Abstract
Continuous effort is dedicated to clinically and computationally discovering potential drugs for the novel coronavirus-2. Computer-Aided Drug Design CADD is the backbone of drug discovery, and shifting to computational approaches has become necessary. Quantitative Structure–Activity Relationship QSAR is a widely used approach in predicting the activity of potential molecules and is an early step in drug discovery. 3-chymotrypsin-like-proteinase 3CLpro is a highly conserved enzyme in the coronaviruses characterized by its role in the viral replication cycle. Despite the existence of various vaccines, the development of a new drug for SARS-CoV-2 is a necessity to provide cures to patients. In the pursuit of exploring new potential 3CLpro SARS-CoV-2 inhibitors and contributing to the existing literature, this work opted to build and compare three models of QSAR to correlate between the molecules’ structure and their activity: IC50 through the application of Multiple Linear Regression(MLR), Support Vector Regression(SVR), and Particle Swarm Optimization-SVR algorithms (PSO-SVR). The database contains 71 novel derivatives of ML300which have proven nanomolar activity against the 3CLpro enzyme, and the GA algorithm obtained the representative descriptors. The built models were plotted and compared following various internal and external validation criteria, and applicability domains for each model were determined. The results demonstrated that the PSO-SVR model performed best in predictive ability and robustness, followed by SVR and MLR. These results also suggest that the branching degree 6 had a strong negative impact, while the moment of inertia X/Z ratio, the fraction of rotatable bonds, autocorrelation ATSm2, Keirshape2, and weighted path of length 2 positively impacted the activity. These outcomes prove that the PSO-SVR model is robust and concrete and paves the way for its prediction abilities for future screening of more significant inhibitors' datasets.
Similar content being viewed by others
Introduction
Corona Virus Disease 2019 (COVID-19) was declared by the World Health Organization WHO as a global pandemic on 12 March 2020 after its first outbreak in Wuhan, China, in December 2019 (Ciotti et al. 2020). The cause of this unprecedented pneumonia was a coronavirus, later named Severe Acute Respiratory Syndrome coronavirus-2 SARS-CoV-2 (Harapan et al. 2020). As of 4:59 pm CEST on 1 April 2022, the world counts 486 761 597 confirmed cases of COVID-19 and 6 142 735 reported deaths (https://covid19.who.int/). SARS-CoV-2 is a beta-enveloped positive-sense single-stranded RNA coronavirus with a genomic sequence identity of 96.2% with BatCoV RaTG13, a bat coronavirus detected in Rhinolophusaffinis from the province of Yunnan province, China (Harapan et al. 2020).
Vaccines have recently been developed and licensed for urgent use to fight against COVID-19. However, they still present potential problems for patients with allergies, pregnancy, and immune disorders (Zhai et al. 2021), in addition to difficulties of access and distribution, especially in poor developing countries (Palestine and some African countries) (Alaran et al. 2021; Feinmann 2021). Moreover, reaching collective immunity to ensure complete control over the pandemic is time-consuming and yet to be achieved (Zhai et al. 2021). The absence of an appropriate antiviral treatment worsens the situation (Kirmani et al. 2021). Henceforth, a specific treatment for infected patients is a must to preserve lives.
The 3-chymotrypsin-like-proteinase 3CLpro is a highly conserved enzyme across coronavirus variants (Zhai et al. 2021). Despite the emergence of new mutated variants (B.1.1.7, B.1.351, P.1, and B.1.617) (Tahir ul Qamar et al. 2020), the structure of the viral protease 3CLpro remained highly conserved and showed no mutation. Moreover, 3CLpro has available biological evaluation experiments and relies on structural and genomic studies, making necessary data available (Tahir ul Qamar et al. 2020). It interferes in the replication process (Yan and Gao 2021), where it is responsible for the cleavage of 11 non-structural proteins (nsp1 to nsp11) (De et al. 2020) essential for viral replication and transcriptase (Dongliang et al. 2022). In light of this evidence, 3CLpro is considered a valuable target in fighting against SARS-CoV-2.
