Abstract
Petal senescence refers to the progressive loss of intracellular structures and functions within plant decorative organs, ultimately leading to cell death. Autophagy involves the degradation of damaged cellular components and nutrient recycling. Plant organ senescence and autophagy are highly coordinated; however, the mechanisms by which autophagy regulates petal senescence remain largely unknown. In this study, by using transmission electron microscopy, we observed that autophagic activity peaked early, at flower opening, without any senescence and other morphological symptoms in petals. We found that darkness positively regulated petal senescence and upregulated autophagy-related genes (ATGs). Dark treatment promoted the accumulation of Rosa hybrida phytochrome-interacting factor 4 (RhPIF4) in petals. RhPIF4 silencing delayed petal senescence and repressed the expression of ATGs. In contrast, silencing of the light-responsive gene Rosa hybrida elongated hypoctyl 5 (RhHY5) promoted petal senescence and ATG gene expression. RhPIF4/8 and RhHY5 could directly interact with RhWRKY40, and RhWRKY40 is directly bound to the promoters of RhATG7 and RhATG11. Silencing RhWRKY40 delayed petal senescence and suppressed RhATG7 and RhATG11 expression. Based on these results, we propose that RhPIF4/8 and RhHY5 transcription factors are involved in regulating petal senescence in response to dark or light conditions by modulating autophagic activity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Senescence is the final stage of plant organ development and involves the degradation of macromolecules and membrane systems, and the recycling of cellular constituents and metabolites (Ma et al. 2018; Woo et al. 2019). Autophagy is an intracellular degradation process closely associated with cellular senescence, where unnecessary or dysfunctional cytoplasmic components are broken down within the vacuole (Üstün et al. 2017). Under normal conditions, autophagy occurs at basal levels to maintain cellular homeostasis (Su et al. 2020) and is induced under stress conditions to protect cells from death (Chi et al. 2020; Wang et al. 2015, 2019; Zhang et al. 2020; Zhu et al. 2018). However, overactivation of autophagy has pro-death functions (van Doorn and Woltering 2005).
The process of autophagy is well-established in plants (Avila-Ospina et al. 2014). Autophagy is initiated by the formation of autophagosomes, double-membrane vesicles that envelop cellular constituents during macroautophagy (Shibutani and Yoshimori 2014). The fusion of the autophagosome's outer membrane with the tonoplast releases single-membrane vesicles called autophagic bodies into the vacuole, where they undergo degradation. The core autophagy components, encoded by autophagy-related genes (ATGs), are conserved across plant species and critical in regulating autophagy processes (Li et al. 2015; Xia et al. 2011; Zhou et al. 2015). ATGs are divided into four functional groups: the ATG1/ATG13 kinase complex, the autophagy-specific phosphatidylinositol 3-kinase (PI3K) complex, the ATG9 cycling complex, and the ATG8 and ATG12 ubiquitin-like conjugation systems (Yang et al. 2020a). ATG8, a ubiquitin-like protein, is used as an autophagosome marker in yeast, mammals, and plants (Chung et al. 2010; Rubinsztein et al. 2009). ATG5 and ATG7 are considered the core components of autophagy induction (Shimizu et al. 2010). Notably, inhibiting autophagy either biochemically or in ATG5/ATG7-RNAi lines significantly suppressed strawberry fruit ripening (Sánchez-Sevilla et al. 2021). ATG11, a scaffold protein, is essential for selective autophagy and is necessary to initiate glucose starvation-induced autophagy (Yao et al. 2020). The expression of ATG11 gradually increased and peaked during the final stage of leaf senescence, suggesting that mitophagy may be significantly induced to accelerate and complete the senescence process (Broda et al. 2018). Regarding the flowers, autophagic structures have been observed in the petals across different species, such as Ipomoea nil and Dendrobium (Shibuya et al. 2009; van Doorn et al. 2015). The induction of PhATG8 homologs in petunias leading to petal senescence is initiated by pollination (Shibuya et al. 2013). EPHEMERAL1 (EPH1)-RNAi suppresses autophagy and delays petal senescence in Ipomoea nil (Shibuya et al. 2014). However, the molecular mechanisms underlying the ATGs' involvement and functions in petal senescence remain unclear.
Darkness is an important external cue that triggers senescence and autophagy (Song et al. 2014; Yang et al. 2020b). The light response factor elongated hypoctyl 5 (HY5) inhibits autophagy by recruiting histone deacetylase 9 (HDA9) to ATG5 and ATG8e loci (Yang et al. 2020b). On the other hand, under dark conditions, HY5 degradation results in the removal of HDA9 from ATG5 and ATG8e, activating ATG gene expression (Yang et al. 2020b). Phytochrome-interacting factors (PIFs) are essential regulators in response to darkness, that accumulate under dark conditions and are degraded upon exposure to light (Al-Sady et al. 2006; Nozue et al. 2007; Shen et al. 2008). The pif3, pif4, and pif5-2 mutants exhibit increased leaf longevity in normal and dark conditions (Song et al. 2014). However, whether PIF genes play a role in darkness-induced autophagy remains unclear.
This study provides evidence for autophagy induction during the early stages of petal senescence. The results highlight the critical role of RhPIF4/8 and RhHY5 in mediating darkness-induced acceleration of autophagy as a mechanism for regulating petal senescence.
Materials and methods
Plant materials and treatments
Flowers of cut roses (Rosa hybrida 'Samantha') grown in a greenhouse were used for sampling. The tobacco (Nicotiana benthamiana) and rose plants were grown at 22 ± 1°C and 50% relative humidity under long-day conditions (16 h light/8 h dark). The light and dark treatments were according to a previous study (Zhang et al. 2021). The flowers were exposed for 2 days to light or dark conditions.
Transmission electron microscopy (TEM) analysis
Samples (2 mm × 2 mm) from rose petals were used to visualize the accumulation of autophagosomes/autophagic bodies. Samples were fixed with 0.1 M paraformaldehyde, 2.5% (v/v) glutaraldehyde, 0.2 M PBS buffer (pH 7.2) and then exposed to a vacuum of -25 kPa for 5 min twice. They were then placed at room temperature for 2 h and 4℃ for 24 h for fixing. The ultrathin sections were prepared according to a previous study (Wang et al. 2019). We observed the autophagosomes and autophagic bodies using a Hitachi transmission electron microscope HT7800 at an accelerating voltage of 80 kV.
