang LH, Scott A, Li E, Pei XH. Estrogen promotes estrogen receptor negative BRCA1-deficient tumor initiation and progression. Breast Cancer Res. 2018;20:74.
Tkocz D, Crawford NT, Buckley NE, Berry FB, Kennedy RD, Gorski JJ, et al. BRCA1 and GATA3 corepress FOXC1 to inhibit the pathogenesis of basal-like breast cancers. Oncogene. 2011;31:3667–78.
Willems E, Cabral-Teixeira J, Schade D, Cai W, Reeves P, Bushway PJ, et al. Small molecule-mediated TGF-beta type II receptor degradation promotes cardiomyogenesis in embryonic stem cells. Cell Stem Cell. 2012;11:242–52.
van de Vijver MJ, He YD, van’t Veer LJ, Dai H, Hart AA, Voskuil DW, et al. A gene-expression signature as a predictor of survival in breast cancer. N. Engl J Med. 2002;347:1999–2009.
Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 2012;486:346–52.
Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol. 2009;27:1160–7.
Taube JH, Herschkowitz JI, Komurov K, Zhou AY, Gupta S, Yang J, et al. Core epithelial-to-mesenchymal transition interactome gene-expression signature is associated with claudin-low and metaplastic breast cancer subtypes. Proc Natl Acad Sci USA. 2010;107:15449–54.
Hollern DP, Contreras CM, Dance-Barnes S, Silva GO, Pfefferle AD, Xiong J, et al. A mouse model featuring tissue-specific deletion of p53 and Brca1 gives rise to mammary tumors with genomic and transcriptomic similarities to human basal-like breast cancer. Breast Cancer Res Treat. 2019;174:143–55.
Zhu X, Shan L, Wang F, Wang J, Wang F, Shen G, et al. Hypermethylation of BRCA1 gene: implication for prognostic biomarker and therapeutic target in sporadic primary triple-negative breast cancer. Breast Cancer Res Treat. 2015;150:479–86.
Hollern DP, Swiatnicki MR, Andrechek ER. Histological subtypes of mouse mammary tumors reveal conserved relationships to human cancers. PLoS Genet. 2018;14:e1007135.
Hennessy BT, Gonzalez-Angulo AM, Stemke-Hale K, Gilcrease MZ, Krishnamurthy S, Lee JS, et al. Characterization of a naturally occurring breast cancer subset enriched in epithelial-to-mesenchymal transition and stem cell characteristics. Cancer Res. 2009;69:4116–24.
Barlow J, Yandell D, Weaver D, Casey T, Plaut K. Higher stromal expression of transforming growth factor-beta type II receptors is associated with poorer prognosis breast tumors. Breast Cancer Res Treat. 2003;79:149–59.
Bhola NE, Balko JM, Dugger TC, Kuba MG, Sanchez V, Sanders M, et al. TGF-beta inhibition enhances chemotherapy action against triple-negative breast cancer. J Clin Invest. 2013;123:1348–58.
Gobbi H, Dupont WD, Simpson JF, Plummer WD Jr., Schuyler PA, Olson SJ, et al. Transforming growth factor-beta and breast cancer risk in women with mammary epithelial hyperplasia. J Natl Cancer Inst. 1999;91:2096–101.
Gobbi H, Arteaga CL, Jensen RA, Simpson JF, Dupont WD, Olson SJ, et al. Loss of expression of transforming growth factor beta type II receptor correlates with high tumour grade in human breast in-situ and invasive carcinomas. Histopathology 2000;36:168–77.
Busch S, Acar A, Magnusson Y, Gregersson P, Ryden L, Landberg G. TGF-beta receptor type-2 expression in cancer-associated fibroblasts regulates breast cancer cell growth and survival and is a prognostic marker in pre-menopausal breast cancer. Oncogene 2015;34:27–38.
De Summa S, Pinto R, Sambiasi D, Petriella D, Paradiso V, Paradiso A, et al. BRCAness: a deeper insight into basal-like breast tumors. Ann Oncol. 2013;24:viii13–viii21.
Lord CJ, Tutt AN, Ashworth A. Synthetic lethality and cancer therapy: lessons learned from the development of PARP inhibitors. Annu Rev Med. 2015;66:455–70.
Maxwell KN, Domchek SM. Cancer treatment according to BRCA1 and BRCA2 mutations. Nat Rev Clin Oncol. 2012;9:520–8.
Jonsson G, Staaf J, Vallon-Christersson J, Ringner M, Gruvberger-Saal SK, Saal LH, et al. The retinoblastoma gene undergoes rearrangements in BRCA1-deficient basal-like breast cancer. Cancer Res. 2012;72:4028–36.
Stefansson OA, Jonasson JG, Olafsdottir K, Hilmarsdottir H, Olafsdottir G, Esteller M, et al. CpG island hypermethylation of BRCA1 and loss of pRb as co-occurring events in basal/triple-negative breast cancer. Epigenetics 2011;6:638–49.
Herschkowitz JI, He X, Fan C, Perou CM. The functional loss of the retinoblastoma tumour suppressor is a common event in basal-like and luminal B breast carcinomas. Breast Cancer Res. 2008;10:R75.
Drost RM, Jonkers J. Preclinical mouse models for BRCA1-associated breast cancer. Br J Cancer. 2009;101:1651–7.
