Abstract
Background
Newcastle disease (ND) is a viral disease affecting a wide range of bird species and has a considerable financial impact on the world's poultry market. The ND virus (NDV) strains currently circulating in poultry throughout Africa, and especially in East Africa, exhibit significant genetic variation.
Objectives
The primary objective of the present investigation was to investigate the NDV genotypes in chickens raised in backyards in Tanzania's Morogoro and Iringa districts, which were associated with ND outbreaks.
Methods
Two tissue samples from chickens taken during a suspected ND outbreak in Tanzania's Morogoro (Eastern zone) and Iringa (Southern highlands zone) were subjected to reverse transcription polymerase chain reaction targeting the fusion (F) and hemagglutinin-neuraminidase (HN) genes, followed by sequencing.
Results
Based on comprehensive analysis of the entire F and HN gene sequences, the viruses were categorized as genotype VII and displayed significant genetic similarity with NDV strains previously identified in Mozambique, South Africa, Zambia, Zimbabwe, Botswana, Southeast Asia, and China. The uniformity in the amino acid cleavage site motif of the F protein across the examined NDV isolates, characterized by 112R–R–Q/K–K–R–F117, indicates their classification as virulent strains.
Conclusion
Regularly characterizing circulating strains and expanding the study to other parts of Tanzania may help to enhance disease control by giving a more precise picture of the situation regarding ND, especially in light of the issues posed by NDV genotype VII elsewhere.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Background
Newcastle disease (ND) impacts a diverse array of avian species and has significant global influence over the commercial poultry sector. Given ND's economic implications on the chicken industry [1], it warrants notification to the World Organisation for Animal Health (WOAH). The causative agent of ND is Orthoavulavirus javaense, previously named Avian orthoavulavirus 1 and commonly known as ND virus (NDV; used herein) within the family Paramyxoviridae and genus Orthoavulavirus [2]. NDV’s single-stranded, negative-sense RNA genome encodes for six proteins: nucleoprotein (N), phospho-protein (P), matrix protein (M), fusion protein (F), hemagglutinin-neuraminidase (HN), and large polymerase (L). RNA editing of P messenger RNA (mRNA) facilitates the expression of supplementary proteins, V and W [3]. The interplay between the envelope surface glycoproteins F and HN mediates infection of host cells by NDV [4]. The HN protein activates the F protein, facilitating the attachment of virus particles to sialic acid-containing receptors on cell surfaces [5]. This process facilitates the fusion between the viral envelope and the host cell membrane [4]. Distinctions among lentogenic, mesogenic, and velogenic strains of Newcastle Disease Virus (NDV) are based on the cleavage site of the F protein: lentogenic strains are characterized by an amino acid sequence of 112 G/E-K/R-Q-G/E-R-L117, while all mesogenic and velogenic NDV strains exhibit the sequence 112R/K-R-Q-R/K-R-F117 within the F protein [6].
All bird species are thought to be susceptible to NDV, which infects at least 250 different species [7]. Non-avian hosts, including humans, cattle, sheep, camels, pigs, rabbits, mink, mice, and hamsters, have been shown to exhibit signs of NDV infections [8]. Although human-to-human transmission has not been reported, bird-to-human transmission has been demonstrated [9]. In humans, NDV has been identified in washings of the conjunctiva sac, nasal discharge, saliva, blood samples, and urine. Thus, ND could be of greater importance to humans than is currently appreciated [10].
Similar to other African nations, ND results in considerable losses to poultry in Tanzania [11]. In Tanzania, 86% of livestock-keeping families keep chickens, making them the most frequent type of livestock [12]. Although the adoption of vaccination against ND with LaSota and I-2 vaccines has become increasingly common in recent years on traditional farms in Tanzania [13], frequent outbreaks still occur in backyard poultry farms (BPFs) [14].
Due to the continuous genetic evolution and wide circulation of NDV in poultry populations, and multiple transmissions in poultry, the emergence of novel genetic NDV variants is ongoing. For these reasons, this study aimed to investigate the phylogenetic and genetic characteristics of NDV isolates from Iringa (Southern highlands zone) and Morogoro (Eastern zone) in January 2022 in Tanzania. These data facilitate tracking NDV evolution and genetic diversity and improving disease surveillance in the country. Furthermore, genetic sequences of NDV can be used in the design and future development of genotype-matched vaccines to enhance the control of NDV in Tanzania and neighboring countries.
