Abstract
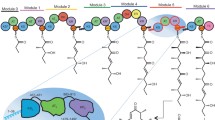
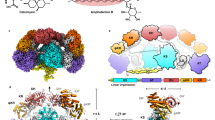
The polyketide synthase (PKS) mega-enzyme assembly line uses a modular architecture to synthesize diverse and bioactive natural products that often constitute the core structures or complete chemical entities for many clinically approved therapeutic agents1. The architecture of a full-length PKS module from the pikromycin pathway of Streptomyces venezuelae creates a reaction chamber for the intramodule acyl carrier protein (ACP) domain that carries building blocks and intermediates between acyltransferase, ketosynthase and ketoreductase active sites (see accompanying paper2). Here we determine electron cryo-microscopy structures of a full-length pikromycin PKS module in three key biochemical states of its catalytic cycle. Each biochemical state was confirmed by bottom-up liquid chromatography/Fourier transform ion cyclotron resonance mass spectrometry. The ACP domain is differentially and precisely positioned after polyketide chain substrate loading on the active site of the ketosynthase, after extension to the β-keto intermediate, and after β-hydroxy product generation. The structures reveal the ACP dynamics for sequential interactions with catalytic domains within the reaction chamber, and for transferring the elongated and processed polyketide substrate to the next module in the PKS pathway. During the enzymatic cycle the ketoreductase domain undergoes dramatic conformational rearrangements that enable optimal positioning for reductive processing of the ACP-bound polyketide chain elongation intermediate. These findings have crucial implications for the design of functional PKS modules, and for the engineering of pathways to generate pharmacologically relevant molecules.




Similar content being viewed by others
Accession codes
Primary accessions
Electron Microscopy Data Bank
Data deposits
The cryo-EM maps have been deposited in the Electron Microscopy Data Bank under accession numbers EMD-5663 (pentaketide–KS5–PikAIII), EMD-5664 (β-ketohexaketide–PikAIII), EMD-5665 (β-hydroxyhexaketide–PikAIII conformation i), EMD-5666 (β-hydroxyhexaketide–PikAIII conformation ii) and EMD-5667 (β-hydroxyhexaketide–PikAIII conformation iii).
References
Newman, D. J. & Cragg, G. M. Natural products as sources of new drugs over the 30 years from 1981 to 2010. J. Nat. Prod. 75, 311–335 (2012)
Dutta, S. et al. Structure of a modular polyketide synthase. Naturehttp://dx.doi.org/10.1038/nature13423 (this issue)
Kittendorf, J. D. & Sherman, D. H. Developing tools for engineering hybrid polyketide synthetic pathways. Curr. Opin. Biotechnol. 17, 597–605 (2006)
Walsh, C. T. Combinatorial biosynthesis of antibiotics: challenges and opportunities. ChemBioChem 3, 124–134 (2002)
Menzella, H. G. & Reeves, C. D. Combinatorial biosynthesis for drug development. Curr. Opin. Microbiol. 10, 238–245 (2007)
Menzella, H. G., Carney, J. R. & Santi, D. V. Rational design and assembly of synthetic trimodular polyketide synthases. Chem. Biol. 14, 143–151 (2007)
Menzella, H. G. et al. Combinatorial polyketide biosynthesis by de novo design and rearrangement of modular polyketide synthase genes. Nature Biotechnol. 23, 1171–1176 (2005)
Sherman, D. H. The Lego-ization of polyketide biosynthesis. Nature Biotechnol. 23, 1083–1084 (2005)
Fischbach, M. A. & Walsh, C. T. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: logic, machinery, and mechanisms. Chem. Rev. 106, 3468–3496 (2006)
Xue, Y., Zhao, L., Liu, H. W. & Sherman, D. H. A gene cluster for macrolide antibiotic biosynthesis in Streptomyces venezuelae: architecture of metabolic diversity. Proc. Natl Acad. Sci. USA 95, 12111–12116 (1998)
Hansen, D. A. et al. Biocatalytic synthesis of pikromycin, methymycin, neomethymycin, novamethymycin, and ketomethymycin. J. Am. Chem. Soc. 135, 11232–11238 (2013)
Tang, Y., Kim, C. Y., Mathews, I. I., Cane, D. E. & Khosla, C. The 2.7-Å crystal structure of a 194-kDa homodimeric fragment of the 6-deoxyerythronolide B synthase. Proc. Natl Acad. Sci. USA 103, 11124–11129 (2006)
Keatinge-Clay, A. T. & Stroud, R. M. The structure of a ketoreductase determines the organization of the β-carbon processing enzymes of modular polyketide synthases. Structure 14, 737–748 (2006)
