Abstract
Purpose
To investigate the methylation status and expression level of G protein-coupled receptor 135 (GPR135) in nasopharyngeal carcinoma (NPC) and determine its prognostic value.
Methods
The GPR135 methylation data of NPC and normal nasopharyngeal tissues were obtained from the Gene Expression Omnibus (GEO) GSE52068 dataset. The GPR135 promoter region methylation level in four normal nasopharyngeal epithelial tissues and eight NPC tissues was detected by bisulfite sequencing. GPR135 expression in NPC and normal nasopharyngeal tissue was obtained from the GEO GSE13597 dataset.The GPR135 mRNA expression levels in 13 NPC and 26 healthy control tissues were assessed with quantitative real-time PCR (qRT-PCR). The GPR135 expression level in 124 NPC tissue sections was analyzed by immunohistochemistry. The correlation between GPR135 expression and clinicopathological features was analyzed by a chi-square test. GPR135 expression in patients with NPC was evaluated by immunohistochemistry, and its influence on prognosis was assessed by Kaplan-Meier and Cox regression analyses.
Results
The bisulfite sequencing demonstrated that the GPR135 promoter region was highly methylated in NPC tissues. The immunohistochemistry results revealed that patients with high GPR135 expression had better overall survival (hazard ratio [HR] = 0.177, 95% confidence interval [95%CI]: 0.072–0.437, P = 0.008), disease-free survival (HR = 0.4401, 95%CI: 0.222–0.871, P = 0.034), and local recurrence-free survival (HR = 0.307, 95%CI: 0.119–0.790, P = 0.046) than those with low GPR135 expression.
Conclusion
GPR135 is hypermethylated in NPC, where high GPR135 expression indicates a positive prognosis. Therefore, GPR135 might be a prognostic indicator.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Nasopharyngeal carcinoma (NPC) is a highly endemic malignant head and neck tumors in Southern China and Southeast Asia [1,2,3], and local recurrence and distant metastasis after standardized treatment remain key challenges [4, 5]. Hence, further exploration of the mechanisms and biomarkers of NPC progression could aid the identification of new therapeutic strategies for patients with NPC.
G protein-coupled receptors (GPCRs) are membrane proteins that contain seven transmembrane structures and are the largest protein superfamily in the human body. GPCRs are widely available in cell membranes and participate in tumor development by affecting tumor proliferation, growth, and tumor cell metastasis [6,7,8,9]. GPCRs are aberrantly expressed in various cancers, such as lung, breast, pancreatic, colorectal and endometrial cancers [10,11,12,13,14]. GPR35 exerts pro-cancer effects by stimulating angiogenesis and tumor tissue remodeling in colon cancer [15], while GPR17 expression is associated with higher survival in glioblastoma [16]. In breast cancer, GPR19 was associated with lower cancer cell invasiveness by inducing increased E-cadherin expression [17].
Abnormal GPCR methylation alterations are closely associated with tumor development in breast, lung, and colorectal cancers [18,19,20]. A recent report indicated that genome-wide methylation and chromosomal copy number variation were detected in small intestine and appendix neuroendocrine tumors, where GPCR family expression was significantly elevated in differentially methylated genes [21]. Previously, we reported aberrant GPR135 methylation and expression in NPC tissues using genome-wide methylation microarray assay [22]. However, GPR135 has been poorly reported as an orphan receptor.
In this study, we investigated the relationship between GPR135 methylation status and expression level. Furthermore, we explored the potential roles of expression correlation with the clinical characteristics of a cohort of patients with NPC, which might provide a novel therapeutic target for individualized NPC treatment.
