Abstract
Leukaemia is caused by the clonal evolution of a cell that accumulates mutations/genomic rearrangements, allowing unrestrained cell growth. However, recent identification of leukaemic mutations in the blood cells of healthy individuals revealed that additional events are required to expand the mutated clones for overt leukaemia. Here, we assessed the functional consequences of deleting the Fanconi anaemia A (Fanca) gene, which encodes a DNA damage response protein, in Spi1 transgenic mice that develop preleukaemic syndrome. FANCA loss increases SPI1-associated disease penetrance and leukaemic progression without increasing the global mutation load of leukaemic clones. However, a high frequency of leukaemic FANCA-depleted cells display heterozygous activating mutations in known oncogenes, such as Kit or Nras, also identified but at low frequency in FANCA-WT mice with preleukaemic syndrome, indicating that FANCA counteracts the emergence of oncogene mutated leukaemic cells. A unique transcriptional signature is associated with the leukaemic status of FANCA-depleted cells, leading to activation of MDM4, NOTCH and Wnt/β-catenin pathways. We show that NOTCH signalling improves the proliferation capacity of FANCA-deficient leukaemic cells. Collectively, our observations indicate that loss of the FANC pathway, known to control genetic instability, fosters the expansion of leukaemic cells carrying oncogenic mutations rather than mutation formation. FANCA loss may contribute to this leukaemogenic progression by reprogramming transcriptomic landscape of the cells.
Similar content being viewed by others
Introduction
Genetic instability, a key characteristic of cancer [1], is considered a major cell autonomous force in both cancer initiation and progression. Indeed, an increased mutational landscape correlates with cancer progression, as supported by three major facts: (a) the majority of solid tumours and leukaemia consistently exhibit genetic instability, as revealed by both cytogenetic and sequencing data analyses [2]; (b) the high risk of cancer incurred by patients bearing inactivating germline mutations in genes encoding DNA damage response (DDR) proteins, which maintain genome stability [3]; and, inversely, (c) the frequent presence of inactivating mutations in DDR protein-encoding genes in sporadic tumours [4].
However, the mutational signature repertoire and the mutational load associated with cancer are clearly tumour-dependent and, comparatively to solid tumours, scarce in leukaemia [2]. Moreover, recently published studies revealed an accumulation of somatic mutations in cancer-associated driver genes in healthy cells that maintain their physiological functions [5]. For example, a pattern of somatic mutations found in haematological cancer-associated genes, including DNMT3A, TET2, ASXL1, TP53 and its regulator PPM1D, has been described in healthy individuals without clinical haematologic abnormalities [6]. These observations raised questions about what additional factors and events, in addition to genetic instability, contribute to leukaemic progression.
SPI1/PU.1 is a master TF involved in normal and stressed haematopoiesis. SPI1 behaves either as a tumour suppressor [7, 8] or an oncogene [9,10,11]. Alteration of SPI1 expression in a mouse model has been identified as a founding event in myeloid transformation, triggering cellular and molecular alterations that are consistent with preleukaemic disorders [12,13,14]. SPI1-overexpressing mice develop such a preleukemic disorder from erythroid origin [14]. SPI1 constitutive expression in mice causes haematological neoplasia characterized by anaemia due to the blockage of erythroid differentiation and splenomegaly due to expansion of blasts whose expansion is still controlled by growth factors [14]. Whereas 60% of mice develop preleukaemic syndrome, they rarely progress spontaneously into leukaemia (identified in less than 10% of the offspring). However, leukaemic progression can be mediated by additional mutations in the c-Kit gene encoding the stem cell factor (SCF) receptor [15]. Thus, transgenic Spi1-overexpressing (TgSpi1) mice represent a preleukaemic state towards the development of leukaemia. Similarly, a block of erythroid differentiation due to high expression of transcriptional regulators has recently been described in over 25% of acute human erythroleukaemia (AEL) cases [16]. The identification of secondary mutations in genes encoding DDR proteins in murine models of AEL [17], including those that code for the proteins of the Fanconi anaemia pathway (FANC), supports that downstream alteration in key oncogenes and abnormalities in DDR plays a role in leukaemic progression.
FANC proteins constitute a pathway embedded in the DDR network and are involved in DNA repair and replication rescue, ensuring chromosomal integrity [18, 19]. Their genetic loss of function leads to Fanconi anaemia (FA), a human bone marrow failure (BMF) syndrome associated with DDR failure, genetic instability, abnormalities in ribosome biogenesis and a proinflammatory and prosenescent phenotype [20, 21]. Counterintuitively, considering hypoproliferative FA cells and clinical phenotypes, FA patients are at high risk for myelodysplastic syndrome (MDS), acute myeloid leukaemia (AML) and solid cancers [22]. FA is due to inactivating biallelic mutations in over 20 FANC genes, some of which have also been involved in other cancer predisposition syndromes [23, 24]. Surprisingly, patients with mutations in the different genes that affect the functionality of the FANC pathway do not have the same spectrum of cancers [23, 25, 26]. Moreover, FANC-knockout (KO) mouse models generally fail to spontaneously develop leukaemia or other cancers, but they present minor BMF defects, infertility and DNA damage hypersensitivity, attesting to the presence of major DDR alterations [27,28,29,30]. Thus, even if myelodysplasia or AML in FA patients appears to be associated with some specific genomic abnormalities [31], genetic instability seems to be insufficient per se for overt leukaemic development in FA.
To provide new insights into additional factors or events that, in addition to genetic instability, contribute to leukaemic progression, we combined alternative genetic preleukaemic settings provided by the SPI1-overexpressing mice and the knockout of the Fanca gene. Since preleukaemic erythroblastic cells expressing high levels of SPI1 exhibit replicative stress [32, 33], a deficiency in DNA repair and/or replicative stress responding proteins could increase the genetic instability and/or tumorigenic progression of SPI1-overexpressing preleukaemic blasts, revealing gene products and pathways involved in allowing leukaemia-initiated cells to progress. We show that loss of function of FANCA and abnormally high expression of SPI1 synergize during leukaemogenesis. Depletion of FANCA in TgSpi1 mice leads to progression from preleukaemia to leukaemia without increasing mutation load. In contrast, selection and/or expansion of cells with oncogenic mutations are promoted and associated with a peculiar transcriptional programme in leukaemic cells.
Results
FANCA deficiency exacerbates TgSpi1-associated haematologic disorder
We crossed Fanca+/− (FA+/−) mice [28] with transgenic mice constitutively expressing a Spi1 transgene (TgSpi1Fanca+/+) [14] and backcrossed the first generation (F1) of TgSpi1FA+/− mice with Fanca+/− mice to obtain TgSpi1FA−/− mice (Fig. 1A). The lack of Fanca expression was verified by RT-qPCR, and the overexpression of SPI1 was confirmed by immunohistochemistry (Supplementary Fig. S1A, B).
