Abstract
Background
Mismatch repair (MMR) deficiency is the hallmark of tumours from Lynch syndrome (LS), sporadic MLH1 hypermethylated and Lynch-like syndrome (LLS), but there is a lack of understanding of the variability in their mutational profiles based on clinical phenotypes. The aim of this study was to perform a molecular characterisation to identify novel features that can impact tumour behaviour and clinical management.
Methods
We tested 105 MMR-deficient colorectal cancer tumours (25 LS, 35 LLS and 45 sporadic) for global exome microsatellite instability, cancer mutational signatures, mutational spectrum and neoepitope load.
Results
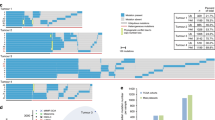
Fifty-three percent of tumours showed high contribution of MMR-deficient mutational signatures, high level of global exome microsatellite instability, loss of MLH1/PMS2 protein expression and included sporadic tumours. Thirty-one percent of tumours showed weaker features of MMR deficiency, 62% lost MSH2/MSH6 expression and included 60% of LS and 44% of LLS tumours. Remarkably, 9% of all tumours lacked global exome microsatellite instability. Lastly, HLA-B07:02 could be triggering the neoantigen presentation in tumours that show the strongest contribution of MMR-deficient tumours.
Conclusions
Next-generation sequencing approaches allow for a granular molecular characterisation of MMR-deficient tumours, which can be essential to properly diagnose and treat patients with these tumours in the setting of personalised medicine.


Similar content being viewed by others
Data availability
The datasets generated and/or analysed during the current study are not publicly available yet but will be following NCI data-sharing policies. However, datasets could be available from the corresponding author on reasonable request.
References
Plotz G, Piiper A, Wormek M, Zeuzem S, Raedle J. Analysis of the human MutLalpha.MutSalpha complex. Biochem Biophys Res Commun. 2006;340:852–9.
Xicola RM, Llor X, Pons E, Castells A, Alenda C, Pinol V, et al. Performance of different microsatellite marker panels for detection of mismatch repair-deficient colorectal tumors. J Natl Cancer Inst. 2007;99:244–52.
Hampel H, Frankel WL, Martin E, Arnold M, Khanduja K, Kuebler P, et al. Screening for the Lynch syndrome (hereditary nonpolyposis colorectal cancer). N Engl J Med. 2005;352:1851–60.
Llor X. When should we suspect hereditary colorectal cancer syndrome? Clin Gastroenterol Hepatol. 2012;10:363–7.
Kane MF, Loda M, Gaida GM, Lipman J, Mishra R, Goldman H, et al. Methylation of the hMLH1 promoter correlates with lack of expression of hMLH1 in sporadic colon tumors and mismatch repair-defective human tumor cell lines. Cancer Res. 1997;57:808–11.
Bessa X, Balleste B, Andreu M, Castells A, Bellosillo B, Balaguer F, et al. A prospective, multicenter, population-based study of BRAF mutational analysis for Lynch syndrome screening. Clin Gastroenterol Hepatol. 2008;6:206–14.
Rodriguez-Soler M, Perez-Carbonell L, Guarinos C, Zapater P, Castillejo A, Barbera VM, et al. Risk of cancer in cases of suspected lynch syndrome without germline mutation. Gastroenterology. 2013;144:926–32 e1.
Geurts-Giele WR, Leenen CH, Dubbink HJ, Meijssen IC, Post E, Sleddens HF, et al. Somatic aberrations of mismatch repair genes as a cause of microsatellite-unstable cancers. J Pathol. 2014;234:548–59.
Mensenkamp AR, Vogelaar IP, van Zelst-Stams WA, Goossens M, Ouchene H, Hendriks-Cornelissen SJ, et al. Somatic mutations in MLH1 and MSH2 are a frequent cause of mismatch-repair deficiency in Lynch syndrome-like tumors. Gastroenterology. 2014;146:643–6.e8.
Sourrouille I, Coulet F, Lefevre JH, Colas C, Eyries M, Svrcek M, et al. Somatic mosaicism and double somatic hits can lead to MSI colorectal tumors. Fam Cancer. 2013;12:27–33.
Jansen AM, van Wezel T, van den Akker BE, Ventayol Garcia M, Ruano D, Tops CM, et al. Combined mismatch repair and POLE/POLD1 defects explain unresolved suspected Lynch syndrome cancers. Eur J Hum Genet. 2016;24:1089–92.
Alexandrov LB, Kim J, Haradhvala NJ, Huang MN, Tian Ng AW, Wu Y, et al. The repertoire of mutational signatures in human cancer. Nature. 2020;578:94–101.
Alexandrov LB, Nik-Zainal S, Wedge DC, Aparicio SA, Behjati S, Biankin AV, et al. Signatures of mutational processes in human cancer. Nature. 2013;500:415–21.