2 Materials and methods
2.1 Sampling
In 2022, two municipalities in different regions of Tanzania, namely Iringa (Southern highlands zone) and Morogoro (Eastern zone) (Fig. 1) in Tanzania reported suspected ND outbreaks. Proventriculus tissues collected from two dead vaccinated chickens showed lesions consistent with NDV infection during postmortem examination.
A cartographic representation of Tanzania, indicating the provenance of the samples tested in this research. The samples were procured from Nduli in the Iringa district, as well as Kihonda in the Morogoro district. The image was generated using QGIS software iteration 3.24.3, leveraging data from the DIVA-GIS repository accessible at https://www.diva-gis.org/Data
2.2 Virus isolation
The virus was isolated according to procedures detailed in prior investigations [15] consistent with the guidelines provided by WOAH [16]. Ten-day-old, specific-pathogen-free (SPF) embryonated chicken eggs (ECEs) were treated with 100 µL of clarified tissue homogenate in their allantoic cavities. Control eggs were treated with sterile saline. Eggs were incubated at 37 °C for 4–5 days. Eggs were candled twice daily, and the embryos were visually examined to determine their viability. Any embryos that perished within 24 h post-infection (PI) were excluded from the study, as their deaths were attributed to non-specific causes. Those embryos that survived longer than 24 h or the entire incubation period were cooled at 4 °C overnight before harvesting the allantoic fluid (AF).
2.3 Hemagglutination assay
A hemagglutination (HA) assay using 1% chicken red blood cells was used to test the extracted AF from each infected egg using established protocols [16]. HA-negative samples were passaged for an additional two passages in ECEs, followed by an HA assay to confirm the absence of hemagglutination patterns. HA-positive samples were further tested for NDV by reverse transcription polymerase chain reaction (RT-PCR).
2.4 Reverse transcription polymerase chain reaction (RT-PCR)
RNA was extracted using the QIAamp Viral RNA Extraction Kit (Qiagen, California, USA) per the manufacturer's instructions. We generated cDNA from purified RNA using the SuperScript III First-Strand Synthesis System (InvitrogenTM, CA, USA). Briefly, 8 µL of RNA was mixed with 1 µL of random hexamer primers (50 ng/μL) and 1 µL of dNTPs mix (10 mM). The mixture was then incubated at 65 °C for 5 min to denature RNA secondary structures and cooled on ice for five minutes. We then added 2 µL of 10 × RT buffer, 4 µL of 25 mM MgCl2, 2 µL of 0.1 M Dithiothreitol (DTT), 1 µL of RNAseOUT (40 U/µL), and 1 µL of SuperScript III RT enzyme (200 U/µL). The mixture was incubated at 25 °C for 5 min, followed by incubation at 50 °C for 50 min and at 85ºC for 5 min to terminate the reaction. After cooling on ice, 1 µL of RNase H was added, and the mixture was incubated at 37 °C for 20 min. The final cDNA products were stored at -80 °C until further analysis. RT-PCR was conducted on the two HA-positive isolates. The initial step involved cDNA amplifications using a SuperScript III RT kit (Invitrogen, Valencia, CA, USA) as per the manufacturer's guidelines with specific primers designed for the F and HN genes [17]. The reactions underwent an initial denaturation at 95 ºC for 10 min, followed by 35 cycles consisting of 30 s at 95 ºC for denaturation, 1 min for annealing, and 2 min at 68 ºC for extension. The final extension step was carried out at 68 ºC for 10 min. The primer annealing temperatures were set at 58 ºC for the F gene and 56 ºC for the HN gene. Subsequently, electrophoresis was used to separate the PCR products on a 1.5% agarose gel stained with GelRed (Phenix, Hong Kong, China).