Given the tedious process of drug discovery and drug repurposing, there is an urgent need to move towards computational approaches. They are a quick and cost-effective choice in disease control (Hamadache et al. 2017). Quantitative Structure–Activity Relationship QSAR is a promising in silico method used to predict the biological activities of different compounds based on the structure (Hamadache et al. 2016), including molecules that have not yet been synthesized. It is established on a solid mathematical algorithm that can produce a predictive correlation model (Laidi et al. 2020). The theory is that the structure contains the element responsible for its activity (Janairo et al. 2021; Yu 2021).
Yu (Yu 2021), in his study, established a QSAR model to predict the inhibitory constant pKi of 89 compounds against 3CLprousing support vector machine and genetic algorithm methods. Six descriptors were selected, and the optimal hyperparameters were C = 90.234 and γ = 1.19810–5. Coefficients of determination of 0.839 and 0.747 were obtained for the training, and test sets, respectively, with a root, mean square error RMSE of 0.435 and 0.525 in the same order.
In a related study, Janairo et al. (Janairo et al. 2021) built and compared different models to predict the inhibitory activity of the 3CLpro enzyme. SVR, Classification and Regression Trees CART, random forest, and MLR were tested. The best-performing model was found to be MLR with values of R2 = 0.806 and RMSE = 0.430 for the test set.
Ishola et al. (Ishola et al. 2021) have developed a QSAR model using MLR to predict the inhibitory activity of known compounds from ChEMBL targeting the viral 3CLpro. The model constructed had a correlation coefficient R2 of 0.907 and a cross-validation coefficient Q2 of 0.866.
One major challenge facing machine learning techniques is the considerable number of parameters to tune before running the algorithm. The performance of the regression algorithms is highly affected by the values of their parameters. These are either set by default or manually configured based on trial and error (Probst et al. 2019). Despite the advantages of the support vector regression technique compared to other supervised learning techniques: suitable for nonlinear phenomena, flexibility thanks to the kernel functions, and easy implementation (Peng and Nagata 2020), it is not an exception from the challenges mentioned above (Calvet et al. 2017). Alternatively, metaheuristic optimization algorithms can potentially solve the parameter tuning and the overfitting of machine learning techniques thanks to their guided search of near-optimal approximates in very little computational time (Shen et al. 2004). Particularly, particle swarm optimization PSO has proved its efficiency in improving the robustness and the performance of machine learning algorithms compared to other optimization algorithms, such as the genetic algorithm GA (Wihartiko et al. 2018). It is advantageous over other techniques as it is easier to implement, has fewer parameters, and yields better accuracy with fewer iterations (Alkabbani et al. 2021). However, very little attention was drawn to it, and very few works have considered its application in predicting inhibitors against the 3CLpro enzyme. Furthermore, the combination of Support Vector Regression-PSO algorithms can outperform other methods thanks to their individual and combined advantages.
To this end, this work aims to build a model able to accurately predict the activity (IC50) of a set of 71 molecules to inhibit the viral SARS-CoV-2 3CLpro using a PSO-SVR hybrid algorithm and to compare it to existing regression algorithms: MLR and SVR. The characteristics have been calculated using the online Blue-Desc Descriptor Calculator software. Representative descriptors were then selected using the Genetic Algorithm to serve as inputs, along with the converted pIC50, to the regression algorithms. A comparison has been performed based on validation criteria.