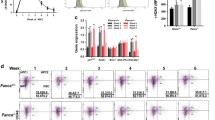
A To generate mice harbouring the Fanca deletion and Spi1 transgene, homozygous Spi1 transgenic male mice (TgSpi1) were crossed with Fanca-deleted heterozygous females. From this first cross (F1), TgSpi1 and Fanca-deleted heterozygous males (TgSpi1Fanca+/−) were then used for breeding with Fanca-deleted heterozygous females (Fanca+/−), and the resulting TgSpi1 and Fanca-deleted homozygous (TgSpi1FA−/−) mice were identified by genotyping. Mice in grey boxes were not used. Fanca-deleted mice are further indicated as FA−/− or FA+/−. B Kaplan‒Meier curve of the survival of mice bearing TgSpi1FA−/− (N = 39), TgSpi1FA+/− (N = 81) or allelic controls (FA+/+, N = 12; FA−/−, N = 12; TgSpi1FA+/+, N = 57). ****P < 0.0001, Mantel‒Cox test. Haemoglobin concentration (C), haematocrit level (D) and cell count (E) of peripheral blood from moribund TgSpi1FA−/− and TgSpi1FA+/+ mice and from 3- to 4-month-old FA+/+ and FA−/− mice. Histograms indicate the mean ± SEM from 6 to 13 individual mice according to the groups. Statistical analysis was carried out using the Mann‒Whitney test. F Histograms indicate the mean ± SEM of spleen weight of moribund TgSpi1FA−/− and TgSpi1FA+/+ mice and from 3- to 4-month-old FA+/+ and FA−/− mice from 4 to 15 individual mice according to the groups. Statistical analysis was carried out using the Mann‒Whitney test. G Representative images of the spleens of the mice with the 4 genotypes. Scale bar, 1 cm. H Haematoxylin and eosin-stained splenic sections of moribund TgSpi1FA−/− and TgSpi1FA+/+ mice or 3- to 4-month-old healthy FA+/+ and FA−/− mice. Magnification, ×200.
All mice appeared healthy at birth without congenital malformations or growth problems. At later ages, TgSpi1FA+/+, TgSpi1FA+/− and TgSpi1FA−/− mice developed anaemia and splenomegaly, leading to their death. However, all of the TgSpi1FA−/− mice died within 10 months after birth, while only 67% of the TgSpi1FA+/+ mice died within this time frame (Fig. 1B and Supplementary Fig. S1C). The median survival of 67 days for the TgSpi1FA−/− mice and 115 days for the TgSpi1FA+/+ mice demonstrated acceleration of disease onset (Fig. 1B). TgSpi1FA+/− mice present with intermediate behaviour, with a median survival of 105 days, and 20% of the animals were still alive at 10 months of age. None of the wild-type Fanca+/+ (WT, FA+/+) or Fanca−/− (FA−/−) mice developed a disease, indicating that the pathology requires the unrestrained expression of SPI1. At death, TgSpi1FA+/+ and TgSpi1FA−/− mice showed a similar severity of anaemia with a reduction in both the haemoglobin and haematocrit levels (Fig. 1C, D). TgSpi1FA+/+ and TgSpi1FA−/− mice displayed a high number of total white blood cells (>100 × 103/μL) (Fig. 1E) and comparable splenomegaly (Fig. 1F, G) with disrupted splenic architecture (Fig. 1H). All these parameters indicate that their death occurs at a similar level of disease progression.
Taken together, our data demonstrate that FANCA loss of function accelerates the progression of the haemopathy associated with the unrestrained expression of SPI1 and allows for full disease penetrance.
Lack of FANCA does not modify the lineage specificity of leukaemic cells associated with SPI1 overexpression
To shed light on how loss of FANCA function led to the observed acceleration and full penetrance of SPI1-associated disease, we searched for the haematopoietic populations contributing to the disease. At death, the proportions of CD4+CD8- and CD4-CD8+ T lymphoid or CD11b+ myeloid cells in the bone marrow (BM) were similar in all genotypes, while B lymphoid, CD19+B220+, cells were equally reduced in TgSpi1FA+/+ and TgSpi1FA−/− mice compared to non-TgSpi1, FA+/+ and FA−/− mice (Supplementary Fig. S2).
Subsequently, we profiled erythroid subpopulations according to CD71 and Ter119 cell-surface markers and the forward scatter (FSC) parameter [34, 35]. Flow cytometry analysis identified the following six cellular subpopulations: CD71−Ter119− (I); CD71+Ter119− (II, colony-forming unit erythroid, CFU-E); CD71+Ter119Med (III, pro-erythroblasts, Pro-E); and the three Ter119High subpopulations (IV) that are CD71+FSCHigh or basophilic, CD71+FSCLow or polychromatic, and CD71-FSCLow or orthochromatic cells (Fig. 2A, B). Compared to FA+/+ and FA−/− mice, which show similar profiles, TgSpi1FA+/+ and TgSpi1FA−/− mice displayed an increased proportion of CFU-E progenitors in their BM to more than 50% of the total cell count. This is at the expense of basophilic, polychromatic and orthochromatic erythroblasts, indicating erythroid differentiation blockage consistent with the observed anaemia (Fig. 2B, C). Altered erythroid differentiation was also evident in the spleen, with a clear accumulation of CFU-E progenitors in TgSpi1 mice (Fig. 2C). Similar populations were found between TgSpi1FA+/+ and TgSpi1FA−/− mice.
A Schematic diagram representing erythroid differentiation with the specific phenotypic markers used. B Representative profiles of cells incubated with antibodies against Ter119 and CD71 and analysed by flow cytometry. C Histograms represent the mean % of each population for mice analysed (from 4 to 8 mice) according to Ter119 and CD71 markers and basophilic, polychromatic and acidophilic cells relative to 100% of bone marrow or spleen cells as described in A and B. Statistical analysis was carried out using the Mann‒Whitney test. Three types of comparisons were performed: FA−/− to TgSpi1FA−/−; FA+/+ to TgSpi1FA+/+ and TgSpi1FA−/− to TgSpi1FA+/+. For better visualization, only one * was written for all P values (P < 0.05), even if more highly significant.
In conclusion, the proliferating blasts had an erythroid lineage origin in both TgSpi1FA+/+ and TgSpi1FA−/− mice. Moreover, accelerated death and full penetrance of the TgSpi1-associated disease due to FANCA loss of function were not caused by additional alterations of other lineages such as myeloid, B or T lymphoid cells.