Protein extraction and western blotting
The RhATG8 and RhATG8-PE proteins were detected by immunoblot assays. Total proteins were extracted from the petals, homogenized using 2 × SDS-PAGE sample buffer (100 mM Tris–HCl, pH 6.8, 4% [w/v] SDS, 20% [v/v] glycerol, and 0.2% [w/v] bromophenol blue). The samples were heated to 95°C for 5 min, and then the supernatants were obtained by centrifugation at 12,000 rpm for 10 min. The ATG8 lipidation assays were performed according to a previous study (Chung et al. 2010). Supernatants were loaded to a 15% SDS PAGE gel containing 6 M urea and immunoblotted with anti-ATG8 antibodies. To assess the RhPIF4 functions in vivo, the 35Spro::RhPIF4-3Flag. 35Spro::RhPIF4-3Flag and 35Spro::GFP constructs were transformed into Agrobacterium tumefaciens, and the respective bacterial suspensions were injected into N. benthamiana leaves (OD600 = 1.0). After 2 d, leaves were treated in dark for 24 h, and samples were collected after 3 d of infiltration. RhPIF4-3Flag was detected by immunoblot assays. The N. benthamiana leaves were homogenized in 2 × SDS-PAGE sample buffer. The GFP protein was used as the control reporter of expression. Anti-ATG8 and anti-FLAG antibodies were obtained from ABclonal (A22294, 1:1000 dilution; AE005, 1:5000 dilution). Anti-GFP antibodies were obtained from Easybio (BE2001, 1: 5000 dilution). Anti-β-Tubulin antibodies were obtained from Abmart (M2000055, 1:5000 dilution). Goat anti-rabbit secondary antibodies and goat anti-mouse secondary antibodies were obtained from Easybio (BE0101, 1:5000 dilution; BE0102, 1:5000 dilution). The primers used are listed in Table S1.
RNA extraction and quantitative reverse transcription-PCR (qRT-PCR)
Total RNA was extracted from rose petals using the hot borate method (Zhang et al. 2019). Reverse transcription was performed with 1 μg total RNA as a template, using the HiScript III All-in-one RT SuperMix Perfect for qPCR (Vazyme, Nanjing, China). The qRT-PCR reactions were carried out in a Step One Plus real-time PCR system (Applied Biosystems). The RhUBI2 and RheIF5A genes were used as the reference genes for normalization to calculate the relative gene expression using the 2−ΔΔCT method. qRT-PCR was performed using gene-specific primers (Table S1).
Transient overexpression (OE) assays in rose petals
In brief, the A. tumefaciens suspensions (OD600 = 1.0) of 35Spro::RhHY5-GFP, 35Spro::RhPIF4-GFP, 35Spro::RhPIF8-GFP were individually injected into the outer petals of stage 2 flowers. The empty vector 35Spro::GFP was used as the negative control. After 2 d of plant growth, RhPIF4-GFP-OE, RhPIF8-GFP-OE, and GFP-OE flowers were placed under constant dark (DD) conditions for 1 d. RhHY5-GFP-OE and GFP-OE flowers were placed under constant light (LL) conditions for 1 d. Gene-specific primers in this assay are listed in Table S1.
Virus-induced gene silencing (VIGS)
Nucleotide sequence fragments corresponding to RhPIF4, RhHY5, and RhWRKY40 were cloned into the tobacco rattle virus (pTRV2) vector for gene silencing, as described in a previous study (Chen et al. 2023). The pTRV2, pTRV2-fusion plasmids, and pTRV1 were introduced individually into A. tumefaciens strain GV3101. Rose plants were immersed in an infiltration buffer containing pTRV1 and pTRV2 (TRV2 control), pTRV1, and each recombinant pTRV2 A. tumefaciens. Then, they were exposed to a vacuum of -25 kPa. Three independent experiments were performed with 35–40 plants in each. The days from fully opening (stage 5) to the complete senescent stage of flowers corresponded to the duration of flower senescence. The gene-specific primers used for the VIGS assays are listed in Table S1.
Yeast assays
Yeast assays were performed as previously described (Chen et al. 2023; Zhang et al. 2021). For yeast one-hybrid (Y1H) assay, AD-RhWRKY40 and RhATG7/11pro::LacZ, AD-RhHY5/RhPIF4/8 and RhATG11pro::LacZ were co-transformed into the yeast strain EGY48. The RhHY5/RhPIF4/8-interacting proteins were screened in the cDNA library (from rose petals) using a yeast two-hybrid (Y2H) system. Different combinations of BD-fusion and AD-fusion plasmids were co-transformed into the yeast strain Y2H Gold. Yeast transformants were selected using SD/-Trp/-Ura agar plates for Y1H assay and SD/-Trp/-Leu agar plates for Y2H assay for 3 days at 30℃. Positive interactors were selected using SD/Gal/Raf/-Trp/-Ura/X-Gal agar plates for the Y1H assay and SD/-Trp/-Leu/-His/3-AT agar plates for the Y2H assay. Yeast assays were performed using gene-specific primers (Table S1).
Luciferase complementation imaging (LCI) assay
Transient LCI assays were carried out in N. benthamiana as previously described (Jiang et al. 2022). RhWRKY40-nLUC and the cLUC-RhHY5/RhPIF4/RhPIF8 recombinant vector were introduced into A. tumefaciens strain GV3101 and suspensions were injected into the N. benthamiana leaves. RhWRKY40-nLUC and cLUC, nLUC and cLUC-RhHY5/RhPIF4/RhPIF8, and nLUC and cLUC were used as negative controls. After three days, LUC signals were imaged after incubating the leaves with the fluorescein substrate (1 mM D-luciferin, Promega). A CCD camera (Berthold, LB985, Germany) was used to capture the LUC signal. LCI assays were performed using gene-specific primers (Table S1).
Co-immunoprecipitation (Co-IP) assays
For the Co-IP assays, different combinations of A. tumefaciens cells harboring 35Spro::RhHY5-3Flag, 35Spro::RhPIF4-3Flag, or 35Spro::RhPIF8-3Flag and 35Spro::RhWRKY40-GFP were co-infiltrated into N. benthamiana leaves. The samples were ground in protein extraction buffer (50 mM Tris–HCl, pH 7.5, 5 mM EGTA, 5% (v/v) glycerol, 10 mM DTT, 1 mM PMSF, 1 × protease inhibitor cocktail (Roche)). DYKDDDDK Fab-Trap agarose (ChromoTek, ffa) was used to bind Flag-tag fusion proteins from supernatants at 4℃, which were then washed five times with wash buffer. Protein extracts were eluted with 2 × SDS loading buffer at 95 ℃ for 5 min. 10% SDS-PAGE gels were used to separate the input and eluted proteins and then exposed to anti-FLAG or anti-GFP antibodies. Co-IP was performed using gene-specific primers (Table S1).
Chromatin immunoprecipitation–qPCR (ChIP-qPCR)
The ChIP-qPCR assay was performed following a previously published method (Cheng et al. 2021). In brief, a 35Spro::RhWRKY40-GFP A. tumefaciens suspension (OD600 = 1.0) was injected into rose petals. The empty vector 35Spro::GFP was used as a negative control. After 3 d of growth, a 2 g petal sample was treated with 1% (v/v) formaldehyde, and the reaction was terminated by a final concentration of 0.125 M glycine. After sonication and centrifugation, the supernatants were incubated with GFP-Trap agarose (ChromoTek, gta) at 4℃. The precipitated DNA was purified using the QIAquick PCR purification kit (Qiagen), and the binding sites were analyzed by qRT-PCR using gene-specific primers (Table S1).
Dual-luciferase reporter assay
The 2000/1100 bp promoter sequences were cloned into pGreenII 0800-LUC vectors to construct the RhATG7pro::LUC and RhATG11pro::LUC reporter vectors. The coding sequences of RhHY5, RhPIF4, and RhPIF8 were inserted into 35Spro::GFP vectors to construct the 35Spro::RhHY5, 35Spro::RhPIF4, 35Spro::RhPIF8 effector vectors. 35Spro::GFP was used as a negative control. The RhATG7pro::LUC and RhATG11pro::LUC reporter vectors were transformed into A. tumefaciens strain GV3101 (pSoup). The mixed A. tumefaciens cultures were injected into young but fully expanded leaves of N. benthamiana. After 2 d, LUC and Renilla luciferase (REN) were detected and quantified using the dual-luciferase reporter assay system (Promega). The dual-LUC assays were performed using gene-specific primers (Table S1).