Alekseyev, V. Y., Liu, C. W., Cane, D. E., Puglisi, J. D. & Khosla, C. Solution structure and proposed domain domain recognition interface of an acyl carrier protein domain from a modular polyketide synthase. Protein Sci. 16, 2093–2107 (2007)
Aldrich, C. C., Beck, B. J., Fecik, R. A. & Sherman, D. H. Biochemical investigation of pikromycin biosynthesis employing native penta- and hexaketide chain elongation intermediates. J. Am. Chem. Soc. 127, 8441–8452 (2005)
Aldrich, C. C., Venkatraman, L., Sherman, D. H. & Fecik, R. A. Chemoenzymatic synthesis of the polyketide macrolactone 10-deoxymethynolide. J. Am. Chem. Soc. 127, 8910–8911 (2005)
Wong, F. T., Chen, A. Y., Cane, D. E. & Khosla, C. Protein–protein recognition between acyltransferases and acyl carrier proteins in multimodular polyketide synthases. Biochemistry 49, 95–102 (2010)
Caffrey, P. Conserved amino acid residues correlating with ketoreductase stereospecificity in modular polyketide synthases. ChemBioChem 4, 654–657 (2003)
Keatinge-Clay, A. T. A tylosin ketoreductase reveals how chirality is determined in polyketides. Chem. Biol. 14, 898–908 (2007)
Bonnett, S. A. et al. Structural and stereochemical analysis of a modular polyketide synthase ketoreductase domain required for the generation of a cis-alkene. Chem. Biol. 20, 772–783 (2013)
Lyon, A. M., Dutta, S., Boguth, C. A., Skiniotis, G. & Tesmer, J. J. Full-length Gαq–phospholipase C-β3 structure reveals interfaces of the C-terminal coiled-coil domain. Nature Struct. Mol. Biol. 20, 355–362 (2013)
Strunk, B. S. et al. Ribosome assembly factors prevent premature translation initiation by 40S assembly intermediates. Science 333, 1449–1453 (2011)
Nguyen, C. et al. Trapping the dynamic acyl carrier protein in fatty acid biosynthesis. Nature 505, 427–431 (2014)
Masoudi, A., Raetz, C. R., Zhou, P. & Pemble, C. W. IV Chasing acyl carrier protein through a catalytic cycle of lipid A production. Nature 505, 422–426 (2014)
Bunkoczi, G. et al. Mechanism and substrate recognition of human holo ACP synthase. Chem. Biol. 14, 1243–1253 (2007)
Beck, B. J., Aldrich, C. C., Fecik, R. A., Reynolds, K. A. & Sherman, D. H. Iterative chain elongation by a pikromycin monomodular polyketide synthase. J. Am. Chem. Soc. 125, 4682–4683 (2003)
Yin, Y., Lu, H., Khosla, C. & Cane, D. E. Expression and kinetic analysis of the substrate specificity of modules 5 and 6 of the picromycin/methymycin polyketide synthase. J. Am. Chem. Soc. 125, 5671–5676 (2003)
Pfeifer, B. A., Admiraal, S. J., Gramajo, H., Cane, D. E. & Khosla, C. Biosynthesis of complex polyketides in a metabolically engineered strain of E. coli. Science 291, 1790–1792 (2001)
Ohi, M., Li, Y., Cheng, Y. & Walz, T. Negative staining and image classification—powerful tools in modern electron microscopy. Biol. Proced. Online 6, 23–34 (2004)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nature Methods 9, 853–854 (2012)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nature Methods 10, 584–590 (2013)
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013)
Fernández, J. J., Luque, D., Caston, J. R. & Carrascosa, J. L. Sharpening high resolution information in single particle electron cryomicroscopy. J. Struct. Biol. 164, 170–175 (2008)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Zheng, J., Taylor, C. A., Piasecki, S. K. & Keatinge-Clay, A. T. Structural and functional analysis of A-type ketoreductases from the amphotericin modular polyketide synthase. Structure 18, 913–922 (2010)
Bonnett, S. A. et al. Acyl-CoA subunit selectivity in the pikromycin polyketide synthase PikAIV: steady-state kinetics and active-site occupancy analysis by FTICR-MS. Chem. Biol. 18, 1075–1081 (2011)
Acknowledgements
This work was supported by the Pew Scholar Program in Biomedical Sciences (G.S.), the University of Michigan Biological Sciences Scholars Program (G.S.), Rackham Merit and American Foundation for Pharmaceutical Education predoctoral fellowships (D.A.H.), a National Research Service Award postdoctoral fellowship (J.A.C.), the Life Sciences Research Foundation (A.R.H.N.), National Institutes of Health grant 1R21CA138331-01A1 (K.H.), GM076477 (D.H.S. and J.L.S.), DK042303 (J.L.S.), and DK090165 (G.S.), and the Hans W. Vahlteich Professorship (to D.H.S.).