2.5 Sequencing, sequence assembly, alignment, and phylogeny
Dideoxynucleotide Sanger sequencing of PCR products was conducted at Macrogen Laboratory (Macrogen Inc., Amsterdam, The Netherlands) on an ABI 3730xl DNA analyzer (Applied Biosystems, Foster City, CA).. Sequence Scanner Software v2.0 (appliedbiosystems) was used to check the quality of the chromatograms and Bioedit Sequence Alignment Editor v7.2.5.0 [18] was used to assemble and edit the obtained nucleotide sequences. The obtained sequences and those found in the NCBI nucleotide database were compared using the NCBI BLAST tool. Open reading frames (ORF) were predicted using the GENEQUEST program of DNASTAR Lasergene 17 (version 17.5.0.48; Madison, WI). Nucleotide alignment was performed by the ClustalW method built in MEGAX software [19]. Comparisons were made between the sequences from this work and reference sequences from the NCBI database. The reference sequences selected for phylogenetic analysis were chosen according to a set of criteria designed to capture global genetic diversity. These criteria included representation from diverse geographic regions, strains from different genotypes, availability of complete or near-complete genome sequences, and sourcing from reputable databases with comprehensive metadata. The full nucleotide sequence of the F and HN genes was used as the basis for a phylogenetic analysis using MEGA X software. The Maximum Likelihood method utilized the General Time-Reversible (GTR) model incorporating a discrete gamma distribution (+ G) and permitting invariant sites (+ I). The statistical analysis was based on 1000 bootstrap re-sampling [19]. Analysis of amino acid substitutions in F and HN proteins were assessed [20]. The sequences derived from this investigation were uploaded to the NCBI GenBank with accession numbers PP195786 for the HN gene and PP216548 and PP216549 for the F genes.
3 Results
3.1 Virus isolation and confirmation
The inoculated embryos showed extensive hemorrhage and congestion and died within 48–96 h after the first passage (Fig. 2). AF from the inoculated ECE was examined for hemagglutinating activity by HA followed by confirmation using RT-PCR. Both samples tested positive for hemagglutination activity and were positive for the presence of F and HN genes of NDV by RT-PCR, confirming the presence of NDV in the two isolates.
Findings from the isolation of NDV in the allantoic cavities of chicken embryos that were 10 days old. A The embryo that was injected with the negative control mixture. B An infected embryo with a positive sample that exhibits dwarfism, aberrant feathering, and subcutaneous hemorrhages throughout the body
3.2 Sequence and phylogenetic analyses
We obtained two full F gene sequences from Iringa (Iri1921; accession number PP216548) and Morogoro (Mor1221; accession number PP216549). Both sequences were 1662 bp in length.
The nucleotide assembly sequences generated for the HN genes of our two isolates yielded a single good-quality result, namely one from the Morogoro isolate (Mor1221). We were unable to obtain the HN gene for the second isolate from Iringa (Iri1921). The full HN gene of the Morogoro strain (Mor1221) has the accession number PP195786 (1755 bp in length).
3.2.1 F gene
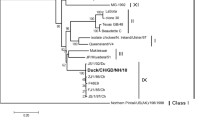
A phylogenetic analysis of the F genes of strains Iri1921 and Mor1221 showed that both viruses clustered within sub-genotype VII.2 (Fig. 3). Based on sequence similarity, NDV strains previously isolated from Africa and Asia were closely related to the two strains presented here (Table 1). The sequence at the cleavage site suggested that both viruses are virulent due to the polybasic amino acid motif 112RRQKRF117, typical of virulent viruses [6].
Phylogenetic analysis using the whole NDV strain F gene sequences. The two nucleotide sequences that are the subject of this investigation are designated in red (subgenotypes VII.2) and indicated with ●. We included 94 class II sub-genotype sequences. The sequences were aligned using ClustalW in MEGA X. We reconstructed a phylogenetic tree using the maximum likelihood approach with 1000 bootstraps. To estimate the evolutionary history while taking into consideration invariant sites (+ I), the general time-reversible (GTR) model with a discrete Gamma distribution (+ G). The first, second, third, and noncoding codon locations are all included in the inquiry. The names given to each sequence in the phylogenetic tree correspond to the GenBank accession number, followed by the sub-genotype, isolation country, strain identifier, and isolation year
3.2.2 HN genes
A phylogenetic analysis of the HN gene of virus strain Mor1221 revealed that it clustered within sub-genotype VII.2 (Fig. 4). The MR1221 HN was closely related to NDV strains from Southeast Asia and Africa (Table 2).