Results
Autophagic activity peaks at petal senescence onset
To investigate the cellular alterations that occur during petal senescence, we examined the ultrastructure of rose petals at five distinct stages of opening (Ma et al. 2005). Transmission electron microscopy (TEM) depicted several small vacuoles within the epidermal cell cytoplasm in stage 1 (S1) petals, which coalesced into the central vacuole in S2 (Fig. 1a). Further examination revealed a significant reduction in the cytoplasm volume, with enlarged vacuoles observed in the cells in S3 petals (Fig. 1a). In S5, the cells exhibited plasmolysis (Fig. 1a).
Dynamic changes in autophagic activity at different developmental stages of rose petals. a Representative flower phenotypes (upper), ultrastructural characteristics of petal longitudinal section (middle), and autophagic bodies of adaxial epidermal cells (lower) at five stages of flower opening. b Representative transmission electron microscopy (TEM) images of autophagic structures in the adaxial epidermal cells of rose petals. Autophagic bodies are marked with red arrows. M, mitochondrion; V, vacuole. c Autophagic structures from (b) were quantified. The number of autophagic structures in the petals of each stage was quantified to calculate the autophagic activity. Relative autophagic activities in stage 2 (S2) to 5 petals were compared to stage 1 petals, which was set to 1. Autophagic structures were used to quantify autophagic activity in petals of different stages (mean ± SE; n = 20; P<0.05; one-way ANOVA). d Autophagy related gene 8 (ATG8) protein levels in S1, S3 and S5 petals. Free and localized in autophagic structures forms of ATG8 are indicated by ATG8 and ATG8-phosphatidylethanolamine (ATG8-PE). Tubulin was used as the loading control. e Relative ATG8-PE levels in S1, S3 and S5 petals in (d). Protein accumulation was analyzed by ImageJ. Protein accumulation in S1 petals was set to 1 (mean ± SE; P<0.05; one-way ANOVA)
Autophagic bodies in vacuoles were scarce in S1, with a slight increase in S2 petals (Fig. 1b, c). A significant increase in autophagic bodies was observed in S3), which remained at the same levels in S4) and 5 (Fig. 1b, c). The modification of ATG8 by phosphatidylethanolamine (PE) was evaluated during petal senescence to confirm the formation of autophagosomes and autophagic bodies. The ATG8-PE bands notably increased in the S3 and S5 petals compared to those in the S1 petals (Fig. 1d, e). Based on our previous findings which showed that senescence-related genes were upregulated at the stage 3 and onwards (Chen et al. 2023), it was hypothesized that, at the subcellular level, the increase in autophagic activity may be associated with the onset of petal senescence.
Darkness-stabilized RhPIF4 accelerates autophagy
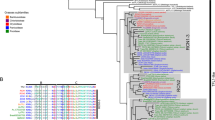
In our previous study, we demonstrated that darkness is a vital factor that aggravates petal senescence (Zhang et al. 2021). To further elucidate the impact of darkness on petal autophagic activity, we first assessed the ATG8-PE bands using western blotting under constant light (LL) and dark (DD) conditions. Notably, ATG8-PE levels exhibited a significant increase in DD compared to LL (Fig. 2a, b). To validate these alterations in petal autophagic activity under dark conditions, we assessed ATG gene expression. We screened the Rosa hybrida 'Samantha' genome, identified all 29 ATG genes (Zhang et al. 2023), and named them based on a phylogenetic analysis (Fig. S1a). RhATG8s were relatively highly expressed in petals, while the transcript levels of RhATG8b/8f notably increased during petal senescence (Fig. S1b). ATG5/7/11 proteins are key components of autophagy induction (Shimizu et al. 2010; Yao et al. 2020). Thus, we chose to investigate the expression levels of five autophagy-related gene members further, RhATG5/7/8b/8f/11, in response to exposure to light and dark conditions. Among the five tested ATG genes, RhATG7/8b/8f/11 were significantly induced in the dark (Fig. 2c).
Effect of darkness on autophagy in rose petals. a Autophagy related gene 8 (ATG8) protein levels in the petals under constant light (LL) and constant dark (DD) conditions. Tubulin was used as the loading control. b Relative autophagy related gene 8-phosphatidylethanolamine (ATG8-PE) levels in LL and DD petals in (a). Protein accumulation was analyzed by ImageJ. Protein accumulation in LL petals was set to 1 (mean ± SE; **P < 0.01; Student's ttest). c Expression of ATGs genes in petals under LL and DD conditions (mean ± SE; n = 3; ns, no significance, *P<0.05, **P<0.01; Student's ttest)
To investigate the role of PIFs in autophagy, we retrieved the sequences of all PIFs present in the genome of the 'Samantha' cultivar (Fig. S2a). Among the five RhPIFs, RhPIF4 and RhPIF8 were relatively highly expressed during petal senescence (Fig. S2b). We have previously demonstrated that RhPIF8 is a negative regulator of petal senescence (Zhang et al. 2021). We analyzed the transcript levels of ATG genes in RhPIF8-overexpressing flowers using qRT-PCR, which revealeda significant decrease in RhATG7/8b/11 expression (Fig. S3).
To determine the function of RhPIF4, we initially examined its expression in response to darkness. We found that RhPIF4 was induced under darkness conditions (Fig. 3a). Additionally, we assessed whether the expression of ATGs was impacted in RhPIF4-overexpressing flowers. In RhPIF4-GFP-overexpressing petals, RhATG7/11 transcript levels were significantly increased, while RhATG8b/f expression remained unchanged compared to GFP-overexpressing petals (Fig. 3b), implying that RhPIF4 is involved in autophagic activity regulation. We then evaluated the influence of RhPIF4 on petal senescence using VIGS. We determined through qRT-PCR that RhPIF4 expression was reduced in petals of VIGS plants compared to the control plants, as depicted in Fig. 3d. RhPIF4-silenced plants exhibited a prolonged senescence phase, lasting an average of 6.4 ± 0.4 d, compared to the control plants, which had an average senescence phase of 4.0 ± 0.3 d (Fig. 3c and e). Furthermore, RhATG7/11 expression was significantly decreased in RhPIF4-silenced lines (Fig. 3f). Therefore, our results suggested that RhPIF4/8 may regulate petal senescence through the modulation of autophagy.