Boon T, Cerottini JC, Van den Eynde B, van der Bruggen P, Van, Pel A. Tumor antigens recognized by T lymphocytes. Annu Rev Immunol. 1994;12:337–65.
Linnebacher M, Gebert J, Rudy W, Woerner S, Yuan YP, Bork P, et al. Frameshift peptide-derived T-cell epitopes: a source of novel tumor-specific antigens. Int J Cancer. 2001;93:6–11.
Le DT, Durham JN, Smith KN, Wang H, Bartlett BR, Aulakh LK, et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science. 2017;357:409–13.
Overman MJ, Lonardi S, Wong KYM, Lenz HJ, Gelsomino F, Aglietta M, et al. Durable clinical benefit with nivolumab plus ipilimumab in DNA mismatch repair-deficient/microsatellite instability-high metastatic colorectal cancer. J Clin Oncol. 2018;36:773–9.
Fabrizio DA, George TJ Jr., Dunne RF, Frampton G, Sun J, Gowen K, et al. Beyond microsatellite testing: assessment of tumor mutational burden identifies subsets of colorectal cancer who may respond to immune checkpoint inhibition. J Gastrointest Oncol. 2018;9:610–7.
Xicola RM, Gagnon M, Clark JR, Carroll T, Gao W, Fernandez C, et al. Excess of proximal microsatellite-stable colorectal cancer in African Americans from a multiethnic study. Clin Cancer Res. 2014;20:4962–70.
Pico MD, Castillejo A, Murcia O, Giner-Calabuig M, Alustiza M, Sanchez A, et al. Clinical and pathological characterization of Lynch-like syndrome. Clin Gastroenterol Hepatol. 2020;18:368–74.e1.
Grossman RL, Heath AP, Ferretti V, Varmus HE, Lowy DR, Kibbe WA, et al. Toward a shared vision for cancer genomic data. N Engl J Med. 2016;375:1109–12.
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol. 2013;31:213–9.
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164.
Shyr C, Tarailo-Graovac M, Gottlieb M, Lee JJ, van Karnebeek C, Wasserman WW. FLAGS, frequently mutated genes in public exomes. BMC Med Genomics. 2014;7:64.
Kautto EA, Bonneville R, Miya J, Yu L, Krook MA, Reeser JW, et al. Performance evaluation for rapid detection of pan-cancer microsatellite instability with MANTIS. Oncotarget. 2017;8:7452–63.
Bonneville R, Krook MA, Kautto EA, Miya J, Wing MR, Chen HZ, et al. Landscape of microsatellite instability across 39 cancer types. JCO Precision Oncology 2017;1:15.
Huang MN, McPherson JR, Cutcutache I, Teh BT, Tan P, Rozen SG. MSIseq: Software for assessing microsatellite instability from catalogs of somatic mutations. Sci Rep. 2015;5:13321.
Blokzijl F, Janssen R, van Boxtel R, Cuppen E. MutationalPatterns: comprehensive genome-wide analysis of mutational processes. Genome Med. 2018;10:33.
Gaujoux R, Seoighe C. A flexible R package for nonnegative matrix factorization. BMC Bioinformatics. 2010;11:367.
Woo J, Winterhoff BJ, Starr TK, Aliferis C, Wang J. De novo prediction of cell-type complexity in single-cell RNA-seq and tumor microenvironments. Life Sci Alliance. 2019;2:e201900443.
Gu Z, Eils R, Schlesner M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics. 2016;32:2847–9.
Sakai R, Winand R, Verbeiren T, Moere AV, Aerts J. dendsort: modular leaf ordering methods for dendrogram representations in R. F1000Res. 2014;3:177.
Schenck RO, Lakatos E, Gatenbee C, Graham TA, Anderson ARA. NeoPredPipe: high-throughput neoantigen prediction and recognition potential pipeline. BMC Bioinformatics. 2019;20:264.
Nielsen M, Andreatta M. NetMHCpan-3.0; improved prediction of binding to MHC class I molecules integrating information from multiple receptor and peptide length datasets. Genome Med. 2016;8:33.
Shukla SA, Rooney MS, Rajasagi M, Tiao G, Dixon PM, Lawrence MS, et al. Comprehensive analysis of cancer-associated somatic mutations in class I HLA genes. Nat Biotechnol. 2015;33:1152–8.
Chen C, Li Z, Huang H, Suzek BE, Wu CH, UniProt C. A fast Peptide Match service for UniProt Knowledgebase. Bioinformatics. 2013;29:2808–9.
Luksza M, Riaz N, Makarov V, Balachandran VP, Hellmann MD, Solovyov A, et al. A neoantigen fitness model predicts tumour response to checkpoint blockade immunotherapy. Nature. 2017;551:517–20.