FANCA deficiency promotes tumorigenicity associated with EPO independence
It was previously shown that cells isolated from the bone marrow and spleen of sick TgSpi1 mice required the erythroid growth factor erythropoietin (EPO) to be maintained in culture and did not give rise to tumour after subcutaneous engraftment [14]. Accordingly, EPO starvation induced growth arrest and death of blasts derived from 83% of TgSpi1FA+/+ mice generated in this study (Fig. 3A, left panel and 3B). In contrast, the blasts derived from only 23% of TgSpi1FA−/− mice required EPO for survival and proliferation (Fig. 3A, right panel and 3B). In addition, EPO-independent cells formed in situ tumours within three weeks of their subcutaneous injection into immunodeficient mice demonstrating their ability for autonomous proliferation (Fig. 3B and Supplementary Fig. S3A). None of the EPO-dependent cells gave rise to tumours. Additionally, blasts were intravenously injected into immunodeficient mice where they were exposed to endogenous EPO. Accordingly, EPO-dependent and EPO-independent cells were able to invade the bone marrow and spleen. However, EPO-independent blasts from the TgSpi1FA−/ − mice displayed a strong proliferation advantage as deduced from the higher fraction of blasts in the bone marrow and spleen compared to EPO-dependent cells (Supplementary Fig. S3B). EPO-dependent cells that do not give rise to subcutaneous tumours, with no autonomous proliferation, are considered as preleukaemic cells.
A Primary cells isolated from the spleens of sick TgSpi1FA+/+ and TgSpi1FA−/− mice with splenomegaly were cultured with (+EPO, full line) or without EPO (−EPO, dashed line). Living cells were counted on the indicated days using the trypan blue exclusion test. Data are the mean ± SEM of a minimum of 3 independent experiments. B EPO requirement and in situ subcutaneous secondary tumour in nude mice from TgSpi1FA+/+ and TgSpi1FA−/− blasts. D: EPO dependent; I: EPO independent; nd: not determined. Filled boxes indicate that primary transforming cells formed secondary tumours in nude mice; empty boxes indicate that no secondary tumours were detected. Histograms represent the percentage of sick mice whose spleen was invaded by EPO dependent or EPO-independent cells among TgSpi1FA+/+ or TgSpi1FA−/− mice. The same is shown for the ability to develop subcutaneous in situ secondary tumours. Statistical analysis was carried out using Chi-square test. * indicates P < 0.05. C Representative immunoblots of whole-cell extracts of TgSpi1FA+/+ and TgSpi1FA−/− cells coupled with anti-pSTAT5, total STAT5, pAKT, total AKT, pERK1/2 and total ERK1/2 antibodies. Cells were cultured in the presence (+) or absence (−) of EPO for 4 h. Histograms represent the mean of the signal ratio of phosphorylated to total protein in cells deprived of EPO relative to their respective control, cells cultured in the presence of EPO from 3 independent mice. Statistical analysis was carried out using unpaired Student t-test. * indicates P < 0.05; **P < 0.01; ***P < 0,001.
To sustain blast survival and proliferation, EPO initiates several signalling pathways via the activation of diverse effectors, including STAT5, AKT1 and ERK1/2 [36]. Accordingly, TgSpi1FA+/+ preleukaemic cells cultured in the presence of EPO presented STAT5, AKT1 and ERK1/2 phosphorylation, an indicator of their activation, which was systematically reduced following EPO starvation (Fig. 3C, #445, #138, #479). In contrast, in TgSpi1FA−/− blasts whose proliferation was independent of EPO, the activation of the AKT1 and ERK1/2 signalling pathways was independent of the presence of EPO (Fig. 3C, #39, #26, #28). Interestingly, the consequence of EPO deprivation on STAT5 activation differs according to the TgSpi1FA−/− mice. Cells from #26 mice still presented STAT5 phosphorylation in the absence of EPO, while those from #28 and #39 mice were not able to maintain STAT5 phosphorylation, indicating that the absence of FANCA was not sufficient for STAT5 constitutive activation. Interestingly, shRNA-mediated Fanca downregulation in EPO-dependent TgSpi1FA+/+ cell lines (#138, #445, #479) was unable to confer EPO independence in vitro 9 days post-infection (Supplementary Fig. S3C, D).
In conclusion, our results show that the absence of FANCA protein contributes to the emergence of blasts that do not require EPO to survive and/or proliferate. However, the sole absence of FANCA protein in a clone is not sufficient to confer EPO independence in a short time, revealing a complex mechanism likely associated with the presence of additional epi- or genetic events.
FANCA loss of function favours the emergence of leukaemic TgSpi1 clones with oncogenic mutations
To gain insight into the causes of EPO independence and tumorigenicity associated with loss of FANCA function, we performed whole-exome sequencing (WES) analysis of blasts isolated from the spleen of 5 TgSpi1FA+/+ and 9 TgSpi1FA−/− mice with anaemia and splenomegaly.
WES analysis demonstrated that the number of small mutations (<200 bp) with a variant allele frequency (VAF) above 15% (Supplementary Table S1) and copy number variation (CNV) (>200 bp) (Supplementary Table S2, S3) per spleen was comparable between TgSpi1FA+/+ and TgSpi1FA−/− mice (Fig. 4A, B). We identified a mean of 11.8 nonsilent mutations (range 8–20) for TgSpi1FA+/+ and 11.2 (range 7–17) for TgSpi1FA−/−. Qualitatively, the type (transversion vs. transition) as well as the single nucleotide variant (SNV) and indels resulting in protein variations (missense, nonsynonymous, splice site, stop-gain, frameshift and nonframeshift indels) were similar between TgSpi1FA+/+ and TgSpi1FA−/− cells (Fig. 4C–E). Hence, even though TgSpi1FA−/− leukaemic cells displayed higher levels of 53BP1 foci, DNA damage, chromosome breakage and intracellular reactive oxygen species (ROS) than TgSpi1FA+/+ preleukaemic cells (Supplementary Fig. S4), which are expected due to the lack of a functional FANCA protein, they did not show an increased level of genetic instability.
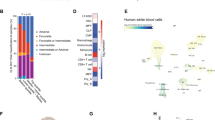
A Somatic mutation rates per animal (dot) are shown. Histograms indicate the mean ± SEM; ns: not significant Mann‒Whitney test. B Bar charts with error bars overlaid with dot plots showing the mean ± SEM of the number of CNV segments. Copy number gains and losses (>200 bp) were defined in TgSpi1FA+/+ preleukaemic and TgSpi1FA−/− leukaemic cells using the Control-FREEC tool. Each dot represents cells from one sick mouse. Histograms indicate the mean ± SEM; ns: nonsignificant Mann‒Whitney test. C Pie charts showing the distribution of the type of mutations in TgSpi1FA+/+ (n = 5) or TgSpi1FA−/− (n = 10) cells derived from sick mice. D Relative proportion of single nucleotide variations, transitions and transversions, and indels identified in TgSpi1FA+/+ (n = 5, outer circle) or TgSpi1FA−/− (n = 10, inner circle) cells derived from sick mice. E Types of transitions and transversions in TgSpi1FA+/+ (n = 5) or TgSpi1FA−/− (n = 10) cells for all sick mice from the indicated genotypes.