Effect of Rosa hybrida PIFphytochrome-interacting factor 4 (RhPIF4) on autophagy during petal senescence. a RhPIF4 protein levels in petals under constant light (LL) and constant dark (DD) conditions. Samples were analyzed by immunoblotting with anti-FLAG, anti-GFP, and anti-Tubulin antibodies. Tubulin served as the loading control, and GFP was used as the expressing control. b Expression of Rosa hybrida autophagy related genes (RhATGs) and RhPIF4-GFP-overexpressing petals (mean ± SE; n = 3; ns, no significance, **P<0.01, ***P<0.001; Student's ttest). c Flower phenotypes of RhPIF4-silenced plants and tobacco rattle virus (TRV) controls. The flowers were photographed daily. Bar, 1 cm. Experiments were independently performed thrice with similar results. d Expression of RhPIF4 in RhPIF4-silenced petals and the TRV controls (mean ± SE; n = 5; **P<0.01; Student's ttest). e Senescence period duration of RhPIF4-silenced flowers and TRV controls (mean ± SE; n = 7; ***P<0.001; Student's ttest). f Expression of RhATG7/11 in RhPIF4-silenced petals and the TRV controls (mean ± SE; n = 5; **P<0.01; Student's ttest)
Light-responsive RhHY5 suppresses autophagy
To explore the involvement of the light regulator HY5 in autophagy processes in the petals, we retrieved the sequences of RhHY5 and its homologous genes, RhHYH-1 and RhHYH-2, from the rose genome database (Fig. S4a). Notably, RhHY5 exhibited markedly higher expression levels than the RhHYHs in the petals (Fig. S4b). RhHY5 was significantly induced under LL conditions, while its expression was decreased under DD conditions (Fig. 4a). To confirm whether RhHY5 regulates ATG genes, we evaluated the ATG gene expression in petals when RhHY5 is overexpressed transiently. RhATG7/11 expression was significantly decreased, while RhATG8b/8f remained relatively unchanged compared to the control wild-type plants (Fig. 4b). VIGS was subsequently performed to further validate the role of RhHY5 in petal senescence. We confirmed the reduction in RhHY5 expression in RhHY5-silenced petals compared to the TRV controls using qRT-PCR (Fig. 4d). RhHY5-silenced plants presented accelerated senescence (3.1 ± 0.3 d) compared to the TRV controls (4.3 ± 0.3 d) (Fig. 4c, e). Additionally, RhATG7/11 expression was notably increased in RhHY5-silenced petals (Fig. 4f). These findings suggest that RhHY5 may be involved in regulating autophagic activity in the petals under light conditions.
Role of Rosa hybrida elongated hypoctyl 5 (RhHY5) in autophagy-regulated petal senescence. a RhHY5 expression in petals under constant light (LL) and constant dark (DD) conditions (mean ± SE; n = 3; *P<0.05, **P<0.01; Student’s ttest). b Expression of autophagy related genes (ATGs) in GFP- and RhHY5-GFP-overexpressing petals (mean ± SE; n = 3; ns, no significance, *P<0.05, ***P<0.001; Student's ttest). c Flower phenotypes of RhHY5-silenced flowers and the tobacco rattle virus (TRV) controls. The flowers were photographed daily. Bar, 1 cm. Experiments were independently performed thrice with similar results. d RhHY5 expression in RhHY5-silenced petals and the TRV controls (mean ± SE; n = 5; **P<0.01; Student's ttest). e Senescence period duration of RhHY5-silenced flowers and the TRV controls (mean ± SE; n = 7; *P<0.05; Student's ttest). f Expression of RhATG7/11 in RhHY5-silenced petals and the TRV controls (mean ± SE; n = 5; **P<0.01; Student's ttest)
RhPIF4 or RhHY5 interacts with RhWRKY40
Next, we analyzed the RhATG7/11 promoter sequences for the presence of the G-box element (CACGTG), a known binding site for HY5 and PIFs (Burko et al. 2020; Castillon et al. 2007). The RhATG11 promoter contained the G-box element, which was absent from the RhATG7 promoter (Fig. S5a). The direct binding of RhPIF4/8 and RhHY5 to the RhATG11 promoter was evaluated using the Y1H method, revealing that neither was bound to the RhATG11 promoter (Fig. S5b). These findings indicate that RhPIF4/8 and RhHY5 do not play a direct role, by promoter binding, in regulating RhATG7/11.
To identify the proteins that potentially interact with RhPIF4/8 and RhHY5, we conducted a Y2H library screening of the proteins encoded by the three genes individually. Notably, RhWRKY40 emerged as a shared interacting candidate for RhPIF4/8 and RhHY5. To validate this interaction, we introduced the coding sequence of RhWRKY40 into the pGBKT7 (BD-) vector as bait, whereas the RhPIF4/8 and RhHY5 coding sequences were each inserted into the prey pGADT7 (AD-) vector. All three proteins were confirmed to interact with RhWRKY40 based on the Y2H assays (Fig. 5a).
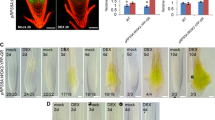
Interactions of Rosa hybrida phytochrome-interacting factor 4 (RhPIF4) or Rosa hybrida elongated hypoctyl 5 (RhHY5) with Rosa hybrid WRKY40 (RhWRKY40). a Interactions between RhWRKY40 and RhPIF4/8, RhHY5, as demonstrated by yeast two-hybrid assays. AD corresponds to the negative control, an empty pGADT7 vector. b Interactions between RhWRKY40 and RhPIF4/RhHY5, were assessed by an luciferase complementation imaging (LCI) assay. c Interactions between RhWRKY40 and RhPIF4/RhHY5 were assessed using co-immunoprecipitation (Co-IP)
To further confirm the interactions between RhWRKY40 and RhHY5/RhPIF4/RhPIF8 in plant cells, we performed LCI and Co-IP assays. RhWRKY40-nLUC and cLUC-RhHY5/RhPIF4/RhPIF8 were co-infiltrated into N. benthamiana leaves, with RhWRKY40-nLUC/cLUC, nLUC/cLUC-RhHY5/RhPIF4/RhPIF8, and nLUC/cLUC as negative controls. After three days, LUC signals were imaged after the incubation with the fluorescein substrate D-luciferin. RhWRKY40-nLUC and cLUC-RhHY5/RhPIF4/RhPIF8 co-expression resulted in robust LUC activity, whereas the controls showed no LUC signal (Fig. 5b, S6a).
Additionally, we performed Co-IP assays following the transient co-infiltration of RhHY5/RhPIF4/RhPIF8-3Flag and RhWRKY40-GFP constructs in N. benthamiana leaves, using RhHY5/RhPIF4/RhPIF8-3Flag and GFP individually as the comparative control. A clear Co-IP of RhHY5/RhPIF4/RhPIF8-3Flag with RhWRKY40-GFP was evident (Fig. 5c, S6b), while no interaction occurred in the control.
RhWRKY40 directly activates the expression of RhATG7/11 genes
We isolated the 2000/1100 bp promoter region upstream of the RhATG7/11 coding sequence, where we identified numerous W-box elements and predicted WRKY binding sites (Eulgem et al. 2000) (Fig. S7). Y1H assays were subsequently performed to determine whether RhWRKY40 could bind to the RhATG7/11 promoters. Based on the results, RhWRKY40 activated LacZ reporter expression under RhATG7/11 promoter control (Fig. 6a). We then performed a dual-LUC reporter assay to analyze the role of RhWRKY40 in RhATG7/11 transcription. Co-infiltration of RhWRKY40 with RhATG7pro::LUC or RhATG11pro::LUC significantly activated LUC reporter expression in the leaves compared to the 35Spro::GFP control (Fig. 6b). These results indicated that RhWRKY40 acts as an activator of RhATG7/11 transcription.