Jiang Z, Deng T, Jones R, Li H, Herschkowitz JI, Liu JC, et al. Rb deletion in mouse mammary progenitors induces luminal-B or basal-like/EMT tumor subtypes depending on p53 status. J Clin Invest. 2010;120:3296–309.
Chang CJ, Chao CH, Xia W, Yang JY, Xiong Y, Li CW, et al. p53 regulates epithelial-mesenchymal transition and stem cell properties through modulating miRNAs. Nat Cell Biol. 2011;13:317–23.
Herschkowitz JI, Simin K, Weigman VJ, Mikaelian I, Usary J, Hu Z, et al. Identification of conserved gene expression features between murine mammary carcinoma models and human breast tumors. Genome Biol. 2007;8:R76.
Wang H, Bierie B, Li AG, Pathania S, Toomire K, Dimitrov SD, et al. BRCA1/FANCD2/BRG1-Driven DNA Repair Stabilizes the Differentiation State of Human Mammary Epithelial Cells. Mol Cell. 2016;63:277–92.
Wang H, Xiang D, Liu B, He A, Randle HJ, Zhang KX, et al. Inadequate DNA Damage Repair Promotes Mammary Transdifferentiation, Leading to BRCA1 Breast. Cancer Cell 2019;178:135–51 e19.
Pei XH, Bai F, Smith MD, Usary J, Fan C, Pai SY, et al. CDK inhibitor p18(INK4c) is a downstream target of GATA3 and restrains mammary luminal progenitor cell proliferation and tumorigenesis. Cancer Cell. 2009;15:389–401.
Goulding H, Pinder S, Cannon P, Pearson D, Nicholson R, Snead D, et al. A new immunohistochemical antibody for the assessment of estrogen receptor status on routine formalin-fixed tissue samples. Hum Pathol. 1995;26:291–4.
Reich M, Liefeld T, Gould J, Lerner J, Tamayo P, Mesirov JP. GenePattern 2.0. Nat Genet. 2006;38:500–1.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Verrecchia F, Chu ML, Mauviel A. Identification of novel TGF-beta /Smad gene targets in dermal fibroblasts using a combined cDNA microarray/promoter transactivation approach. J Biol Chem. 2001;276:17058–62.
Schon M, Wienrich BG, Kneitz S, Sennefelder H, Amschler K, Vohringer V, et al. KINK-1, a novel small-molecule inhibitor of IKKbeta, and the susceptibility of melanoma cells to antitumoral treatment. J Natl Cancer Inst. 2008;100:862–75.
Bai F, Liu S, Liu X, Hollern DP, Scott A, Wang C, et al. PDGFRbeta is an essential therapeutic target for BRCA1-deficient mammary tumors. Breast Cancer Res. 2021;23:10.
Acknowledgements
We thank Drs. Beverly Koller, Chuxia Deng, and Lothar Hennighausen for Brca1 mutant and MMTV-cre mice, Yue Xiong for discussion, Charles M. Perou for microarray analysis and discussion, Emely Pimentel and Alexandria Scott for technical support, the FACS core facility at University of Miami and Shenzhen University for cell sorting, the DVR core facility for animal husbandry. Chuying Wang thanks Xi’an Jiaotong University for financial support. This study was supported by the Guangdong Provincial Science and Technology Program (2019B030301009), National Natural Science Foundation of China (81972637), High-level university phase 2 construction funding from Shenzhen University (860-00000210), Natural Science Foundation of Shenzhen City (JCYJ20190808115603580 and JCYJ20190808165803558), Natural Science Foundation of Guangdong Province (2019A1515011343 and 2021A1515011145), and the Bankhead-Coley Cancer Research grant (TBC07).
Author information
Authors and Affiliations
Contributions
FB and XHP designed the research studies. FB, CW, XL, SL, CL, SR, and XHP conducted experiments and analyzed data. DH, CF, and JIH carried out microarray analysis and breast cancer dataset analysis. WGZ provided technical and material support. FB, DH, and XHP wrote the manuscript. XHP supervised the project. All authors made comments on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics Statement
The Institutional Animal Care and Use Committee at the University of Miami and Shenzhen University approved all animal procedures.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Dr Satoshi Inoue
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bai, F., Wang, C., Liu, X. et al. Loss of function of BRCA1 promotes EMT in mammary tumors through activation of TGFβR2 signaling pathway. Cell Death Dis 13, 195 (2022). https://doi.org/10.1038/s41419-022-04646-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-022-04646-7
- Springer Nature Limited
This article is cited by
-
PALB2-mutated human mammary cells display a broad spectrum of morphological and functional abnormalities induced by increased TGFβ signaling
Cellular and Molecular Life Sciences (2024)
-
Loss of function of GATA3 regulates FRA1 and c-FOS to activate EMT and promote mammary tumorigenesis and metastasis
Cell Death & Disease (2023)
-
Therapeutic targeting approach on epithelial-mesenchymal plasticity to combat cancer metastasis
Medical Oncology (2023)
-
miR-17-5p slows progression of hepatocellular carcinoma by downregulating TGFβR2
Clinical and Translational Oncology (2023)
-
Epithelial-mesenchymal transition in cancer stemness and heterogeneity: updated
Medical Oncology (2022)