2 Methods
2.1 Preparation of clinical specimens
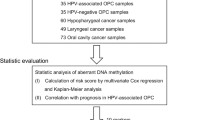
The GPR135 methylation data of 24 pairs of NPC and normal nasopharyngeal tissues were obtained from the Gene Expression Omnibus (GEO) GSE52068 dataset (https://www.ncbi.nlm.nih.gov/geo/). We also accessed the GPR135 mRNA expression values of three normal control tissues and 25 primary NPC tissues from the GEO GSE13597 dataset [23]. Furthermore, we obtained 158 NPC and 15 normal nasopharyngeal tissues from Guilin Medical University Affiliated Hospital, which we used for GPR135 promoter region bisulfite sequencing and detection of GPR135 expression for prognostic analysis. All patients were newly diagnosed with NPC and had not undergone any anti-tumor therapy. All the patients were staged according to 7th Union for International Cancer Control and American Joint Committee on Cancer staging system. The patients were followed-up every 3 months through outpatient examination and telephone interviews until 5 years after radiotherapy or death. Overall survival (OS) was calculated from the time of diagnosis to death from any cause. Disease-free survival (DFS) was calculated from the time of diagnosis to treatment failure or death from any cause; locoregional relapse-free survival (LRRFS) to the date of first locoregional relapse or death from any cause; and distant metastasis-free survival (DMFS) to the date of first detection of metastasis or death from any cause. The Guilin Medical University Affiliated Hospital Medical Ethics Committee approved this study (ethics number: QTLL202140). All patients provided written informed consent prior to enrollment.
2.2 DNA extraction and bisulfite pyrosequencing
DNA (1–2 µg) was extracted from four NPC tissues and eight normal tissues using an AllPrep RNA/DNA Mini Kit (Qiagen, Hilden, Germany) and treated with sodium bisulfite using an EpiTect Bisulfite Kit (Qiagen). The kits were all used according to the manufacturer’s instructions. PCR was performed under the following conditions: initial denaturation at 95 °C for 3 min, followed by 50 cycles of denaturation at 95 °C for 15 s, annealing at 54 °C for 20 s, and extension at 72 °C for 30 s. The final extension step was conducted at 72 °C for 5 min. The GPR135 bisulfite pyrosequencing primers (Supplementary Table 1) were designed using PyroMark Analysis Software 2.0 (Qiagen). The sequencing reactions and methylation level quantification were performed using a PyroMark Q96 System (Qiagen).
2.3 Quantitative real-time reverse transcription PCR (qRT-PCR)
Total RNA was extracted from tissues or cells using TRIzol (Invitrogen), and complementary DNA (cDNA) was synthesized using a reverse transcription kit (Promega, Madison, WI, USA). The kits were all used according to the manufacturer’s instructions. Using the cDNA as a template, qRT-PCR was performed using SYBR Green qPCR SuperMix-UDG (Invitrogen) on a CFX96 System (Bio-Rad, Hercules, CA, USA). Supplementary Table 1 lists the primers used.
2.4 Immunohistochemistry
GPR135 expression was detected with immunohistochemistry. First, tissue sections were baked in a 60 °C oven for 2 h, then dewaxed in xylene and rehydrated with gradient ethanol. Endogenous peroxidase was blocked using 3% hydrogen peroxide. Antigen retrieval was performed by heating the sections in 100 °C water for 10 min, followed by washing with phosphate-buffered saline containing Tween-20 (PBST) and blocking with goat serum (Beyotime). Then, the sections were incubated overnight with the primary antibody (anti-GPR135, Invitrogen) and subsequently incubated with a biotin-labeled secondary antibody. The sections were stained using 3,3’-diaminobenzidine (DAB, Agilent Technologies, Santa Clara, CA, USA) and counterstained with hematoxylin. Immunohistochemical scoring was evaluated according to a previously described method [24]. Staining intensity was graded from 0 to 3 (no staining, weak staining, moderate staining, strong staining), and the positivity rate was graded from 0 to 4 (< 25%, 26–50%, 51–75%, > 75%). The total score was calculated by multiplying the staining intensity and positivity rate scores. Patients were categorized as having low expression (< 4) or high expression (≥ 4) of GPR135 based on their total scores, which were determined by two pathologists who were unaware of the patient’s condition.