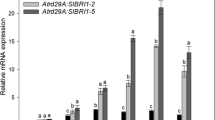
Impact of Rosa hybrid WRKY40 (RhWRKY40) on RhATG7/11 transcript levels. a Binding of the RhWRKY40 protein to the RhATG7/11 promoters, as demonstrated by yeast one-hybrid assays. b Activation of RhATG7/11 by RhWRKY40. Quantitative analysis of the transcriptional activation of the RhATG7/11 promoters by RhWRKY40 (mean ± SE; n = 4; *P<0.05, **P<0.01; Student's ttest). c Chromatin immunoprecipitation-qPCR (ChIP-qPCR) assay demonstrating RhWRKY40 binding to the RhATG7/11 promoters in planta (mean ± SE; n = 3; **P < 0.01; Student's ttest). Petals expressing 35Spro::GFP were used as the negative control. d Dual-luciferase (Dual-LUC) analyses to assess the effects of RhHY5 and RhPIF4 on RhWRKY40 (mean ± SE; n = 3; *P<0.05, **P<0.01; Student's ttest). Leaves expressing RhHY5/RhPIF4 and GFP were used as negative controls
We then examined the in vivo interaction between RhWRKY40 and the RhATG7/11 promoters using ChIP-qPCR. The results verified RhWRKY40 binding to the P3 (-726 to -568 bp) and P1 (-183 to -2 bp) regions of the RhATG7 promoter, and the P2 region (-637 to -449 bp) of the RhATG11 promoter (Fig. 6c). To verify the type of interaction between RhHY5/RhPIF4/RhPIF8 and RhWRKY40, a firefly LUC reporter under each of the RhATG7/11 promoters was co-infiltrated with RhHY5/RhPIF4/RhPIF8 and RhWRKY40 effector constructs in N. benthamiana leaves. Co-transfection of RhHY5 or RhPIF8 with RhWRKY40 suppressed reporter gene expression, whereas co-transfection of RhPIF4 with RhWRKY40 significantly increased LUC activity compared with the control (Fig. 6d, S8). Based on these findings, the RhHY5-RhWRKY40 and RhPIF8-RhWRKY40 modules have a negative regulatory effect on RhATG7/11, while the RhPIF4-RhWRKY40 module acts as an activator.
To investigate the involvement of RhWRKY40 in autophagy-induced petal senescence, we initially evaluated its expression patterns. We found that RhWRKY40 expression increased significantly during senescence and was significantly upregulated under dark conditions (Fig. 7a, b). To explore the involvement and functions of RhWRKY40 on petal senescence, we used VIGS to silence RhWRKY40. Reduced RhWRKY40 expression in RhWRKY40-silenced petals compared to that in the TRV control was confirmed with qRT-PCR (Fig. 7c). The senescence duration in RhWRKY40-silenced petals was extended to 6.6 ± 0.2 d, compared to 4.4 ± 0.4 d in the TRV control (Fig. 7d, e). RhATG7/11 expression was also decreased in RhWRKY40-silenced flowers relative to the controls (Fig. 7f, g), suggesting that RhWRKY40 regulates petal senescence through autophagy modulation.
Phenotype of Rosa hybrid WRKY40 (RhWRKY40)-silenced flowers. a RhWRKY40 expression in petals at the stage 1 (S1), S3 and S5 (mean ± SE; n = 3; *P<0.05, ***P <0.001; Student’s ttest). b RhWRKY40 expression in petals under constant light (LL) and constant dark (DD) conditions (mean ± SE; n = 3; *P <0.05; Student's ttest). c RhWRKY40 expression in RhWRKY40-silenced petals and the TRV controls (mean ± SE; n = 3; *P<0.05; Student's ttest). d Flower phenotypes of RhWRKY40-silenced petals and the tobacco rattle virus (TRV) controls. The flowers were photographed daily. Bar, 1 cm. Experiments were independently performed thrice with similar results. e Senescence period duration of RhWRKY40-silenced flowers and the TRV controls (mean ± SE; n = 7; ***P<0.001; Student's ttest). f, g Expression of Rosa hybrida autophagy related gene 7 (RhATG7) and RhATG11 in RhWRKY40-silenced petals and the TRV controls (mean ± SE; n = 3; **P<0.01; Student’s ttest)
Discussion
Autophagy plays a key role in petal senescence
Autophagy is a degradation and recycling process crucial for the functioning and survival of eukaryotic cells. Although the autophagy-associated molecular mechanisms during leaf senescence have been extensively studied, only a few studies on autophagy have been performed in the flowers. Wilting is a visible symptom of petal senescence (Ma et al. 2005), accompanied by the collapse of epidermal cells (Hoeberichts et al. 2005; Smith et al. 1992; van Doorn et al. 2003). In this study, we characterized rose petal senescence at the cellular level by surveying the ultrastructural changes in adaxial epidermal cells. We observed an abrupt surge in autophagic bodies in rose petals during the early stage (stage 3) of flower opening, as depicted in Fig. 1. Notably, no signs of senescence were observable in the flower at that point. Based on our previous studies, multiple major senescence-related genes were upregulated at the stage 3, so we considered that the petal senescence in roses was initiated at this stage (Chen et al. 2023). Therefore, cellular changes associated with petal senescence occur earlier than the corresponding phenotypic changes. We propose that an increase in autophagy may serve as an indicator of the petal senescence onset. Furthermore, we observed that ATG8-PE bands were significantly elevated in stage 3 petals (Fig. 1d). Similar findings have been reported in the leaves, in which a similar upregulation of ATG genes is concurrent to the onset of leaf senescence (Guo et al. 2021).
Autophagy is involved in programmed cell death (PCD) during petal senescence
Petal senescence is a form of PCD, and tonoplast rupture is the primary cause of this type of developmental cell death (van Doorn and Woltering 2008). Damaged tonoplasts and plasmolysis have been observed in the opened flowers (stage 4 and stage 5) (Fig. 1). In Ipomoea nil, the transcription factor EPH1 promotes PCD to regulate petal senescence (Shibuya et al. 2014). Autophagy has a significant role in PCD and contributes to chloroplast degradation during natural leaf senescence (Otegui 2018). Studies in rice have indicated that OsATG7 promotes pollen maturation by regulating PCD and facilitating tapetal degradation (Kurusu et al. 2014). Autophagy deficiency leads to delayed cell death and hinders corpse clearance in the columella (Feng et al. 2022). In atg5 plants, tracheary elements (TE) formation, a typical process involving PCD, is inhibited, resulting in a reduction in protoxylem and metaxylem cells (Kwon et al. 2010). Silencing ATG4 in tobacco suspension-cultured cells reduces vacuolar processing enzyme (VPE) activity and cell death (Teper-Bamnolker et al. 2021). We propose that autophagy regulates PCD in the rose flower petals, thereby participating in petal senescence, a swift and irreversible process.