Phylogenetic analysis using the whole NDV strain HN gene sequence. MR1221 HN is designated in red (subgenotypes VII.2) and indicated with ●. We used 87 class II sub-genotype sequences. The sequences were aligned using ClustalW in MEGA X. We reconstructed a phylogenetic tree using the maximum likelihood approach with 1000 bootstraps. To estimate the evolutionary history while taking into consideration invariant sites (+ I), the general time-reversible (GTR) model with a discrete Gamma distribution (+ G). The first, second, third, and noncoding codon locations are all included in the inquiry. The names given to each sequence in the phylogenetic tree correspond to the GenBank accession number, followed by the sub-genotype, isolation country, strain identifier, and isolation year
3.3 Analyze of the amino acid substitution of F and HN proteins
Amino acid substitution analysis of the F (Fig. 5A) and HN (Fig. 5B) proteins of Tanzanian subgenotype VII.2 in the current study was conducted in comparison with some African strains (Uganda, Zambia, Mozambique, South Africa, Botswana, and Zimbabwe) of NDV under genotype VII. This analysis revealed strong conservation in the F and HN protein sequence regions of the Tanzanian NDVs identified in the current study (shown in italics) with other strains.
4 Discussion
We obtained HN and F gene sequences of NDV isolates from BPFs in Tanzania in 2022. To improve knowledge of the distribution and genetic variability of NDV in Tanzania, we characterized the sequences phylogenetically. The viruses sequenced here were categorized as subgenotype VII.2 according to the classification scheme suggested by Dimitrov et al. [31]. We added one complete HN gene and two complete F gene sequences to the GenBank database. Both viruses identified are closely related to previously reported isolates from Mozambique (KU523528) based on the F gene. Sequencing of the HN gene showed similarities with other sequences previously isolated in Asian countries such as Malaysia, China, and Indonesia.
Previous studies have suggested that subgenotype VII strains isolated in southern Africa originated in Asia, which could explain the similarity shared by the Tanzania HN gene with strains identified in Southeast Asian countries [32, 33]. This provides more evidence that genotype VII viruses in southern Africa originated in Southeast Asia.
Since its first detection in Mozambique in 2011 [30], subgenotype VII has spread to Zimbabwe, Malawi, Zambia, South Africa, and Botswana. Before its appearance in South Africa and Mozambique [21], sub-genotype VII.2 viruses, which have the potential to cause ND panzootics, were endemic in some Southeast Asian countries, such as Indonesia [26], Malaysia [34], Cambodia [35], and Vietnam [36]. In recent years, this sub-genotype has spread to many parts of Tanzania [15, 37]. Sub-genotype VII.2 was recently identified in a live bird market in Zambia and was related to strains previously isolated in other southern African nations [32]. Notably, several sub-lineages of genotype VII have appeared in the Far East and spread to other geographical areas in South America [38, 39], Europe [40, 41], Asia [24, 25, 42, 43] and Africa [15, 30, 37, 44]you. This suggests global circulation of NDV sub-genotype VII.2.
ND outbreaks in vaccinated commercial flocks caused by genotype VII have resulted in up to 60–80% of deaths in Indonesia [45], and Japan [46, 47], indicating that more extensive NDV surveillance is needed. Since 2020, a new, rigorous program for monitoring vaccinations to combat several animal diseases, including NDV, has been in place in Tanzania. This program ensures compliance with a vaccination schedule for NDV and manages the quality and pricing of vaccines on the market [48, 49]. However, despite these measures, breeders still suffer outbreaks of ND, causing huge losses in their flocks.
The numerous NDV outbreaks in vaccinated chickens necessitate ongoing observation of the evolution and transmission of NDV. In addition, ongoing assessment of ND vaccination effectiveness is required to prevent future outbreaks. Several factors may be responsible for vaccine failure in poultry [50]: problems associated with the vaccine (inactivation of the vaccine due to expiration, antigenic differences between existing vaccine and field strains, inadequate level of protection; factors associated with the host/birds (stress factors, interference with maternal antibodies/immunity, immunosuppressive and coexisting diseases, genetic factor); factors associated with administration of vaccines (inappropriate route of administration, insufficient dosage, improper formulation of vaccine and diluent used). Notably, both strains identified here were from vaccinated chickens, highlighting the ineffectiveness of current vaccination strategies. The results of the alignment of amino acid substitutions in the F and HN proteins of subgenotype VII.2 present a compelling rationale for considering this subgenotype as a potential vaccine target. This is particularly true given that conserved regions could be targeted by vaccines [51].