2.5 Statistical analysis
Statistical analysis was conducted using Statistical Package for the Social Sciences (SPSS) 23 and data were visualized using GraphPad Prism 8. Differences between two groups were assessed with t-tests, and the composition ratios of different groups were compared using chi-square tests. Survival was analyzed with the Kaplan-Meier test, while the influence of single and multiple factors on patient prognosis was analyzed with the Cox proportional hazards model. P < 0.05 was considered statistically significant.
3 Results
3.1 GPR135 was frequently methylated in NPC tissues
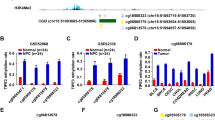
Figure 1 A depicts the frequency of GPR135 methylation in the 24 pairs of NPC tissues and normal nasopharyngeal tissues from the GSE52068 dataset. Eight CpG sites in GPR135 (cg15436096, cg09859398, cg22821834, cg16411251, cg04186360, cg16322262, cg21893185, cg10014563) were significantly hypermethylated in the NPC tissues but not in the normal nasopharyngeal tissues (Fig. 1B). We selected a -87 to -154 bp region upstream of the GPR135 promoter transcription start site for bisulfite sequencing, and determined that the GPR135 promoter region methylation level was generally higher in the NPC tissues than in the normal tissues (Fig. 1C and D).
The GPR135 promoter region was hypermethylated and downregulated in NPC. A Heatmap cluster of GPR135 CG sites between NPC (n = 24) and normal nasopharyngeal tissue samples (n = 24). B The methylation level of GPR135 CG sites in the GSE52068 dataset. C, D Bisulfite sequencing analysis of the GPR135 promoter region in normal tissues (n = 4) and NPC tissues (n = 8). *P < 0.05, ***P < 0.001
3.2 GPR135 expression was decreased in NPC tissues
The GSE13597 dataset analysis demonstrated that the GPR135 mRNA levels were downregulated in NPC (Fig. 2A). qRT-PCR analysis of 13 randomly selected normal tissues and 26 NPC tissues demonstrated that GPR135 mRNA expression levels was significantly lower in NPC tissues compared to normal tissues (Fig. 2B).
3.3 Higher GPR135 expression was associated with better prognosis in patients with NPC
To validate the clinical significance of GPR135, we obtained NPC specimens from 124 diagnosed patients at the Affiliated Hospital of Guilin Medical University between 2013 and 2017. Table 1 lists the patients’ clinicopathological characteristics and their GPR135 expression levels. The patients were divided into GPR135 low-expression and high-expression groups based on their immunohistochemical scores (Fig. 3A and B). Survival analysis of the two groups demonstrated that patients in the GPR135 high-expression group had better OS (Fig. 3C, P = 0.008), DFS (Fig. 3D, P = 0.034), and LRRFS (Fig. 3E, P = 0.046) compared to those in the GPR135 low-expression group, but no significant difference was observed for DMFS (Fig. 3F, P = 0.174).
Immunohistochemical risk stratification and prognostic analysis of GPR135. A, B Immunohistochemical staining of representative samples from the GPR135 low-expression group (A) and GPR135 high-expression group (B). C–F Comparison of OS (C), DFS (D), LRRFS (E), and DMFS (F) between the GPR135 low-expression and high-expression groups
3.4 GPR135 was an independent prognostic factor
In the multivariate analysis, the variables of age, sex (male vs. female), overall stage (II–III vs. IVA–B), induction chemotherapy (no vs. yes), concurrent chemotherapy (no vs. yes), and GPR135 expression (high vs. low) were considered in the Cox proportional hazards model for OS, DFS, LRRFS, and DMFS. The results demonstrated that GPR135 was an independent prognostic factor for OS (hazard ratio [HR] = 0.634, 95% confidence interval [95%CI]: 0.408–0.985, P = 0.043), DFS (HR = 0.567, 95%CI: 0.345–0.932, P = 0.025), DMFS (HR = 0.607, 95%CI: 0.387–0.952, P = 0.030), and LRRFS (HR = 0.577, 95%CI: 0.370–0.901, P = 0.016) (Table 2).