RhHY5 and RhPIF4/8 finely tune petal senescence under light and dark conditions
Our experimental findings suggested that RhHY5 and RhPIF4/8 participate in regulating petal senescence under light/dark conditions (Figs. 3, 4, S3). RhHY5 expression levels were higher in LL compared to DD, consistent with a previous study that suggested HY5 was more highly expressed in seedlings grown under white light (15–20-fold higher compared to dark conditions) (Osterlund et al. 2000). Additionally, HY5 expression decreased within 5 h of darkness, due to photoactivated cryptochromes interacting with SUPPRESSOR OF PHYA-105 (SPA) 1, disrupting the CONSTITUTIVE PHOTOMORPHOGENIC 1 (COP1) ubiquitin ligase complex (Saijo et al. 2003). In the dark, PIFs accumulate and, upon light exposure, are degraded by the 26S proteasome pathway (Shen et al. 2007). A competitive mechanistic interaction exists between HY5 and the PIFs. In Arabidopsis thaliana, HY5 competitively binds to the PIF4-targeted gene promoters, repressing PIF4-mediated thermosensory cell elongation (Gangappa and Kumar 2017). Our results revealed distinct roles of RhPIF4 and RhPIF8 in petal senescence (Fig. 3, S3), suggestive of a competitive relationship between RhPIF4 and RhPIF8 with RhWRKY40. RhPIF4 expression remained stable under dark conditions, whereas RhPIF8 expression increased during the late stage (Zhang et al. 2021). We hypothesize that RhPIF8 competes with RhPIF4 for Rh WRKY40 under dark conditions to slow down autophagy-induced senescence.
We demonstrated in our previous study that the RhPIF8-RhBBX28 module significantly affected mitochondrial ROS homeostasis (Zhang et al. 2021). The levels of mRNA of ATG genes were lower in RhPIF8-overexpressing petals (Fig. S3), potentially due to the reduced ROS levels in the petals. Elevated ROS levels and the resulting downstream signaling could induce autophagy, which eliminated ROS and damaged organelles, thereby cells are protected against excessive ROS-induced rapid cell death (Guan et al. 2019; Scherz-Shouval and Elazar 2007; Zhou et al. 2018; Zhu et al. 2018). The ATG4 functions as a central redox signal integrator (Pérez-Pérez et al. 2012). In tomato, local virus inoculation in the leaves induces ROS production in the root-tip cells, activating ROS-triggered autophagy to counter intracellular ROS oxidative damage (Zhou et al. 2018). In tomato, alternative oxidase (AOX), an oxidase of the mitochondrial alternative oxidation pathway, mediates autophagy activation by balancing ROS levels in ethylene-mediated drought tolerance (Zhu et al. 2018). Further investigation is necessary to unveil the regulatory mechanisms and interactions between ROS, autophagy, and petal senescence.
Conclusions
Based on our results, we propose a model in which RhPIF4, RhPIF8, and RhHY5 regulate petal senescence through the modulation of autophagic activities in response to light or dark conditions (Fig. 8). In the presence of light, the RhHY5-RhWRKY40 module inhibits RhATG7/11 expression, hence delaying petal senescence. However, when the RhPIF4-RhWRKY40 module is activated under dark conditions, it results in the enhancement of autophagic activity and accelerates petal senescence, while the RhPIF8-RhWRKY40 module acts antagonistically on the same process. Therefore, a competitive relationship may exist among RhPIF4 and RhPIF8 with RhWRKY40 in the dark, which regulates autophagy and, thereby, petal senescence.
A proposed model of Rosa hybrida phytochrome-interacting factor 4/8 (RhPIF4/8) and Rosa hybrida elongated hypoctyl 5 (RhHY5) regulating autophagy-mediated senescence under dark conditions in rose petals. In rose petals, autophagy is induced under dark conditions and inhibited in the light. In the presence of light, the RhHY5-Rosa hybrida WRKY40 (RhWRKY40) module delays petal senescence by suppressing autophagic activity. Conversely, under dark conditions, RhPIF4-RhWRKY40 accelerates petal senescence by promoting Rosa hybrida autophagy related gene 7/11 (RhATG7/11) expression, while RhPIF8-RhWRKY40 delays petal senescence by repressing autophagic activity
Availability of data and materials
The datasets generated and/or analyzed are available from the corresponding author upon reasonable request.
References
Al-Sady B, Ni W, Kircher S, Schäfer E, Quail PH. Photoactivated phytochrome induces rapid PIF3 phosphorylation prior to proteasome-mediated degradation. Mol Cell. 2006;23:439–46. https://doi.org/10.1016/j.molcel.2006.06.011.
Avila-Ospina L, Moison M, Yoshimoto K, Masclaux-Daubresse C. Autophagy, plant senescence, and nutrient recycling. J Exp Bot. 2014;65:3799–811. https://doi.org/10.1093/jxb/eru039.
Broda M, Millar AH, Van Aken O. Mitophagy: a mechanism for plant growth and survival. Trends Plant Sci. 2018;23:434–50. https://doi.org/10.1016/j.tplants.2018.02.010.
Burko Y, Seluzicki A, Zander M, Pedmale UV, Ecker JR, Chory J. Chimeric activators and repressors define HY5 activity and reveal a light-regulated feedback mechanism. Plant Cell. 2020;32:967–83. https://doi.org/10.1105/tpc.19.00772.
Castillon A, Shen H, Huq E. Phytochrome interacting factors: central players in phytochrome-mediated light signaling networks. Trends Plant Sci. 2007;12:514–21. https://doi.org/10.1016/j.tplants.2007.10.001.
Chen C, Ma Y, Zuo L, Xiao Y, Jiang Y, Gao J. The calcineurin B-like 4/CBL-interacting protein 3 module degrades repressor JAZ5 during rose petal senescence. Plant Physiol. 2023;193:1605–20. https://doi.org/10.1093/plphys/kiad365.
Cheng C, Yu Q, Wang Y, Wang H, Dong Y, Ji Y, et al. Ethylene-regulated asymmetric growth of the petal base promotes flower opening in rose (Rosa hybrida). Plant Cell. 2021;33:1229–51. https://doi.org/10.1093/plcell/koab031.
Chi C, Li X, Fang P, Xia X, Shi K, Zhou Y, et al. Brassinosteroids act as a positive regulator of NBR1-dependent selective autophagy in response to chilling stress in tomato. J Exp Bot. 2020;71:1092–106. https://doi.org/10.1093/jxb/erz466.
Chung T, Phillips AR, Vierstra RD. ATG8 lipidation and ATG8-mediated autophagy in Arabidopsis require ATG12 expressed from the differentially controlled ATG12A and ATG12B loci. Plant J. 2010;62:483–93. https://doi.org/10.1111/j.1365-313X.2010.04166.x.
Eulgem T, Rushton PJ, Robatzek S, Somssich IE. The WRKY superfamily of plant transcription factors. Trends Plant Sci. 2000;5:199–206. https://doi.org/10.1016/s1360-1385(00)01600-9.
Feng Q, De Rycke R, Dagdas Y, Nowack MK. Autophagy promotes programmed cell death and corpse clearance in specific cell types of the Arabidopsis root cap. Curr Biol. 2022;32:2110–19.e3. https://doi.org/10.1016/j.cub.2022.03.053.
Gangappa SN, Kumar SV. DET1 and HY5 control PIF4-mediated thermosensory elongation growth through distinct mechanisms. Cell Rep. 2017;18:344–51. https://doi.org/10.1016/j.celrep.2016.12.046.
Guan B, Lin Z, Liu D, Li C, Zhou Z, Mei F, et al. Effect of waterlogging-induced autophagy on programmed cell death in Arabidopsis roots. Front Plant Sci. 2019;10:468. https://doi.org/10.3389/fpls.2019.00468.