We compared the list of the 128 genes with identified variants (Supplementary Table S1) to a comprehensive list of 1081 “cancer genes” obtained by combining the Catalogue of Validated Oncogenic Mutations of the Cancer Genome Interpreter (CGI) database and the Cancer Gene Census (CGC) of the COSMIC database. Twenty-three genes displaying pathogenic mutations were identified as human cancer genes (Fig. 5 and Supplementary Fig. S5). Figure 5 also includes a variant of Fgd2, which is not considered to be a cancer gene in the CGI and CGC databases but has been repeatedly found to display this specific variant (at least 5 times) in the cancer samples recorded in the COSMIC database. Mutations in fifteen out of the 23 genes were also identified in human erythroleukaemia (Fig. 5) [16, 37].
Mutations in the Fgfr3, Ptpn11, Smad4 and Trp53 genes were identified in both genotypes that excludes that observed variants were directly responsible for the EPO independency and tumorigenicity of the TgSpi1FA−/− blasts.
The JAK2 V617F mutation, which is associated with myeloproliferative disorders and EPO hypersensitivity [38], was identified exclusively in blasts of TgSpi1FA+/+ mice, including #445 and #451, with VAF values of 28 and 18%, respectively. JAK2 Y931C is another mutation conferring gain of function [39] and was found in #474 mice. However, none of the JAK2 mutations conferred per se a proliferative advantage allowing in vivo clonal selection.
Interestingly, mutations in Flt3, Fgd2, Nras and Kit were identified exclusively in TgSpi1FA−/− cells. All of these genes are key oncogenes in leukaemia. The Kit D818Y (#26, #57, #92, #223) or Nras Q61H (#28, #39) mutations, identified in 45 and 20% of mice, respectively, have been previously shown to confer growth factor independency in the TgSpi1FA+/+ cells or myeloma cells, respectively [15, 40], indicating a causal relationship between constitutively activated KIT and RAS pathways and growth factor independence.
Our results show that the somatic mutation rate was similar between TgSpi1FA+/+ and TgSpi1FA−/− mice, but specific mutations in known oncogenes activating signalling pathways, Kit and Ras, were identified in FANCA-deficient cells. Considering the stochasticity of the mutational process, these data suggest that FANCA deficiency supports the selection of oncogenic mutations known to favour cell overgrowth. This hypothesis predicts that the oncogenic mutations identified in TgSpi1FA−/− leukaemic cells should be present in the cell populations issued from the bone marrow or spleen of the TgSpi1FA+/+ mice in low-prevalence subclones. Accordingly, searching for Kit or NRas variants with low VAF in the preleukaemic cells from TgSpi1FA+/+ mice, we found KIT D818Y (mice #445 VAF 1.9%) and NRAS Q61H or G60V mutations (mice #479 VAF 2.1% and mice #138 VAF 1.6%, respectively). However, such mutated cells did not expand to become a major population of the spleen of TgSpi1FA+/+ mice displaying active FANCA, even when maintained in culture for several months.
Altogether, our results indicate that loss of FANCA establishes a favourable cellular environment for the selection of cells carrying genes with oncogenic mutations.
A unique transcriptional deregulation is associated with the leukaemic status of FANCA-deficient cells
We hypothesized that transcriptomic modifications may contribute to the selection of leukaemic mutated clones in the absence of FANCA. We first compared RNA-seq data between TgSpi1FA−/− leukaemic and preleukaemic cells (model 1). To exclude interference due to EPO signalling, we performed RNA-seq on blasts cultured in the presence of EPO. We identified 3005 differentially regulated transcripts (|FC| ≥ 1.5; Padjusted < 0.05), of which 1181 were repressed and 1824 were increased, in TgSpi1FA−/− leukaemic compared to TgSpi1FA−/− preleukaemic cells (Fig. 6A, model 1 and Supplementary Table S4).
A Top, model 1 (FA loss-of-function): volcano plots of differentially expressed genes between the preleukaemic TgSpi1FA+/+ versus preleukaemic TgSpi1FA−/− cells or between the TgSpi1FA−/− preleukaemic and leukaemic cells. Bottom, model 2 (blood transfusion): volcano plots of differentially expressed genes between the TgSpi1FA+/+ preleukaemic versus leukaemic cells. Increased genes in leukaemic cells (FC ≥ 1.5, P < 0.05) are indicated in red; repressed genes (FC ≤ 0.67, P < 0.05) are indicated in green. B Relative expression of genes from NOTCH or WNT/β-catenin pathways measured by RNA-seq between leukaemic (L) and preleukaemic (PL) cells from mice with the indicated genotypes. TgSpi1FA+/+(#138, #445, #451, #479) or TgSpi1FA−/− (#40, #77, #224) preleukaemic cells or TgSpi1FA−/− leukaemic cells (#26, #28, #39). Adjusted P values for multiple testing using the Benjamini and Hochberg method are indicated. ns: not significant. C TgSpi1FA+/+(#138, #445, #451, #479) or TgSpi1FA−/− (#40, #77, #224) preleukaemic cells or TgSpi1FA−/− leukaemic cells (#26, #28, #39) were seeded at 2 × 104 cells/mL and treated with the indicated doses of the NOTCH inhibitor (LY411575). Living cells were counted 3 days post-treatment using a DAPI exclusion test and quantified as a percentage of the control DMSO. Data are the mean ± SEM of 3 independent experiments. Statistical analysis was carried out using two-way ANOVA against the DMSO control sample. * indicates P < 0.05; **P < 0.01. PL preleukaemic, L Leukaemic. D Relative expression of Mdm4 mRNA measured by RNA-seq in preleukaemic or leukaemic cells of mice with the indicated genotypes (PL preleukaemic, L leukaemic). P values (RNA-seq analysis) were adjusted for multiple testing using the Benjamini and Hochberg method.
GSEA analysis revealed that leukaemic cells were significantly enriched for functional annotations related to WNT/β-catenin signalling that also include genes of NOTCH signalling (Supplementary Fig. S6A). Relative expression of Lrp5, Vangl2, Notch1 or Jag1 analysed by RNA-seq experiments are shown as examples of DEG encoding proteins of the WNT/β-catenin or NOTCH signalling, respectively (Fig. 6B, middle red bars), and was validated by RT-qPCR in the TgSpi1FA−/− leukaemic compared to TgSpi1FA−/− preleukaemic cells (Supplementary Fig. S6B). Interestingly, only 173 transcripts were differentially expressed between the TgSpi1FA+/+ and TgSpi1FA−/− preleukaemic cells (Fig. 6A), and none of the genes of the WNT/β-catenin and NOTCH signalling pathways were deregulated (Fig. 6B and Supplementary Table S5), indicating a specific modification of the transcriptional profile in the FANCA-deficient leukaemic cells.