Materials and methods
Data set retrieval
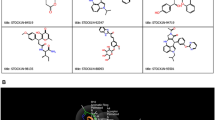
The logic behind the selected nonpeptidic and noncovalent molecules is their proven activity against 3CLpro SARS-CoV-2 and their novelty (Han et al. 2021). The selected compounds are derivatives of ML300 (Fig. 1 a), a potent probe noncovalent inhibitor of the SARS-CoV-1 3CLpro developed by Molecular Libraries Probe Production Centers Network (MLPCN), that despite its nanomolar activity, it was far from being optimized and suffered from poor metabolic stability. A novel benzotriazole-based derivatives series was synthesized. They have a much more robust nanomolar and submicromolar inhibitory potential in live SARS-CoV-2, with a better metabolic profile (Han et al. 2021). These derivatives were intriguing for several reasons. Their experimental activity values were determined, and thus the interpretation of a QSAR model is applicable. Otherwise, data may risk being heterogenous (same molecule but with two or more activity values from different resources). Also, their mechanism of action is well-determined and different from the recently proven inhibitor GC376 (Fig. 1 b). These molecules follow a noncovalent binding mode, while the GC376 follows a covalent binding mode with the residues in the enzyme’s active site. The original articles (Han et al. 2021) and (Fu et al. 2020) explain the detailed mechanisms for the ML300 derivatives series and GC376, respectively. It is worth mentioning that, generally, noncovalent inhibitors are advantageous over covalent inhibitors in terms of low toxicity and broader inhibition spectrum (Aljoundi et al. 2020). This means that noncovalent inhibitors are not only safer and more likely to reach clinical trials and approval, but they can also affect similar targets, which means they are more likely to be repurposed or repositioned. For example, ML300 analogues can target HKU4, a suspected reservoir host for MERS (John et al. 2015). Finally, the quantitative structure–activity for these derivatives remains unknown (Han et al. 2021), making modelling this relationship necessary. Thus, the QSAR models built on the selected database open the door to discovering novel potent noncovalent inhibitors, different from the proven covalent ones, and pave the way for further docking, virtual screening, and synthesis research to identify potential hits based on its findings.
The visualization of the molecules' structures and the generation of their 3D structures with minimal energy were carried out using the Data-Warrior software (Sander et al. 2015). All the used molecules are presented in the supplementary material (Table S-I), along with their IC50 values and.
Other information. For linearization purposes, the experimental values of the half maximal inhibitory concentration (IC50, nM) were converted to 9 minus the negative decimal logarithmic scale (pIC50 = 9-log (IC50)).
Descriptors generation and selection
Molecular descriptors are the numerical representation of molecules’ structural features; they were calculated by Blue-Desc Descriptor Calculator (http://www.scbdd.com/blue_desc/index/) through the online platform ChemDes (Dong et al. 2015)to obtain134 3D, 2D, and 1D descriptors of various families (topological, electronegativity, autocorrelation, functional groups). The genetic algorithm and Double Cross-Validation DCV tool (Roy and Ambure 2016) were used to select only the representative descriptors. This step removes highly correlated descriptors, null, infinity, constant, and near constant descriptors (minimal deviation of the descriptors’ values for the molecules). It leaves only the descriptors that induce a change in the activity value when they change. Hence, the number of descriptors was reduced to 7 impactful descriptors and thus forming a matrix of 71 × 7.
Data division and model development
The data set was optimally distributed into a train set, containing 80% of the dataset, and a test set, including 20% of the dataset. QSAR models were developed using MLR, SVR, and PSO-SVR. The MLR was performed using MLRplusValidation 1.3 software (http://teqip.jdvu.ac.in/QSAR_Tools/) whereas the SVR and PSO-SVRalgorithmswere conducted using MATLAB environment.
Regression methods
Multiple linear regression
Multiple Linear Regression MLR is a type of linear regression that takes more than one explanatory (independent) variable (x1, …, xn) to build a multivariate predictive model(Uyanık and Güler 2013). The equation governing MLR is as follows:
where \({y}_{i}\) is the dependent variable (output), \({\beta }_{0}\) is the intercept corresponding to the dependent variable's valuable when all the independent variables are equal to zero, \(\beta\) is the coefficient of each independent variable, i.e. each descriptor contribution, and \(e\) is the error (residual) (Roy et al. 2015a). The independent variables should not be correlated but uniquely describe the outcome (dependent variable), while their number should be kept below one-fifth of the number of observations. A fast statistical analysis on the descriptors will reveal the over-fitting problem. Accordingly, the number of independent variables can be decreased to avoid over-fitting in the MLR model (Awad and Khanna 2015).
Support vector regression
Support Vector Regression SVR is a regression technique rooted in Vapnik’s concept of support vector. SVR is a kernel-based technique with a solid theoretical basis and is generally easier to implement than artificial neural networks, its major competitor (Rupp 2015).SVR is a supervised-learning machine characterized by using Kernel functions, the number of support vectors, penalty parameters C, coefficient of non-sensitivity ε, and the Gaussian nucleus bandwidth σ (Liu et al. 2018). These parameters can be manually defined or by the mean of heuristic optimization algorithms.