5 Conclusions
Our study's results imply that BPFs in the Morogoro and Iringa regions of Tanzania are home to virulent genotype VII NDV. As the chickens tested in this study had been vaccinated, these results suggest that current vaccines are ineffective at preventing severe ND. Regular surveillance will lead to improved disease control and a better understanding of the ND status, characterization of circulating strains.
Data availability
The nucleotide sequences generated in this study are available at NCBI GenBank under Accession Numbers PP216548 and PP216549 for the F genes and PP195786 for the HN gene.
Abbreviations
- AOaV-1:
-
Avian Orthoavulavirus 1
- ND:
-
Newcastle disease
- NDV:
-
Newcastle disease virus
- OIE:
-
Office International des Epizooties
- WOAH:
-
World Organisation for Animal Health
- ACDPa:
-
Advisory Committee on Dangerous Pathogens
- BPFs:
-
Backyard poultry farms
- SPF:
-
Specific-pathogen-free
- ECEs:
-
Embryonated chicken eggs
- PBS:
-
Phosphate-buffered saline
- HA:
-
Hemagglutination assay
- RNA:
-
Ribonucleic acid
- cDNA:
-
Complementary deoxyribonucleic acid
- RT-PCR:
-
Reverse transcription polymerase chain reaction
References
Roky SA, Das M, Akter S, Islam A, Paul S. Determinants of Newcastle disease in commercial layer chicken farms in two districts of Bangladesh: a case-control study. Heliyon. 2022;8.
Walker PJ, Siddell SG, Lefkowitz EJ, Mushegian AR, Dempsey DM, Dutilh BE, et al. Changes to virus taxonomy and the International Code of Virus Classification and Nomenclature ratified by the International Committee on Taxonomy of Viruses (2019). Arch Virol. 2019;164:2417–29.
Bahoussi AN, Shah PT, Zhao JQ, Wang PH, Guo YY, Wu C, et al. Multiple potential recombination events among Newcastle disease virus genomes in China between 1946 and 2020. Front Vet Sci. 2023;10:1136855.
Lu X, Wang X, Liu X, Liu X. The multifaceted interactions between Newcastle disease virus proteins and host proteins: a systematic review. Virulence. 2024. https://doi.org/10.1080/21505594.2023.2299182.
Zhang G, Xue J, Li X. The influence of Newcastle disease virus major proteins on virulence. Vet Sci Res. 2021;3:1–6.
Lu X, Liu X, Song Q, Wang X, Hu S, Liu X. Amino acid mutations in hemagglutinin-neuraminidase enhance the virulence and pathogenicity of the genotype III Newcastle disease vaccine strain after intravenous inoculation. Front Vet Sci. 2022. https://doi.org/10.3389/fvets.2022.890657.
Suarez DL, Miller PJ, Koch G, Mundt E, Rautenschlein S. Newcastle disease, other avian paramyxoviruses, and avian metapneumovirus infections. In: Swayne DE, Boulianne M, Logue CM, McDougald LR, Nair V, Suarez DL, Wit S, Grimes T, Johnson D, Kromm M, Prajitno TY, Rubinoff I, Zavala G, editors. Diseases of poultry. Wiley; 2019. p. 109–66. https://doi.org/10.1002/9781119371199.ch3.
Ul-Rahman A, Ishaq HM, Raza MA, Shabbir MZ. Zoonotic potential of Newcastle disease virus: old and novel perspectives related to public health. Rev Med Virol. 2022. https://doi.org/10.1002/rmv.2246.
Kuiken T, Buijs P, Van Run P, Van Amerongen G, Koopmans M, Van Den Hoogen B. Pigeon paramyxovirus type 1 from a fatal human case induces pneumonia in experimentally infected cynomolgus macaques (Macaca fascicularis). Vet Res. 2017;48:1–9.
Prajna NV, Lalitha P, Chen C, Zhong L, Lietman TM, Doan T, et al. Acute keratoconjunctivitis resulting from coinfection with avian Newcastle virus and human adenovirus. Cornea. 2022;41:630–1.