4 Discussion
This study describes the methylation status of the potential cancer suppressor gene GPR135 in NPC, where the GPR135 expression level could serve as a prognostic indicator for patients with NPC. We determined that GPR135 was significantly hypermethylated in NPC, which led to a decrease in its expression level. Furthermore, we determined that patients with lower GPR135 expression had poorer prognosis, which could serve as an independent prognostic factor for such patients.
NPC has a high metastasis risk, and even patients with the same clinical stage have differing prognoses [25]. Accordingly, it is necessary to search for more reliable prognostic markers. Recent studies have shown that specific biomarkers such as lncRNA and intratumoral microbiota can be used for risk stratification in NPC patients, which can help make personalized treatment decisions [26, 27]. Research evidence demonstrated that changes in epigenetics, specifically DNA methylation, are key in NPC metastasis, chemotherapy resistance and radioresistance [28,29,30]. Previously, we screened and verified a methylation gene panel that could be used to determine the prognosis of patients with NPC. However, there remain many potential methylation molecular targets for NPC that should be reported. In the present study, we report for the first time the methylation changes of GPR135 in NPC. These results can provide new ideas and insights for future research on the roles and mechanisms of GPCR family genes in NPC.
We demonstrated that patients with NPC with low GPR135 expression had significantly poorer OS, DFS, and LRRFS rates than those with high GPR135 expression. In multivariate analysis, we determined that GPR135 expression was an independent prognostic factor for OS, DFS, LRRFS, and DMFS in NPC. These results suggested that the GPR135 expression level can predict the prognosis of patients with NPC and might provide a new therapy target for individualized treatment of such patients. Similar to our results, another GPCR family gene, GPR68, exhibits abnormal methylation and expression in head and neck tumors, but its expression and prognosis were not significantly correlated [31]. In breast cancer, GPR30 hypermethylation is key in regulating its low expression, and patients with low GPR30 expression have shorter OS and RFS [18]. However, GPR87 overexpression might be involved in cell proliferation, angiogenesis, and chemotherapeutic resistance in pancreatic cancer, and GPR87-overexpressing patients with pancreatic cancer might have reduced OS [14]. These studies indicated the abnormal methylation changes and expression of GPCR family genes in tumors, which might serve as prognostic indicators. As the orphan receptor of the GPCR family, GPR135 function has rarely been reported in tumors. We determined that GPR135 expression and methylation regulation in NPC were abnormal, which can promote mechanistic research of GPR135 in NPC.
In the present study, we did not observe a difference in GPR135 expression in the blood of patients with NPC before and after treatment. Furthermore, we did not perform cellular and animal experiments focusing on the molecular mechanism of GPR135 inhibition of NPC occurrence, development, and malignant biological behavior. Future research on the mechanism of the GPR135 gene in NPC is needed.
In conclusion, GPR135 is hypermethylated in NPC, which might downregulate its expression levels. Patients with NPC with high GPR135 expression had a positive prognosis, and GPR135 was a standalone prognostic factor in NPC. Therefore, GPR135 has the potential to be a novel tumor marker and therapeutic target in NPC.
Availability of data and materials
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Abbreviations
- NPC:
-
Nasopharyngeal carcinoma
- GPR135:
-
G protein-coupled receptor 135
- GPCRs:
-
G protein-coupled receptors
- GEO:
-
Gene Expression Omnibus
- qRT-PCR:
-
quantitative real-time PCR
- cDNA:
-
complementary DNA
- OS:
-
Overall survival
- DFS:
-
Disease free survival
- LRRFS:
-
Locoregional relapse-free survival
- DMFS:
-
Distant metastasis-free survival
- SPSS:
-
Statistical package for the social sciences
- HR:
-
Hazard ratio
- CI:
-
Confidence interval
References
Chen YP, Chan ATC, Le QT, et al. Nasopharyngeal carcinoma. Lancet. 2019;394:64–80.
Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Chen Q, Tang L, Liu N, et al. Famitinib in combination with concurrent chemoradiotherapy in patients with locoregionally advanced nasopharyngeal carcinoma: a phase 1, open-label, dose-escalation study. Cancer Commun (London). 2018;38:66.
Sang Y, Cheng C, Zeng YX, et al. Snail promotes metastasis of nasopharyngeal carcinoma partly by down-regulating TEL2. Cancer Commun (London). 2018;38:58.
Zhao Y, Lei Y, He SW, et al. Hypermethylation of UCHL1 promotes metastasis of nasopharyngeal carcinoma by suppressing degradation of cortactin (CTTN). Cells. 2020;9:559.
O’hayre M, Vázquez-prado J, Kufareva I, et al. The emerging mutational landscape of G proteins and G-protein-coupled receptors in cancer. Nat Rev Cancer. 2013;13:412–24.
Lappano R, Maggiolini M. G protein-coupled receptors: novel targets for drug discovery in cancer. Nat Rev Drug Discov. 2011;10:47–60.
Nieto Gutierrez A, Mcdonald PH. GPCRs: emerging anti-cancer drug targets. Cell Signal. 2018;41:65–74.
Liu Y, An S, Ward R, et al. G protein-coupled receptors as promising cancer targets. Cancer Lett. 2016;376:226–39.
Ahn JI, Yoo JY, Kim TH, et al. G-protein coupled receptor 64 (GPR64) acts as a tumor suppressor in endometrial cancer. BMC Cancer. 2019;19:810.
Chan KK, Lo RC. Deregulation of Frizzled Receptors in Hepatocellular Carcinoma. Int J Mol Sci. 2018;19(1):313.
Hasenoehrl C, Feuersinger D, Sturm EM, et al. G protein-coupled receptor GPR55 promotes colorectal cancer and has opposing effects to cannabinoid receptor 1. Int J Cancer. 2018;142:121–32.
Shen Y, Li C, Zhou L, et al. G protein-coupled oestrogen receptor promotes cell growth of non-small cell lung cancer cells via YAP1/QKI/circNOTCH1/m6A methylated NOTCH1 signalling. J Cell Mol Med. 2021;25:284–96.
Wang L, Zhou W, Zhong Y, et al. Overexpression of G protein-coupled receptor GPR87 promotes pancreatic cancer aggressiveness and activates NF-κB signaling pathway. Mol Cancer. 2017;16:61.
Pagano E, Elias JE, Schneditz G, et al. Activation of the GPR35 pathway drives angiogenesis in the tumour microenvironment. Gut. 2022;71:509–20.
Doan P, Nguyen P, Murugesan A, et al. Targeting Orphan G protein-coupled receptor 17 with T0 ligand impairs Glioblastoma Growth. Cancers. 2021;13(15):3773.
Rao A, Herr DR. G protein-coupled receptor GPR19 regulates E-cadherin expression and invasion of breast cancer cells. Biochim Biophys Acta Mol Cell Res. 2017;1864:1318–27.
Manjegowda MC, Gupta PS, Limaye AM. Hyper-methylation of the upstream CpG island shore is a likely mechanism of GPER1 silencing in breast cancer cells. Gene. 2017;614:65–73.
Pal U, Ghosh S, Limaye AM. DNA methylation in the upstream CpG island of the GPER locus and its relationship with GPER expression in colon cancer cell lines. Mol Biol Rep. 2020;47:7547–55.
Wu K, Xu L, Cheng L. PAR2 promoter hypomethylation regulates PAR2 gene expression and promotes lung Adenocarcinoma Cell Progression. Comput Math Methods Med. 2021;2021:5542485.