Particle swarm optimization PSO
Inspired by the behaviour of swarms such as birds’ flocks and ant colonies, Particle Swarm optimization is a widespread metaheuristic that Kennedy and Eberhart first developed. It solves complex nonlinear optimization problems with the advantages of having fewer parameters to adjust and an already available literature discussion and community for the parameters that should be adjusted (Almeida 2019). It consists of finding the solution (global best gbest) by comparing and updating an individual’s information of position and velocity (personal best pbest) (Mitikiri et al. 2018). PSO is coupled with machine learning algorithms to determine their optimal parameters.
Statistical validation
Statistical validation establishes the reliability and relevance of the built model for the specific target (Roy et al. 2015a). One of the challenges of validating the QSAR model is to prove it statically accurate in predicting the activity of new molecules unknown to the model (Roy et al. 2016). Internal and external validation criteria were calculated using the external validation tool downloaded from https://sites.google.com/site/dtclabxvplus/ and the Leave One Out validation from https://dtclab.webs.com/software-tools. An extensive explanation of the tool and the metrics used has been detailed in the literature(Roy and Ambure 2016; Hammoudan et al. 2021). Another recent and exciting criterion to compare different models’ fit is Akaike’ sinformation criterion AIC. The lowest value of the AIC corresponds to the model that is best fitted to the experimental data (Falyouna et al. 2020). The used statistical validation criteria and their formulas are detailed in Table 1.
Applicability domain
The Applicability Domain AD is a theoretical space defined by the model’s descriptors and the predicted response of the molecules. The model is considered reliable in the AD space. Thus, the modelled responses outside the AD are considered outliers (Roy et al. 2015b). The AD estimation approaches can be classified as descriptor-space-based and training-set-response-space-based approaches.
The first includes ranges in the descriptor space; the geometric training-set-response-space-based approach includes the range of the variable response technique. Thus, a molecule is considered outside the AD of a particular model if it satisfies the following conditions: (i) at least one descriptor is outside of the ranges approach, and (ii) the distance between the chemical and the centre of the training data set is greater than the acceptable threshold for distance approaches(Roy et al. 2015a). In the present work, the AD was calculated using the Applicability Domain MATLAB toolbox downloaded from https://michem.unimib.it/.
Results and discussion
Selection of representative descriptors
The selection was performed using the GA in Build-QSAR software, revealing the seven most representative descriptors: numberOfHal, SPC-6, WTPT2, MOMIXZ, KierShape2, Numberrotatablebonds, ATSm2.
Constitutional descriptors
Number of hal
A 1-dimensional descriptor representing the number of halogen atoms in the molecule. Halogens are highly electronegative species and are thus highly reactive as they interact with a donor of electron density (Titov et al. 2015). The organic halogens form halogen bindings with the amino acid residues in the active site, specifically chlorine that highly interacts with leucine (Kortagere et al. 2008).
Number rotatable bonds
It is a one-dimensional descriptor that counts the number of bonds that allow free rotations around themselves (Khanna and Ranganathan 2009). The existence of rotatable bonds increases the bioavailability of the molecule and thus indicates the ability for good drug efficacy. The bioactivity depends on the number of rotatable bonds: ten rotatable bonds or fewer bonds have a significant positive impact on the bioavailability and, henceforth, on the molecule’s activity (Veber et al. 2002).
Topological descriptors
Topological descriptors are determined from the graphical representation of molecules without hydrogen atoms. They are thus two-dimensional descriptors that value the weighted arrangement of atoms and their bonds (Roy et al. 2015a).
SPC-6
In general, connectivity indexes measure the possibilities of a molecule interacting with another molecule in a milieu. SPC-6 is a 2-dimensional descriptor that stands for Chi Path/Cluster index. It evaluates the connectivity index at the sixth order or the sixth degree of branching (Dong et al. 2015). SPC-6 considers an atom’s number of electrons and valence electrons. It represents the hypervolume of order 6 in a molecule resulting from contributions of cluster/path fragments (Estrada 2002) and, therefore, the accessible reactivity space of a molecule with the target, in this case, the active site of 3CLpro.