Mngumi EB, Bunuma E. Seroprevalence and risk factors of Newcastle disease virus in local chickens in Njombe and Bahi districts in Tanzania. Trop Anim Health Prod. 2022;54:53.
Silva M Da, Desta S, Stapleton J. Development of the chicken sector in the Tanzanian livestock master plan. Tanzania Livestock Master Plan Brief. 2017;7 October:1–4.
Campbell ZA, Marsh TL, Mpolya EA, Thumbi SM, Palmer GH. Newcastle disease vaccine adoption by smallholder households in Tanzania: identifying determinants and barriers. PLoS ONE. 2018;13:1–16.
Amoia CFANG, Nnadi PA, Ezema C, Couacy-Hymann E. Epidemiology of Newcastle disease in Africa with emphasis on Côte d’Ivoire: A review. Vet World. 2021;14:1727–40.
Amoia CF, Hakizimana JN, Duggal NK, Chengula AA, Rohaim MA, Munir M, et al. Genetic diversity of Newcastle disease virus involved in the 2021 outbreaks in backyard poultry farms in Tanzania. Vet Sci. 2023;10:477.
OIE, 2012. Manual of diagnostic tests and vaccines for terrestrial animals: mammals, birds and bees. In: Biological Standards Commission. World Organization for Animal Health, Paris, pp. 1–19.
Murulitharan K, Yusoff K, Omar AR, Molouki A. Characterization of Malaysian velogenic NDV strain AF2240-I genomic sequence: a comparative study. Virus Genes. 2013;46:431–40.
Hall TA. BIOEDIT: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/ NT. Nucleic Acids Symp Ser. 1999;41:95–8.
Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 2018;35:1547–9.
Omony JB, Wanyana A, Mugimba KK, Kirunda H, Nakavuma JL, Otim-Onapa M, et al. Epitope peptide-based predication and other functional regions of antigenic F and HN proteins of waterfowl and poultry avian avulavirus serotype-1 isolates from Uganda. Front Vet Sci. 2021;8: 610375.
Mapaco LP, Monjane IVA, Nhamusso AE, Viljoen GJ, Dundon WG, Achá SJ. Phylogenetic analysis of Newcastle disease viruses isolated from commercial poultry in Mozambique (2011–2016). Virus Genes. 2016;52:748–53.
Mubamba C, Gummow DBRWRHB, Bisschop GDSPR, Abolnik C, Mubamba C, Wandrag DBR, et al. Tracing the origins of genotype VIIh Newcastle disease in southern Africa. Transbound Emerg Dis. 2018;65:e393-403.
Kgotlele T, Modise B, Nyange JF, Thanda C, Cattoli G, Dundon WG. First molecular characterization of avian paramyxovirus-1 (Newcastle disease virus) in Botswana. Virus Genes. 2020;56:646–50.
Syamsiah AS, Leow BL, Faizul FMY, Muhammad RS, Faizah HMS. Identification of Newcastle disease virus sub-genotype VII 1.1 isolated from chickens in Sabah, Malaysia. Trop Biomed. 2022;39:579–86.
Liu H, Wang J, Ge S, Lv Y, Li Y, Zheng D, et al. Molecular characterization of new emerging sub-genotype VIIh Newcastle disease viruses in China. Virus Genes. 2019;55:314–21.
Xiao S, Paldurai A, Nayak B, Samuel A, Bharoto EE, Prajitno TY, et al. Complete genome sequences of Newcastle disease virus strains circulating in chicken populations of Indonesia. J Virol. 2012;86:5969–70.
Satharasinghe DA, Murulitharan K, Tan SW, Yeap SK, Munir M, Ideris A, et al. Detection of inter-lineage natural recombination in avian paramyxovirus serotype 1 using simplified deep sequencing platform. Front Microbiol. 2016;7(NOV):1–14.
Song HS, Kim HS, Kim JY, Kwon YK, Kim HR. The development of novel reverse transcription loop-mediated isothermal amplification assays for the detection and differentiation of virulent Newcastle disease virus. Int J Mol Sci. 2023;24:13847.