Byun S, Affolter KE, Snow AK, et al. Differential methylation of G-protein coupled receptor signaling genes in gastrointestinal neuroendocrine tumors. Sci Rep. 2021;11:12303.
Jiang W, Liu N, Chen XZ, et al. Genome-wide identification of a methylation gene panel as a prognostic biomarker in nasopharyngeal carcinoma. Mol Cancer Ther. 2015;14:2864–73.
Bose S, Yap LF, Fung M, et al. The ATM tumour suppressor gene is down-regulated in EBV-associated nasopharyngeal carcinoma. J Pathol. 2009;217:345–52.
Guo Z, Zhang X, Zhu H, et al. TELO2 induced progression of colorectal cancer by binding with RICTOR through mTORC2. Oncol Rep. 2021;45:523–34.
Chen YP, Wang ZX, Chen L, et al. A bayesian network meta-analysis comparing concurrent chemoradiotherapy followed by adjuvant chemotherapy, concurrent chemoradiotherapy alone and radiotherapy alone in patients with locoregionally advanced nasopharyngeal carcinoma. Ann Oncol. 2015;26:205–11.
Liang YL, Zhang Y, Tan XR, et al. A lncRNA signature associated with tumor immune heterogeneity predicts distant metastasis in locoregionally advanced nasopharyngeal carcinoma. Nat Commun. 2022;13:2996.
Qiao H, Tan XR, Li H, et al. Association of Intratumoral Microbiota with prognosis in patients with nasopharyngeal carcinoma from 2 hospitals in China. JAMA Oncol. 2022;8:1301–309.
Ren X, Yang X, Cheng B, et al. HOPX hypermethylation promotes metastasis via activating SNAIL transcription in nasopharyngeal carcinoma. Nat Commun. 2017;8:14053.
Li Y, Yang X, Du X, et al. RAB37 hypermethylation regulates metastasis and resistance to Docetaxel-Based induction chemotherapy in nasopharyngeal carcinoma. Clin Cancer Res. 2018;24:6495–508.
Chen Y, Zhao Y, Yang X, et al. USP44 regulates irradiation-induced DNA double-strand break repair and suppresses tumorigenesis in nasopharyngeal carcinoma. Nat Commun. 2022;13:501.
Zhang W, Han Y, Li W, et al. Clinical data analysis reveals the role of OGR1 (GPR68) in head and neck squamous cancer. Anim Model Exp Med. 2020;3:55–61.
Acknowledgements
We thank all patients who participated in this study for their cooperation. We also thank the native English speaking scientists of Elixigen Company (Huntington Beach, California) for editing our manuscript.
Funding
This work was supported by the Natural Science Foundation Key Projects of Guangxi [grant number 2018GXNSFDA050021]; the “139” Talent Cultivation Plan Program of Guangxi Medical High-level Backbone, the National Natural Science Foundation of China [grant number 82160479]; the CSCO Youth Innovative Oncology Research Fund [grant number Y-Young2020-0520]; and the Scientific Research Program of the Health Commission of Guangxi Zhuang Autonomous Region [grant numbers Z20201200, Z20170816, Z20170207].
Author information
Authors and Affiliations
Contributions
WJ designed this study and revised the manuscript. CG analyzed the data and wrote original manuscript. GQ, SL, XL and JX examined the data and followed up the patients. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The study protocol was approved by the Medical Ethics Committee of Guilin Medical University Affiliated Hospital (No. QTLL202140) and was performed in accordance with the ethical standards laid down in the 1964 Declaration of Helsinki and its later amendments. All patients provided written informed consent prior to enrollment.
Consent for publication
Not applicable.
Competing interests
The authors declare no potential conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1:
Supplementary Table 1. Primers used in this study.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gan, C., Qin, G., Liao, S. et al. Hypermethylated GPR135 gene expression is a favorable independent prognostic factor in nasopharyngeal carcinoma. Holist Integ Oncol 2, 25 (2023). https://doi.org/10.1007/s44178-023-00048-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44178-023-00048-7