We next sought to determine whether the transcriptional deregulation was specific to the FANCA-depleted context. One strategy to answer this question is to characterize genes differentially expressed between preleukaemic and leukaemic TgSpi1FA+/+ cells. However, the rarity of leukaemic TgSpi1FA+/+ mice in model 1 (Fig. 3B) prevents such analysis. To resolve this question, we used another previously generated model of leukaemic progression initiated from TgSpi1FA+/+ preleukaemic mice (leukaemic model 2) [14, 15], whose leukaemic cells emerged after serial blood transfusions of preleukaemic TgSpi1FA+/+ mice. Blood transfusion lowered circulating EPO levels in anaemic mice and triggered in vivo selection and expansion of EPO-independent and tumorigenic cells with KIT gain-of-function mutations [14, 15]. Therefore, in this model, blood transfusion allows the emergence of leukaemic cells, instead of FANCA loss of function.
Only 310 transcripts were differentially expressed between preleukaemic and leukaemic TgSpi1FA+/+ cells (Fig. 6A, model 2, Supplementary Table S6). Among them, 4 increased genes and 29 repressed genes were also found to be differentially expressed between preleukaemic and leukaemic TgSpi1FA−/− cells (model 1), none of which belonged to the WNT/β-catenin or NOTCH signalling pathways (Supplementary Table S7, Fig. 6B and Supplementary Fig. S6B, model 2). Therefore, FANCA deficiency is associated with a specific transcriptomic programme in leukaemic cells. Consistent with the specific transcriptional modification, the TgSpi1FA−/− leukaemic cells displayed hypersensitivity to inhibition of NOTCH signalling compared to preleukaemic TgSpi1FA−/− or TgSpi1FA+/+ cells (Fig. 6C). These data demonstrate the requirement of NOTCH signalling in the long-term proliferation of TgSpi1FA-/ - leukaemic cells.
Recent demonstration shows that MDM4 amplification in the bone marrow cells of FA patients confers greater fitness to the cells and yields clonal expansion preceding transformation to myeloid leukaemia [41]. In agreement, we found a higher expression of Mdm4 mRNA in the TgSpi1FA−/− leukaemic compared to preleukaemic cells, that was not observed in the leukaemic TgSpi1FA+/+ mice (Fig. 6D, model 1 versus model 2). These data reinforce the hypothesis that FANCA deficiency leads to transcriptional changes that are required for leukaemic transformation and suggest that, as observed in FA patients, MDM4 contributes to the proliferative advantage of FA murine leukemic cells [41].
In conclusion, we demonstrate a peculiar transcriptional deregulation associated with the leukaemic status of cells devoid of FANCA activity. We propose that these specific transcriptional modifications represent a key point participating in the clonal selection of TgSpi1FA−/− leukaemic cells by conferring advantage to the cells with secondary mutations in Kit or Ras genes (Fig. 7).
WT and Fanca-deficient mice exhibit normal erythropoiesis (top). SPI1 overexpression blocks erythroid differentiation at the CFU-E stage and maintains the proliferation of progenitors, allowing the accumulation of spontaneously arising mutations without leading to growth factor independence (middle). The additional absence of the DDR protein FANCA increases the transcription of genes encoding proteins favouring proliferation pathways, such as MDM4, NOTCH and WNT/β-catenin and favours the expansion of clones harbouring Kit or Nras oncogenic mutations conferring EPO independency. This combination of events underlies a rapid leukaemic progression (bottom).
Discussion
This work provides unexpected evidence that deficiency in the FANCA DDR protein fosters leukaemic progression not by increasing the rate of mutations but by allowing the proliferation of cells with oncogenic variants associated with a reprogrammed transcriptomic landscape.
Although preleukaemic erythroblastic cells expressing high levels of SPI1 exhibit replicative stress [32, 33], the additional deficiency in DNA repair and replication rescue by FANCA loss-of-function did not further increase the gene and genomic instability of the cells. This outcome is in agreement with published data demonstrating that FANC pathway deficiency leads to a reduced mutational load in surviving cells both in vitro and in vivo [42,43,44,45]. A recent study performing exome sequencing on FA patient-derived hematopoietic cells with clonal evolution also showed that FA patient samples did not display more somatic mutations than control AMLs [41].
Leukaemic TgSpi1 cells due to FANCA loss-of-function displayed constitutive activation of EPO signalling pathways that supported their proliferation, similarly to the blood transfusion-derived leukaemic TgSpi1 cells with a normal FANC pathway [14]. In both models, EPO independency and tumorigenicity were associated with activating mutations in genes encoding key signalling proteins, i.e., Kit or Nras. Cells with NRAS activation displayed strong and/or constitutive activation of ERK and AKT, respectively, while STAT5 was not constitutively increased. In contrast, KIT activation was associated with constitutive activation of ERK, AKT and STAT5. These results support the major function of different oncogenic mutations in the autonomous proliferation and survival of leukaemic cells, together with FANCA depletion and SPI1 overexpression. Our work demonstrated the existence of cells with Kit and Nras mutations with a low VAF in EPO-dependent, preleukaemic cells isolated from TgSpi1FA+/+ mice. These cells rarely spontaneously invade the bone marrow or spleen of mice, in contrast to TgSpi1FA−/− mice. The leukaemic progression from the TgSpi1 model described by Moreau-Gachelin et al. [14] (leukaemic model 2) were due to serial blood transfusion in anaemic animals that transitorily palliates to their anaemic status and blocks EPO production. This established a selective pressure allowing in vivo expansion of rare EPO-independent cells with Kit mutations [15]. In contrast, TgSpi1FA−/− mice develop spontaneously overt leukaemia. Such observations strengthen the hypothesis that leukaemic transition in TgSpi1FA−/− mice is sustained by additional changes in the intracellular or extracellular environment, which favours the expansion and selection of cells with oncogenic mutations. Our findings show that in contrast to leukaemic cells with normal FANCA, cells devoid of FANCA and mutated in signalling oncogenes displayed a reprogrammed transcriptional landscape. We hypothesized that transcriptional reprogramming might be involved in the emergence of leukaemic cells. Examples of the contribution of transcriptional deregulation to cellular expansion in oncogene mutated cells are reported in the literature. Indeed, transcriptional deregulation may favour the expansion of cells with NRASG12D oncoproteins whose proliferation required a reduction in Spry2 transcription, encoding a negative regulator of the RAS signalling pathway and, thus represents an additional event, which augments MAPK signalling and contributes to transformation [46]. Similarly, the finding that epiallele, referring to variation in methylation of CpGs in discrete sets and differing among AML cells, may have functional consequences on transcription is of significance [47]. Indeed, it was recently proposed that this epiallele and, thus, transcriptional diversity can occur prior to overt transformation, possibly enabling preleukaemic cells to overt leukaemogenesis [48].