Kariithi HM, Volkening JD, Chiwanga GH, Goraichuk IV, Olivier TL, Msoffe PLM, et al. Virulent Newcastle disease virus genotypes V.3, VII.2, and XII.1.1 and their coinfections with infectious bronchitis viruses and other avian pathogens in backyard chickens in Tanzania. Front Vet Sci. 2023;10:1272402.
Abolnik C, Mubamba C, Wandrag DBR, Horner R, Gummow B, Dautu G, et al. Tracing the origins of genotype VIIh Newcastle disease in southern Africa. Transbound Emerg Dis. 2018;65:e393-403.
Dimitrov KM, Abolnik C, Afonso CL, Albina E, Bahl J, Berg M, et al. Updated unified phylogenetic classification system and revised nomenclature for Newcastle disease virus. Infect Genet Evol. 2019;74: 103917.
Kalonda A, Saasa N, Kajihara M, Nao N, Moonga L, Ndebe J, et al. Phylogenetic analysis of Newcastle disease virus isolated from poultry in live bird markets and wild waterfowl in Zambia. Microorganisms. 2024;12:354.
Tsaxra JB, Abolnik C, Kelly TR, Chengula AA, Mushi JR, Msoffe PLM, et al. Molecular characterization of Newcastle disease virus obtained from Mawenzi live bird market in Morogoro, Tanzania in 2020–2021. Braz J Microbiol. 2023;54:3265–73.
Berhanu A, Ideris A, Omar AR, Bejo MH. Molecular characterization of partial fusion gene and C-terminus extension length of haemagglutinin-neuraminidase gene of recently isolated Newcastle disease virus isolates in Malaysia. Virol J. 2010. https://doi.org/10.1186/1743-422X-7-183.
Choi KS, Kye SJ, Kim JY, Damasco VR, Sorn S, Lee YJ, et al. Molecular epidemiological investigation of velogenic Newcastle disease viruses from village chickens in Cambodia. Virus Genes. 2013;47:244–9.
Choi KS, Kye SJ, Kim JY, To TL, Nguyen DT, Lee YJ, et al. Molecular epidemiology of Newcastle disease viruses in Vietnam. Trop Anim Health Prod. 2014;46:271–7.
Kibasa MI. Epidemiology of Newcastle disease in backyard chickens rearing system in Iringa rural district, Tanzania. 2020.
Absalón AE, Cortés-Espinosa DV, Lucio E, Miller PJ, Afonso CL. Epidemiology, control, and prevention of Newcastle disease in endemic regions: Latin America. Trop Anim Health Prod. 2019;51:1033–48.
Fernández-Díaz M, Montalván-Avalos A, Isasi-Rivas G, Villanueva-Pérez D, Quiñones-Garcia S, Tataje-Lavanda L, et al. Draft genome sequence of an isolate of genotype VII Newcastle disease virus isolated from an outbreak in fighting cock in Peru. Microbiol Resour Announc. 2023. https://doi.org/10.1128/mra.01293-22.
Ross CS, Skinner P, Sutton D, Mayers J, Nunez A, Brookes SM, et al. Game birds can act as intermediaries of virulent genotype VII avian orthoavulavirus-1 between wild birds and domestic poultry. Viruses. 2023. https://doi.org/10.3390/v15020536.
Steensels M, Van Borm S, Mertens I, Houdart P, Rauw F, Roupie V, et al. Molecular and virological characterization of the first poultry outbreaks of Genotype VII.2 velogenic avian orthoavulavirus type 1 (NDV) in North-West Europe, BeNeLux, 2018. Transbound Emerg Dis. 2021;68:2147–60.
Molouki A, Sotani M, Fallah Mehrabadi MH, Shoushtari A, Abtin A, Mahmoudzadeh Akhijahani M, et al. Predominance of fourth panzootic newcastle disease virus subgenotype VII.1.1 in Iran and its relation to the genotypes circulating in the region. Curr Microbiol. 2021;78:3068–78.
Nooruzzaman M, Hossain I, Begum JA, Moula M, Khaled SA, Parvin R, et al. The first report of a virulent newcastle disease virus of genotype VII.2 causing outbreaks in chickens in Bangladesh. Viruses. 2022. https://doi.org/10.3390/v14122627.
da Silva AP, Aston EJ, Chiwanga GH, Birakos A, Muhairwa AP, Kayang BB, et al. Molecular characterization of newcastle disease viruses isolated from chickens in Tanzania and Ghana. Viruses. 2020. https://doi.org/10.3390/v12090916.