Guo Y, Ren G, Zhang K, Li Z, Miao Y, Guo H. Leaf senescence: progression, regulation, and application. Mol Hortic. 2021;1:5. https://doi.org/10.1186/s43897-021-00006-9.
Hoeberichts FA, de Jong AJ, Woltering EJ. Apoptotic-like cell death marks the early stages of gypsophila (Gypsophila paniculata) petal senescence. Postharvest Biol Technol. 2005;35:229–36. https://doi.org/10.1016/j.postharvbio.2004.10.005.
Jiang C, Liang Y, Deng S, Liu Y, Zhao H, Li S, et al. The RhLOL1–RhILR3 module mediates cytokinin-induced petal abscission in rose. New Phytol. 2022;237:483–96. https://doi.org/10.1111/nph.18556.
Kurusu T, Koyano T, Hanamata S, Kubo T, Noguchi Y, Yagi C, et al. OsATG7 is required for autophagy-dependent lipid metabolism in rice postmeiotic anther development. Autophagy. 2014;10:878–88. https://doi.org/10.4161/auto.28279.
Kwon SI, Cho HJ, Jung JH, Yoshimoto K, Shirasu K, Park OK. The Rab GTPase RabG3b functions in autophagy and contributes to tracheary element differentiation in Arabidopsis Plant J. 2010;64:151–64. https://doi.org/10.1111/j.1365-313X.2010.04315.x.
Li F, Chung T, Pennington JG, Federico ML, Kaeppler HF, Kaeppler SM, et al. Autophagic recycling plays a central role in maize nitrogen remobilization. Plant Cell. 2015;27:1389–408. https://doi.org/10.1105/tpc.15.00158.
Ma N, Cai L, Lu W, Tan H, Gao J. Exogenous ethylene influences flower opening of cut roses (Rosa hybrida) by regulating the genes encoding ethylene biosynthesis enzymes. Sci China Life Sci. 2005;48:434–44. https://doi.org/10.1360/062004-37.
Ma N, Ma C, Liu Y, Shahid MO, Wang C, Gao J. Petal senescence: a hormone view. J Exp Bot. 2018;69:719–32. https://doi.org/10.1093/jxb/ery009.
Nozue K, Covington MF, Duek PD, Lorrain S, Fankhauser C, Harmer SL, et al. Rhythmic growth explained by coincidence between internal and external cues. Nature. 2007;448:358–61. https://doi.org/10.1038/nature05946.
Osterlund MT, Hardtke CS, Wei N, Deng XW. Targeted destabilization of HY5 during light-regulated development of Arabidopsis Nature. 2000;405:462–6. https://doi.org/10.1038/35013076.
Otegui MS. Vacuolar degradation of chloroplast components: autophagy and beyond. J Exp Bot. 2018;69:741–50. https://doi.org/10.1093/jxb/erx234.
Pérez-Pérez ME, Lemaire SD, Crespo JL. Reactive oxygen species and autophagy in plants and algae. Plant Physiol. 2012;160:156–64. https://doi.org/10.1104/pp.112.199992.
Rubinsztein DC, Cuervo AM, Ravikumar B, Sarkar S, Korolchuk V, Kaushik S, et al. In search of an "autophagomometer" Autophagy. 2009;5:585–9. https://doi.org/10.4161/auto.5.5.8823.
Saijo Y, Sullivan JA, Wang H, Yang J, Shen Y, Rubio V, et al. The COP1-SPA1 interaction defines a critical step in phytochrome A-mediated regulation of HY5 activity. Genes Dev. 2003;17:2642–7. https://doi.org/10.1101/gad.1122903.
Sánchez-Sevilla JF, Botella MA, Valpuesta V, Sanchez-Vera V. Autophagy is required for strawberry fruit ripening. Front Plant Sci. 2021;12:688481. https://doi.org/10.3389/fpls.2021.688481.
Scherz-Shouval R, Elazar Z. ROS, mitochondria and the regulation of autophagy. Trends Cell Biol. 2007;17:422–7. https://doi.org/10.1016/j.tcb.2007.07.009.
Shen Y, Khanna R, Carle CM, Quail PH. Phytochrome induces rapid PIF5 phosphorylation and degradation in response to red-light activation. Plant Physiol. 2007;145:1043–51. https://doi.org/10.1104/pp.107.105601.
Shen H, Zhu L, Castillon A, Majee M, Downie B, Huq E. Light-induced phosphorylation and degradation of the negative regulator phytochrome-interacting facotr1 from Arabidopsis depend upon its direct physical interactions with photoactivated phytochromes. Plant Cell. 2008;20:1586–602. https://doi.org/10.1105/tpc.108.060020.
Shibutani ST, Yoshimori T. A current perspective of autophagosome biogenesis. Cell Res. 2014;24:58–68. https://doi.org/10.1038/cr.2013.159.
Shibuya K, Yamada T, Suzuki T, Shimizu K, Ichimura K. InPSR26, a putative membrane protein, regulates programmed cell death during petal senescence in Japanese morning glory. Plant Physiol. 2009;149:816–24. https://doi.org/10.1104/pp.108.127415.
Shibuya K, Niki T, Ichimura K. Pollination induces autophagy in petunia petals via ethylene. J Exp Bot. 2013;64:1111–20. https://doi.org/10.1093/jxb/ers395.
Shibuya K, Shimizu K, Niki T, Ichimura K. Identification of a NAC transcription factor, EPHEMERAL1, that controls petal senescence in Japanese morning glory. Plant J. 2014;79:1044–51. https://doi.org/10.1111/tpj.12605.
Shimizu S, Arakawa S, Nishida Y. Autophagy takes an alternative pathway. Autophagy. 2010;6:290–1. https://doi.org/10.4161/auto.6.2.11127.
Smith MT, Sarks Y, van Staden J. Ultrastructural changes in the petals of senescing flowers of Dianthus caryophyllus L. Ann Bot. 1992;69:277–85. https://doi.org/10.1093/oxfordjournals.aob.a088341.
Song Y, Yang C, Gao S, Zhang W, Li L, Kuai B. Age-triggered and dark-induced leaf senescence require the bHLH transcription factors PIF3, 4, and 5. Mol Plant. 2014;7:1776–87. https://doi.org/10.1093/mp/ssu109.
Su T, Li X, Yang M, Shao Q, Zhao Y, Ma C, et al. Autophagy: an intracellular degradation pathway regulating plant survival and stress response. Front Plant Sci. 2020;11:164. https://doi.org/10.3389/fpls.2020.00164.
Teper-Bamnolker P, Danieli R, Peled-Zehavi H, Belausov E, Abu-Abied M, Avin-Wittenberg T, et al. Vacuolar processing enzyme translocates to the vacuole through the autophagy pathway to induce programmed cell death. Autophagy. 2021;17:3109–23. https://doi.org/10.1080/15548627.2020.1856492.