It was recently reported in FA patients with clonal evolution of bone marrow cells that 1q trisomy is the most frequent chromosomal event, driving enforced MDM4 oncogene expression through gene dosage and attenuating the p53 anti-cancer barrier in FA [41]. Such p53 attenuation confers a keen advantage to MDM4 overexpressing cells in FA cells, triggering stem/progenitor cell survival and clonal expansion but is not transforming per se; additional oncogenic mutations are required (3q+, RUNX1, RAS…). In the erythroleukaemic model used in our study, we also found overexpression of Mdm4 during the emergence of FANCA-depleted leukaemic clones, as well as additive mutations, including Ras and Kit genes. Additionally, FANCA loss of function in leukaemic SPI1-overexpressing cells deregulates a complex network of genes known for their strong association with leukaemia development. In particular, several membrane receptors/ligands or downstream signalling pathways, including WNT/β-catenin and NOTCH signalling, were transcriptionally deregulated in Fanca-deleted leukaemic cells. Increased expression of WNT/β-catenin or NOTCH and associated pathways has already been reported in FANCC-deficient B lymphoid human and mouse cells [49, 50]. Nevertheless, Fanca deletion alone in TgSpi1 preleukaemic cells or the presence of oncoproteins of signalling pathways alone in TgSpi1 leukaemic cells with functional FANCA were not sufficient to strongly activate WNT/β-catenin or NOTCH pathways. These data reveal a strict association between transcriptomic deregulation in cells with FANCA loss-of-function and activating mutations of signalling pathways. We suggest that MDM4, WNT/β-catenin or NOTCH signalling participates in the selection and expansion of cells with constitutively increased KIT or RAS signalling pathways.
Accordingly, activation of the β-Catenin pathway was recently demonstrated to be responsible for the preleukaemic to AML transition in distinct preleukaemic diseases due to MDMX overexpression [13]. In the TgSpi1FA−/−-derived leukaemia cells, the inhibition of WNT/β-catenin signalling did not affect cell proliferation, at least in vitro, consistent with a role of WNT/β-catenin signalling in the preleukaemic to leukaemic transition rather than in leukaemia maintenance. In contrast, the NOTCH pathway was required to maintain the proliferative status of TgSpi1FA−/− leukaemic cells. Future studies are required to characterize the mechanisms responsible for the overactivation of genes controlling proliferation pathways. One hypothesis to investigate is WNT/β-catenin and NOTCH signalling, such as MDM4, plays a role in repression of the p53 anti-tumoral barrier in FA−/− cells.
Together with reported studies [41, 51], our work supports that transcriptional deregulation is one of the mechanisms by which the absence of Fanconi proteins may contribute to abnormal haematopoietic cell function. We propose that cells with FANC deficiency are prone to transcriptional modifications of genes encoding proteins participating in multiple signalling pathways, favouring the expansion of clones carrying Ras or Kit oncogenic drivers and thus supporting leukaemogenesis (Fig. 7).
Although it remains to be determined how these functions ultimately exert a positive effect on the emergence of cells carrying oncogenic mutations, they represent targets that can be disrupted to subvert leukaemic progression.
Methods
Mice and cell culture
Fanca+/− and TgSpi1 mice were derived from the FVB/N and DBA2/J backgrounds, respectively [14, 28]. Mice deficient in the Fanca gene and expressing TgSpi1 (TgSpi1Fanca−/− or TgSpi1FA−/−) were generated by mating Fanca+/− mice with homozygous TgSpi1 mice to obtain double-heterozygous mice (F1) (Fig. 1A). These F1 mice were then mated with Fanca+/− mice to generate mice harbouring homozygous mutations of Fanca and heterozygous TgSpi1. The project was officially approved by the Animal Experimentation Ethics Committee (registered by the French Department of Research) and conducted in accordance with French laws and regulations. The resulting mice of the indicated genotypes were monitored for survival. When needed, moribund mice were sacrificed for analysis of different biological parameters.
Preleukaemic and leukaemic cells derived from TgSpi1 mice were obtained previously [14, 15].
For NOTCH inhibition, cells were treated for 3 days with LY411575 (Sigma‒Aldrich), a NOTCH and γ secretase inhibitor. The number of viable cells was determined with DAPI labelling (0.5 µM) in 96-well plates by the BD High-Throughput Sampler (HTS) option from the CytoFLEX flow cytometer (Beckman Coulter). Flow cytometry data were analysed using FlowJo v10.6.2 software (Tree Star).
Real-time quantitative PCR (RT-qPCR)
Total RNA was isolated from cells using the RNeasy Plus Mini Kit or Maxwell RSC simplyRNA Cell Kit (Promega), and RT‒qPCR was performed using TaqMan or SYBR Green technology as described in detail in the Supplementary data.
Bone marrow and spleen immunophenotyping by flow cytometry
Single-cell suspensions of freshly collected bone marrow cells were immunostained with APC-H7 (560185, BD Biosciences) or APCeFluor780 (eBioscience 47-1172-82) -conjugated rat anti-KIT, FITC-conjugated rat anti-CD71 (553266, BD Biosciences), APC-conjugated rat anti-Ter119 (557909, BD Biosciences) and PE-conjugated streptavidin (554061, BD Biosciences) and biotin-conjugated rat anti-CD123 (555070, BD Biosciences) antibodies for the detection of erythroid progenitor (CFU-E) cells, as previously described [52]; FITC-conjugated rat anti-CD11b (553310, BD Biosciences), PerCP-CyTM5.5-conjugated rat anti-mouse CD19 (45-0193-82, eBioscience) and PE-conjugated rat anti-mouse CD4 (553049, BD Biosciences), and APC-conjugated rat anti-myeloid CD8 (553035, BD Biosciences) and PE-CyTM7- conjugated rat anti-myeloid B220 (25-0452-82, eBioscience) antibodies were used for the detection of freshly collected bone marrow cells. Stained cells were analysed with a BD Accuri C6 or a Fortessa analyser (BD Biosciences), and FlowJo (Tree Star) was used for data acquisition and analysis.
Additional details of the methods are included in the supplementary data.
Data availability
The high-throughput data generated are provided in the Gene Expression Omnibus under accession number GSE164411.
References
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Alexandrov LB, Kim J, Haradhvala NJ, Huang MN, Tian Ng AW, Wu Y, et al. The repertoire of mutational signatures in human cancer. Nature. 2020;578:94–101.
Sharma R, Lewis S, Wlodarski MW. DNA repair syndromes and cancer: insights into genetics and phenotype patterns. Front Pediatr. 2020;8:570084.