Author information
Authors and Affiliations
Contributions
J.R.W. produced PikAIII variants and conducted enzyme assays. S.D. carried out cryo-EM data collection and processing. A.M.D. assisted with cryo-EM image processing. W.A.H., A.R.H.N. and K.H. carried out mass-spectrometry analysis. D.A.H. synthesized the thiophenol-activated pentaketide substrate. J.A.C. produced initial PikAIII samples. S.D., J.R.W., K.H., J.L.S. and G.S. analysed the data and interpreted results. D.H.S., J.L.S. and G.S. designed research. J.R.W., S.D., K.H., D.H.S., J.L.S. and G.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Partial mass spectra of active site peptides from LC/FT-ICR MS of trypsin-digested PikAIII.
a–c, KS5 peptides observed after incubation with thiophenol-activated pentaketide. Pentaketide was detected on KS5 Cys 209. The pentaketide thioester at KS5 Cys 209 is labile38, and substantial quantities of unloaded KS5 Cys 209 are expected to form during trypsin digestion and handling for MS analysis. d–i, Example mass spectra of active site KS5, AT5 and ACP5 peptides after incubation with both MM–CoA and thiophenol-activated pentaketide. Pentaketide was detected on KS Cys 209, MM was detected on the AT Ser 655, and β-ketohexaketide was detected on the ACP Ser 1438. j, Active site ACP5 peptide after incubation with MM–CoA and thiophenol-activated pentaketide (top), and MM–CoA, thiophenol-activated pentaketide and buffered NADPH (bottom). After addition of NADPH, β-hydroxyhexaketide was detected on the ACP Ser 1438.
Extended Data Figure 2 Estimation of cryo-EM map resolution by Fourier shell correlation and assessment of over-refinement.
a, b, Conventional Fourier shell correlation (FSC) curves (blue), gold-standard FSC (red) and high-resolution phase randomization test (green) for pentaketide–KS5–PikAIII (a) and pentaketide–KS5–PikAIII (b) three-dimensional reconstructions. The indicated resolution at FSC = 0.5 of the conventional FSC is in agreement with FSC = 0.143 of the gold-standard FSC. FSC calculations with phase randomized data show a sharp drop-off at the expected resolution level (10 Å). c, Conventional FSC curves for each conformer of β-hydroxyhexaketide–PikAIII showing a resolution between 10.7 Å and 11.6 Å at FSC = 0.5. d, Summary of conventional and gold-standard FSC resolution indications. Additionally provided is the agreement by FSC (at 0.5) between the average map of the two gold-standard half-maps and the conventional map (full data set), showing consistency with conventional and gold-standard FSC calculations. The right column of the table lists the cross-correlation values between each PikAIII three-dimensional reconstruction and the produced pseudo-atomic resolution model of the docked crystal structures at the indicated resolution range.
Extended Data Figure 3 Stereo views of the pentaketide–PikAIII three-dimensional map fit with the DEBS module 5 KS (blue) and AT (green) crystal structures.
a, A view looking towards the side of the KS. Upon formation of the pentaketide–KS intermediate, the side entrance to the KS active site (red star) is partially occluded and certain loop regions on the top of KS (indicated with an arrow) appear to move as indicated by differences between the EM map and KS crystal structure. b, Cross-section of the view looking towards the bottom entrance to the KS active site. Upon KS–pentaketide formation, new density (blue) appears around the KS bottom entrance that is not accounted for by the KS crystal structure.
Extended Data Figure 4 PikAIII–TE functional assays.
a, Example high-performance liquid chromatography traces of PikAIII–TE assay. The levels of 10-dml produced by wild type PikAIII–TE (red trace), R747A PikAIII–TE (green), R1133E PikAIII–TE (orange), H1137E PikAIII–TE (blue), R1308 PikAIII–TE (brown) and a no enzyme control (yellow) are shown. b, Activity of PikAIII–TE mutants. All reactions were completed in triplicate and the data are presented as average ± standard deviation.
Extended Data Figure 5 PikAIII/PikAIV functional assays.
a, Example high-performance liquid chromatography (HPLC) traces of PikAIII/PikAIV assays with (red trace) and without NADPH (blue). Peaks for 10-dml, narbonolide (nbl) and thiophenol-activated pentaketide (starting material) are indicated. b, Levels of thiophenol-activated pentaketide consumed in the PikAIII/PikAIV reactions with and without NADPH. All reactions were completed in triplicate and the data are presented as average ± standard deviation.
Rights and permissions
About this article
Cite this article
Whicher, J., Dutta, S., Hansen, D. et al. Structural rearrangements of a polyketide synthase module during its catalytic cycle. Nature 510, 560–564 (2014). https://doi.org/10.1038/nature13409
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13409
- Springer Nature Limited
This article is cited by
-
Insights into azalomycin F assembly-line contribute to evolution-guided polyketide synthase engineering and identification of intermodular recognition
Nature Communications (2023)
-
Enzymology of assembly line synthesis by modular polyketide synthases
Nature Chemical Biology (2023)
-
Solution structure of the type I polyketide synthase Pks13 from Mycobacterium tuberculosis
BMC Biology (2022)
-
Structures and function of a tailoring oxidase in complex with a nonribosomal peptide synthetase module
Nature Communications (2022)
-
Bifurcation drives the evolution of assembly-line biosynthesis
Nature Communications (2022)