Üstün S, Hafrén A, Hofius D. Autophagy as a mediator of life and death in plants. Curr Opin Plant Biol. 2017;40:122–30. https://doi.org/10.1016/j.pbi.2017.08.011.
van Doorn WG, Woltering EJ. Many ways to exit? Cell death categories in plants. Trends Plant Sci. 2005;10(3):117–22. https://doi.org/10.1016/j.tplants.2005.01.006.
van Doorn WG, Woltering EJ. Physiology and molecular biology of petal senescence. J Exp Bot. 2008;59:453–80. https://doi.org/10.1093/jxb/erm356.
van Doorn WG, Balk PA, van Houwelingen AM, Hoeberichts FA, Hall RD, Vorst O, et al. Gene expression during anthesis and senescence in Iris flowers. Plant Mol Biol. 2003;53:845–63. https://doi.org/10.1023/B:PLAN.0000023670.61059.1d.
van Doorn WG, Kirasak K, Ketsa S. Macroautophagy and microautophagy in relation to vacuole formation in mesophyll cells of Dendrobium tepals. J Plant Physiol. 2015;177:67–73. https://doi.org/10.1016/j.jplph.2015.01.006.
Wang Y, Cai S, Yin L, Shi K, Xia X, Zhou Y, et al. Tomato HsfA1a plays a critical role in plant drought tolerance by activating ATG genes and inducing autophagy. Autophagy. 2015;11:2033–47. https://doi.org/10.1080/15548627.2015.1098798.
Wang Y, Cao J, Wang K, Xia X, Shi K, Zhou YH, et al. BZR1 mediates brassinosteroid-induced autophagy and nitrogen starvation in tomato. Plant Physiol. 2019;179:671–85. https://doi.org/10.1104/pp.18.01028.
Woo HR, Kim HJ, Lim PO, Nam HG. Leaf senescence: systems and dynamics aspects. Annu Rev Plant Biol. 2019;70:347–76. https://doi.org/10.1146/annurev-arplant-050718-095859.
Xia K, Liu T, Ouyang J, Wang R, Fan T, Zhang M. Genome-wide identification, classification, and expression analysis of autophagy-associated gene homologues in rice (Oryza sativa L). DNA Res. 2011;18:363–77. https://doi.org/10.1093/dnares/dsr024.
Yang C, Luo M, Zhuang X, Li F, Gao C. Transcriptional and epigenetic regulation of autophagy in plants. Trends Genet. 2020;36:676–88. https://doi.org/10.1016/j.tig.2020.06.013.
Yang C, Shen W, Yang L, Sun Y, Li X, Lai M, et al. HY5-HDA9 module transcriptionally regulates plant autophagy in response to light-to-dark conversion and nitrogen starvation. Mol Plant. 2020;13:515–31. https://doi.org/10.1016/j.molp.2020.02.011.
Yao W, Li Y, Wu L, Wu C, Zhang Y, Liu J, et al. Atg11 is required for initiation of glucose starvation-induced autophagy. Autophagy. 2020;16:2206–18. https://doi.org/10.1080/15548627.2020.1719724.
Zhang S, Feng M, Chen W, Zhou X, Lu J, Wang Y, et al. In rose, transcription factor PTM balances growth and drought survival via PIP2;1 aquaporin. Nat Plants. 2019;5:290–9. https://doi.org/10.1038/s41477-019-0376-1.
Zhang B, Shao L, Wang J, Zhang Y, Guo X, Peng Y, et al. Phosphorylation of ATG18a by BAK1 suppresses autophagy and attenuates plant resistance against necrotrophic pathogens. Autophagy. 2020;17:2093–110. https://doi.org/10.1080/15548627.2020.1810426.
Zhang Y, Wu Z, Feng M, Chen J, Qin M, Wang W, et al. The circadian-controlled PIF8-BBX28 module regulates petal senescence in rose flowers by governing mitochondrial ROS homeostasis at night. Plant Cell. 2021;33:2716–35. https://doi.org/10.1093/plcell/koab152.
Zhang Z, Liu Y, Yang T, Wu S, Sun H, Wu J, et al. Haplotype-resolve genome assembly and resequencing provide insights into the origin and domestication of modern rose. bioRxiv. 2023:2023.06.02.543351. https://doi.org/10.1101/2023.06.02.543351.
Zhou X, Zhao P, Wang W, Zou J, Cheng TH, Peng XB, et al. A comprehensive, genome-wide analysis of autophagy-related genes identified in tobacco suggests a central role of autophagy in plant response to various environmental cues. DNA Res. 2015;22:245–57. https://doi.org/10.1093/dnares/dsv012.
Zhou S, Hong Q, Li Y, Li Q, Wang M. Autophagy contributes to regulate the ROS levels and PCD progress in TMV-infected tomatoes. Plant Sci. 2018;269:12–9. https://doi.org/10.1016/j.plantsci.2017.11.002.
Zhu T, Zou L, Li Y, Yao X, Xu F, Deng X, et al. Mitochondrial alternative oxidase-dependent autophagy involved in ethylene-mediated drought tolerance in Solanum lycopersicum Plant Biotechnol J. 2018;16:2063–76. https://doi.org/10.1111/pbi.12939.
Acknowledgements
Not applicable.
Funding
This work was supported by the National Natural Science Foundation of China (No. 32230094 and 32002079), the Construction of Beijing Science and Technology Innovation and Service Capacity in Top Subjects (CEFFPXM2019_014207_000032) and the 2115 Talent Development Program of China Agricultural University.
Author information
Authors and Affiliations
Contributions
JG, CC and XS conceived and coordinated this project; WW performed the experiments with contributions from YZ, BZ and ZW; WW and CC wrote the manuscript under the supervision of and with contributions from JG. All authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests. Prof. Junping Gao is an editorial board member of Horticulture Advances and was not involved in the journal’s review or decisions related to this manuscript.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1
: Fig. S1. Genome-wide expression analysis of Rosa hybrida autophagy related genes (RhATGs). Fig. S2. Phylogenetic tree and expression of Rosa hybrida phytochrome-interacting factors (RhPIFs). Fig. S3. Expression of autophagy related genes (ATGs) in Rosa hybrida phytochrome-interacting factor 8 (RhPIF8) overexpression petals. Fig. S4. Phylogenetic tree and expression patterns of elongated hypoctyl 5 (HY5) and elongated hypoctyl homologs (HYHs). Fig. S5. Rosa hybrida phytochrome-interacting factor 4/8 (RhPIF4/8) and Rosa hybrida elongated hypoctyl 5 (RhHY5) can not bind the promoters of autophagy related genes (ATGs). Fig. S6. Rosa hybrida phytochrome-interacting factor 8 (RhPIF8) interacts with Rosa hybrida WRKY40 (RhWRKY40). Fig. S7. Promoter structure diagrams of the Rosa hybrida autophagy related gene 7/11 (RhATG7/11) genes. Fig. S8. Dual-luciferase (Dual-LUC) analyses of the influence of Rosa hybrida phytochrome-interacting factor 8 (RhPIF8) and Rosa hybrida WRKY40 (RhWRKY40) on Rosa hybrida autophagy related gene 7/11 (RhATG7/11) promoters.
Additional file 2
: Table S1. Primers used in this study.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, W., Chen, C., Zhao, Y. et al. Transcription factors RhPIF4/8 and RhHY5 regulate autophagy-mediated petal senescence in rose (Rosa hybrida). HORTIC. ADV. 1, 17 (2023). https://doi.org/10.1007/s44281-023-00021-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44281-023-00021-4