Knijnenburg TA, Wang L, Zimmermann MT, Chambwe N, Gao GF, Cherniack AD, et al. Genomic and molecular landscape of DNA damage repair deficiency across the cancer genome atlas. Cell Rep. 2018;23:239–54. e236
Martincorena I, Roshan A, Gerstung M, Ellis P, Van Loo P, McLaren S, et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science. 2015;348:880–6.
Jaiswal S, Fontanillas P, Flannick J, Manning A, Grauman PV, Mar BG, et al. Age-related clonal hematopoiesis associated with adverse outcomes. N Engl J Med. 2014;371:2488–98.
Mueller BU, Pabst T, Osato M, Asou N, Johansen LM, Minden MD, et al. Heterozygous PU.1 mutations are associated with acute myeloid leukemia. Blood. 2002;100:998–1007.
Rosenbauer F, Wagner K, Kutok JL, Iwasaki H, Le Beau MM, Okuno Y, et al. Acute myeloid leukemia induced by graded reduction of a lineage-specific transcription factor, PU.1. Nat Genet. 2004;36:624–30.
Moreau-Gachelin F, Tavitian A, Tambourin P. Spi-1 is a putative oncogene in virally induced murine erythroleukemia. Nature. 1988;331:277–80.
Roos-Weil D, Decaudin C, Armand M, Della-Valle V, Diop MK, Ghamlouch H, et al. A recurrent activating missense mutation in waldenstrom macroglobulinemia affects the DNA binding of the ETS transcription factor SPI1 and enhances proliferation. Cancer Discov. 2019;9:796–811.
Seki M, Kimura S, Isobe T, Yoshida K, Ueno H, Nakajima-Takagi Y, et al. Recurrent SPI1 (PU.1) fusions in high-risk pediatric T cell acute lymphoblastic leukemia. Nat Genet. 2017;49:1274–81.
Will B, Vogler TO, Narayanagari S, Bartholdy B, Todorova TI, da Silva Ferreira M, et al. Minimal PU.1 reduction induces a preleukemic state and promotes development of acute myeloid leukemia. Nat Med. 2015;21:1172–81.
Ueda T, Kanai A, Komuro A, Amano H, Ota K, Honda M, et al. KDM4B promotes acute myeloid leukemia associated with AML1-ETO by regulating chromatin accessibility. FASEB Bioadv. 2021;3:1020–33.
Moreau-Gachelin F, Wendling F, Molina T, Denis N, Titeux M, Grimber G, et al. Spi-1/PU.1 transgenic mice develop multistep erythroleukemias. Mol Cell Biol. 1996;16:2453–63.
Kosmider O, Denis N, Lacout C, Vainchenker W, Dubreuil P, Moreau-Gachelin F. Kit-activating mutations cooperate with Spi-1/PU.1 overexpression to promote tumorigenic progression during erythroleukemia in mice. Cancer Cell. 2005;8:467–78.
Fagnan A, Bagger FO, Pique-Borras MR, Ignacimouttou C, Caulier A, Lopez CK, et al. Human erythroleukemia genetics and transcriptomes identify master transcription factors as functional disease drivers. Blood. 2020;136:698–714.
Iacobucci I, Qu C, Varotto E, Janke LJ, Yang X, Seth A, et al. Modeling and targeting of erythroleukemia by hematopoietic genome editing. Blood. 2021;137:1628–40.
Lopez-Martinez D, Liang CC, Cohn MA. Cellular response to DNA interstrand crosslinks: the Fanconi anemia pathway. Cell Mol Life Sci. 2016;73:3097–114.
Renaudin X, Rosselli F. The FANC/BRCA pathway releases replication blockades by eliminating DNA interstrand cross-links. Genes (Basel). 2020;11:585.
Helbling-Leclerc A, Garcin C, Rosselli F. Beyond DNA repair and chromosome instability-Fanconi anaemia as a cellular senescence-associated syndrome. Cell Death Differ. 2021;28:1159–73.
Gueiderikh A, Maczkowiak-Chartois F, Rosselli F. A new frontier in Fanconi anemia: From DNA repair to ribosome biogenesis. Blood Rev. 2022;52:100904.
Nalepa G, Clapp DW. Fanconi anaemia and cancer: an intricate relationship. Nat Rev Cancer. 2018;18:168–85.
Bogliolo M, Surrallés J. Fanconi anemia: a model disease for studies on human genetics and advanced therapeutics. Curr Opin Genet Dev. 2015;33:32–40.
Gueiderikh A, Rosselli F, Neto JBC. A never-ending story: the steadily growing family of the FA and FA-like genes. Genet Mol Biol. 2017;40:398–407.
Bogliolo M, Bluteau D, Lespinasse J, Pujol R, Vasquez N, d'Enghien CD, et al. Biallelic truncating FANCM mutations cause early-onset cancer but not Fanconi anemia. Genet Med. 2018;20:458–63.
Fouquet B, Pawlikowska P, Caburet S, Guigon C, Mäkinen M, Tanner L, et al. A homozygous FANCM mutation underlies a familial case of non-syndromic primary ovarian insufficiency. Elife. 2017;6:e30490.
Carreau M. Not-so-novel phenotypes in the Fanconi anemia group D2 mouse model. Blood. 2004;103:2430.
Cheng NC, van de Vrugt HJ, van der Valk MA, Oostra AB, Krimpenfort P, de Vries Y, et al. Mice with a targeted disruption of the Fanconi anemia homolog Fanca. Hum Mol Genet. 2000;9:1805–11.
Parmar K, D'Andrea A, Niedernhofer LJ. Mouse models of Fanconi anemia. Mutat Res. 2009;668:133–40.
Pulliam-Leath AC, Ciccone SL, Nalepa G, Li X, Si Y, Miravalle L, et al. Genetic disruption of both Fancc and Fancg in mice recapitulates the hematopoietic manifestations of Fanconi anemia. Blood. 2010;116:2915–20.
Quentin S, Cuccuini W, Ceccaldi R, Nibourel O, Pondarre C, Pagès MP, et al. Myelodysplasia and leukemia of Fanconi anemia are associated with a specific pattern of genomic abnormalities that includes cryptic RUNX1/AML1 lesions. Blood. 2011;117:e161–170.
Rimmele P, Komatsu J, Hupe P, Roulin C, Barillot E, Dutreix M, et al. Spi-1/PU.1 oncogene accelerates DNA replication fork elongation and promotes genetic instability in the absence of DNA breakage. Cancer Res. 2010;70:6757–66.
Rimmelé P, Esposito M, Delestré L, Guervilly JH, Ridinger-Saison M, Despras E, et al. The Spi1/PU.1 transcription factor accelerates replication fork progression by increasing PP1 phosphatase in leukemia. Oncotarget. 2017;8:37104–14.