Vita R, Overton JA, Greenbaum JA, Ponomarenko J, Clark JD, Cantrell JR, et al. The immune epitope database (IEDB) 3.0. Nucleic Acids Res. 2015;43:D405–D412.
Team RC. R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2019.
Bailey MH, Tokheim C, Porta-Pardo E, Sengupta S, Bertrand D, Weerasinghe A, et al. Comprehensive characterization of cancer driver genes and mutations. Cell. 2018;173:371–85 e18.
Network TCGA. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–7.
Latham A, Srinivasan P, Kemel Y, Shia J, Bandlamudi C, Mandelker D, et al. Microsatellite instability is associated with the presence of Lynch syndrome pan-cancer. J Clin Oncol. 2019;37:286–95.
Drost J, van Boxtel R, Blokzijl F, Mizutani T, Sasaki N, Sasselli V, et al. Use of CRISPR-modified human stem cell organoids to study the origin of mutational signatures in cancer. Science. 2017;358:234–8.
Zou X, Koh GCC, Nanda AS, Degasperi A, Urgo K, Roumeliotis TI, et al. A systematic CRISPR screen defines mutational mechanisms underpinning signatures caused by replication errors and endogenous DNA damage. Nat Cancer. 2021;2:643–57.
Buckley AR, Ideker T, Carter H, Harismendy O, Schork NJ. Exome-wide analysis of bi-allelic alterations identifies a Lynch phenotype in The Cancer Genome Atlas. Genome Med. 2018;10:69.
Xicola RM, Clark JR, Carroll T, Alvikas J, Marwaha P, Regan MR, et al. Implication of DNA repair genes in Lynch-like syndrome. Fam Cancer. 2019;18:331–42.
Xavier A, Olsen MF, Lavik LA, Johansen J, Singh AK, Sjursen W, et al. Comprehensive mismatch repair gene panel identifies variants in patients with Lynch-like syndrome. Mol Genet Genom Med. 2019;7:e850.
Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science. 2015;348:124–8.
Mandal R, Samstein RM, Lee KW, Havel JJ, Wang H, Krishna C, et al. Genetic diversity of tumors with mismatch repair deficiency influences anti-PD-1 immunotherapy response. Science. 2019;364:485–91.
Chowell D, Morris LGT, Grigg CM, Weber JK, Samstein RM, Makarov V, et al. Patient HLA class I genotype influences cancer response to checkpoint blockade immunotherapy. Science. 2018;359:582–7.
Acknowledgements
The results shown here are in part based on data generated by the TCGA Research Network: https://www.cancer.gov/tcga.
Funding
This work was supported by grants from the National Cancer Institute (1K01CA204431-01A1 RMX), the Prevent Cancer Foundation (RMX), Colorectal Cancer Alliance’s Chris4Life grant (RMX), pre-doctoral grant from Conselleria d’Educació de la Generalitat Valenciana. VALi+d. EXP ACIF/2010/018, ACIF/2016/002 (MGC), Instituto de Salud Carlos III PI17/01756 (RJ), Asociación Española de Gastroenterología. Beca Tamarite 2017 (RJ) and the Donaldson Foundation (SS and CU).
Author information
Authors and Affiliations
Contributions
Conceptualisation (MGC, XL, RMX); Data curation (MGC, SDL, JW, MDP, TF, CU, MH, MCP, IS); Formal analysis (MGC, RMX); Funding acquisition (RJ, JS, XL, RMX); Investigation (MGC, SDL, JW, JG, CA); Project administration (RMX); Resources (TF, CU, SS, MH, MCP, IS, EMS, NAE, JS, MA, MDP, RJ, JR, SPO, AOH); Supervision (RMX); Validation (MGC, SDL); Visualisation (MGC, JW, RMX); Writing—original draft (MGC, XL, RMX); Writing—review and editing (MGC, CU, SS, ES, JG, MC, RJ, NAE, XL, RMX).
Corresponding author
Ethics declarations
Competing interests
SS is a consultant for Myriad Genetics and DC Health Technologies and has rights to an inventor portion of the licensing revenue from PREMM5. The remaining authors declare no competing interests.
Ethics approval and consent to participate
Patients were recruited and consented in their original institutions under projects approved by institutional human research review committees. At Yale University the project was approved by the Biomedical Institutional Review Board. All data were shared in a de-identified manner to unlink any patient’s personal identifiers from their samples. The study was performed in accordance with the Declaration of Helsinki.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Giner-Calabuig, M., De Leon, S., Wang, J. et al. Mutational signature profiling classifies subtypes of clinically different mismatch-repair-deficient tumours with a differential immunogenic response potential. Br J Cancer 126, 1595–1603 (2022). https://doi.org/10.1038/s41416-022-01754-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-022-01754-1
- Springer Nature Limited