Putri DD, Handharyani E, Soejoedono RD, Setiyono A, Mayasari NLPI, Poetri ON. Pathotypic characterization of Newcastle disease virus isolated from vaccinated chicken in West Java, Indonesia. Vet World. 2017;10:438–44.
Umali DV, Ito H, Suzuki T, Shirota K, Katoh H, Ito T. Molecular epidemiology of Newcastle disease virus isolates from vaccinated commercial poultry farms in non-epidemic areas of Japan. Virol J. 2013;10:1–17.
Abd-Ellatieff HA, Abd El Aziem AN, Elbestawy AR, Goda WM, Belih SS, Ellakany HF, et al. Efficacy of vaccination against infection with velogenic newcastle disease virus genotypes VI and VII 1.1 strains in Japanese Quails. J Comp Pathol. 2021;186:35–50.
Enahoro D, Galiè A, Abukari Y, Chiwanga GH, Kelly TR, Kahamba J, et al. Strategies to upgrade animal health delivery in village poultry systems: perspectives of stakeholders from Northern Ghana and Central Zones in Tanzania. Front Vet Sci. 2021. https://doi.org/10.3389/fvets.2021.611357.
Lindahl JF, Young J, Wyatt A, Young M, Alders R, Bagnol B, et al. Do vaccination interventions have effects? A study on how poultry vaccination interventions change smallholder farmer knowledge, attitudes, and practice in villages in Kenya and Tanzania. Trop Anim Health Prod. 2019;51:213–20.
Fesseha H. Vaccine failure in poultry production and its control methods: a review. Biomed J Sci Tech Res. 2020. https://doi.org/10.26717/BJSTR.2020.29.004827.
Wu KW, Chien CY, Li SW, King CC, Chang CH. Highly conserved influenza A virus epitope sequences as candidates of H3N2 flu vaccine targets. Genomics. 2012;100:102–9.
Acknowledgements
The authors are particularly grateful to the Tanzania Veterinary Laboratory Agency at Iringa and Morogoro regions for their skilful assistance in the field. They also thank Jonas C. Fitwangile for his technical assistance in the laboratory.
Funding
C.FA. was awarded a Regional Scholarship and Innovation Fund (RSIF), through the Partnership for Skills in Applied Sciences, Engineering and Technology (PASET) in order to complete doctoral studies at SACIDS Africa Centre of Excellence for Infectious Diseases, SACIDS Foundation for One Health, Sokoine University of Agriculture, Morogoro, Tanzania. South African National Research Foundation (NRF) and Tanzania Commission for Science and Technology (COSTECH) have jointly funded G.M's OR Tambo Africa Research Chair for Viral Epidemics.
Author information
Authors and Affiliations
Contributions
Conceptualization, C.F.A., A.A.C, J.W-L. and G.M.; methodology, C.F.A., J.N.H., A.A.C., M.R.M., J.W-L. and G.M.; software, C.F.A., J.N.H; validation: A.A.C., J.W-L. and G.M; formal analysis: C.F.A., J.N.H. and M.R.M.; data curation: C.F.A.; writing—original draft preparation: C.F.A.; writing—review and editing: C.F.A., J.N.H., A.A.C., M.R.M., J. W-L. and G.M.; Supervision, A.A.C., J.W-L. and G.M. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Ethical clearance was obtained from the Office of the Vice Chancellor, Sokoine University of Agriculture, Tanzania (Ref. No. SUA/ADM/R.1/8A/737) and Permission to undertake animal health research in Tanzania from the United Republic of Tanzania Ministry of Livestock and Fisheries (Ref. No. DB.16/324/01/137).
Consent for publication
Not applicable.
Competing interests
The authors declare no conflict of interest. The funder had no role in the design of the study; in the analyses, and interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Amoia, C.F., Hakizimana, J.N., Chengula, A.A. et al. Phylogenetic analysis of virulent strains of the Newcastle disease virus isolated from deceased chickens in Tanzania's Morogoro and Iringa regions. Discov Anim 1, 10 (2024). https://doi.org/10.1007/s44338-024-00015-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44338-024-00015-0