Liu Y, Pop R, Sadegh C, Brugnara C, Haase VH, Socolovsky M. Suppression of Fas-FasL coexpression by erythropoietin mediates erythroblast expansion during the erythropoietic stress response in vivo. Blood. 2006;108:123–33.
Socolovsky M, Nam H, Fleming MD, Haase VH, Brugnara C, Lodish HF. Ineffective erythropoiesis in Stat5a(−/−)5b(−/−) mice due to decreased survival of early erythroblasts. Blood. 2001;98:3261–73.
Barnache S, Mayeux P, Payrastre B, Moreau-Gachelin F. Alterations of the phosphoinositide 3-kinase and mitogen-activated protein kinase signaling pathways in the erythropoietin-independent Spi- 1/PU.1 transgenic proerythroblasts. Blood. 2001;98:2372–81.
Iacobucci I, Wen J, Meggendorfer M, Choi JK, Shi L, Pounds SB, et al. Genomic subtyping and therapeutic targeting of acute erythroleukemia. Nat Genet. 2019;51:694–704.
Vainchenker W, Constantinescu SN. A unique activating mutation in JAK2 (V617F) is at the origin of polycythemia vera and allows a new classification of myeloproliferative diseases. Hematology Am Soc Hematol Educ Program. 2005;2005:195–200.
Hornakova T, Springuel L, Devreux J, Dusa A, Constantinescu SN, Knoops L, et al. Oncogenic JAK1 and JAK2-activating mutations resistant to ATP-competitive inhibitors. Haematologica. 2011;96:845–53.
Billadeau D, Jelinek DF, Shah N, LeBien TW, Van Ness B. Introduction of an activated N-ras oncogene alters the growth characteristics of the interleukin 6-dependent myeloma cell line ANBL6. Cancer Res. 1995;55:3640–6.
Sebert M, Gachet S, Leblanc T, Rousseau A, Bluteau O, Kim R, et al. Clonal hematopoiesis driven by chromosome 1q/MDM4 trisomy defines a canonical route toward leukemia in Fanconi anemia. Cell Stem Cell. 2023;30:153–70. e159
Sala-Trepat M, Boyse J, Richard P, Papadopoulo D, Moustacchi E. Frequencies of HPRT- lymphocytes and glycophorin A variants erythrocytes in Fanconi anemia patients, their parents and control donors. Mutat Res. 1993;289:115–26.
Papadopoulo D, Guillouf C, Mohrenweiser H, Moustacchi E. Hypomutability in Fanconi anemia cells is associated with increased deletion frequency at the HPRT locus. Proc Natl Acad Sci USA. 1990;87:8383–7.
Mirchandani KD, McCaffrey RM, D'Andrea AD. The Fanconi anemia core complex is required for efficient point mutagenesis and Rev1 foci assembly. DNA Repair. 2008;7:902–11.
Laquerbe A, Guillouf C, Moustacchi E, Papadopoulo D. The mutagenic processing of psoralen photolesions leaves a highly specific signature at an endogenous human locus. J Mol Biol. 1995;254:38–49.
Kunimoto H, Meydan C, Nazir A, Whitfield J, Shank K, Rapaport F, et al. Cooperative epigenetic remodeling by TET2 loss and NRAS mutation drives myeloid transformation and MEK inhibitor sensitivity. Cancer Cell. 2018;33:44–59.e48.
Landau DA, Clement K, Ziller MJ, Boyle P, Fan J, Gu H, et al. Locally disordered methylation forms the basis of intratumor methylome variation in chronic lymphocytic leukemia. Cancer Cell. 2014;26:813–25.
Li S, Chen X, Wang J, Meydan C, Glass JL, Shih AH, et al. Somatic mutations drive specific, but reversible, epigenetic heterogeneity states in AML. Cancer Discov. 2020;10:1934–49.
Sertorio M, Amarachintha S, Wilson A, Pang Q. Loss of Fancc impairs antibody-secreting cell differentiation in mice through deregulating the Wnt signaling pathway. J Immunol. 2016;196:2986–94.
Zanier R, Briot D, Dugas du Villard JA, Sarasin A, Rosselli F. Fanconi anemia C gene product regulates expression of genes involved in differentiation and inflammation. Oncogene. 2004;23:5004–13.
Rodríguez A, Zhang K, Färkkilä A, Filiatrault J, Yang C, Velázquez M, et al. MYC promotes bone marrow stem cell dysfunction in Fanconi anemia. Cell Stem Cell. 2021;28:33–47. e38
Delestre L, Cui H, Esposito M, Quiveron C, Mylonas E, Penard-Lacronique V, et al. Senescence is a Spi1-induced anti-proliferative mechanism in primary hematopoietic cells. Haematologica. 2017;102:1850–60.
Acknowledgements
This work was supported by grants from Inserm (CG), Ligue Nationale contre le Cancer and l’Agence nationale de la recherche (ANR-21-CE17-0052-02)(FR, CG) and Gefluc Ile-de-France (PP). PP and MBD were supported by SIRIC (INCa-DGOS-INSERM 6043); LD was supported by ITMO Cancer de l’alliance nationale pour les sciences de la vie et de la santé (AVIESAN) and Fondation de France; SG was supported by the Ministère de l’Enseignement Supérieur; and OA was supported by the Ministère de l’Enseignement Supérieur and the Fondation pour la Recherche Médicale. ME was supported by the Institut National du Cancer (PLBIO-06-1). The authors wish to thank F. Moreau-Gachelin for helpful scientific discussions; N. Droin (Gustave Roussy genomic platform, France); S. Legras, C. Thibault-Carpentier, D. Plassard and B. Jost for sequencing and bioinformatics analyses (GenomEast platform, France Genomique consortium); Y. Lecluse, P. Rameau and C. Catelain (Imaging and Cytometry Platform, Gustave Roussy, France) for flow cytometry; L. Touchard, C. Metereau, P. Gonin and K. Ser-Leroux (Preclinical Evaluation Unit, Gustave Roussy) for animal care; O. Bawa; N. Signole (Experimental and Translational Platform of Pathology AMMICa Inserm US23/UMS CNRS3655) and R. Corre (UMR9019) for contributing to the IHC analyses.
Author information
Authors and Affiliations
Contributions
PP and LD: conceptualization, performed experiments, methodology, analysed data, contributed to article’s writing. SG, AO and ME: performed experiments, methodology. MBP: bioinformatics analyses. FR and CG: conceived the project, methodology, analysed data, supervised, funding acquisition, wrote the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pawlikowska, P., Delestré, L., Gregoricchio, S. et al. FANCA deficiency promotes leukaemic progression by allowing the emergence of cells carrying oncogenic driver mutations. Oncogene 42, 2764–2775 (2023). https://doi.org/10.1038/s41388-023-02800-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-023-02800-9
- Springer Nature Limited