WTPT2
Weighted path-2 is a two-dimensional descriptor (Guha and Jurs 2005) that measures weighted paths of length two in a molecule divided by the number of atoms in the same molecule (Kombo et al. 2013). In graph theory, a path is an alternating sequence of vertices and edges that begins and ends with a vertex without passing by the same vertex more than once. A weighted path assigns a numerical value (weight) to each edge depending on the type of atom existing, thus providing information about the alignment and sequence of atoms and, henceforth, about the physicochemical properties of a molecule (Attenborough 2003). Additionally, molecules with similar WTPT2 values are more likely to share similar structures and, thus, similar potency. Therefore, WTPT2 can be very useful in similarity screening. More importantly, high values of WTPT2 indicate a more compact and branched structure, with shorter average distances between pairs of atoms. This is useful in inhibiting 3CLpro since the latter has multiple sub-sites and a complex shape with several pockets. A more branched complex molecule is likelier to fit and interact with the 3CLpro active site (Jain 2004).
Kier Shape 2
It calculates the Kier shape for paths with a length of two (Sauer and Schwarz 2003). This descriptor provides information about the molecular shape, normalized by the number of atoms relative to the star and linear graphs: the extreme shapes a molecule can take. The shape of a molecule directly influences the structural complementarity with the active site of the target enzyme. It allows the occurrence of intermolecular forces with minimum energy. K2 index quantitates the contribution of the molecular shape to the molecular activity.
Geometrical descriptors
MOMIXZ
A three-dimensional descriptor that measures the moment of inertia along the axes ratio X to Z (Sauer and Schwarz 2003). The moment of inertia generally describes how the mass of a rigid body is distributed. The descriptors MOMY X and MOMY Z measure how hard it is to spin the molecule along the axes X and Z, respectively. The closer the mass is to the axis, the smaller the moment of inertia. MOMIXZ compares the moment of inertia of axis X to Z. In other words, it provides information about the molecule’s orientation when in contact with the active site of 3CLpro. It is also noted that the moment of inertia evaluates properties such as geometry, shape, and conformation (Torkington 1950). This stands for great importance as the activity of inhibitors is directly related to the spatial conformation complementarity between the enzyme’s active site and the molecules. A molecule must be in the proper position and have the corresponding conformation to exhibit its inhibitory effect correctly. Otherwise, it cannot attach to the enzyme.
The analysis of the descriptors indicates that the nature of atoms and their sequence, bonds, and shapes influence their activity. Considering the properties of the active site of 3CLpro, it is logical that the nature of atoms, represented by the number of halogens, matters because it is the engine of all kinds of bonds between the enzyme and the inhibitors. The molecules’ planetary (rotatable bonds, connectivity, shape, autocorrelation, weighted path) and 3-D dimension (moment of inertia) reflect the aspect of structural complementarity, as they consider the shape, the sequence, the neighbouring skeletons, and the rotation among axes.
The contributions of each descriptor are illustrated in Fig. 2. The relative importance of the selected descriptors was calculated using the Mean Effect method (Abdulfatai et al. 2020). The MOMIXZ descriptor had the greatest positive importance, explaining 40% of the activity, followed by the Kier Shape 2, Numberrota table bonds, ATSM2, and finally WTPT2, the SPC-6 had the most significant negative contribution, followed by the Number of hal, with the most negligible relative importance. The results further sustain the above interpretation, indicating that to maximize the inhibitory activity of these nonpeptidic noncovalent molecules, the structural complementarity in terms of mass centred towards the X axis over the Z axis, the presence of rotatable bonds at least 20% in the molecules, the weighted paths of lag 2 and the shape of fragments of lag 2 influence positively the activity. Whereas the number of halogens slightly decreases the activity, the significant branching of molecules (6th degree) hinders the binding with the active site. This explains the inversely proportional relationship governing the degree of complexity of the hypervolume of interaction and the activity.
QSAR models’ development
MLR model
This study developed an MLR model to predict the inhibitory activity of various molecules against SARS-CoV-2, the 3CLpro viral enzyme. The established MLR model (Eq. 2) shows the linear relationship between the selected descriptors and the activity. The coefficients linked to each descriptor represent the weights and the type of contribution (positive or negative). The validation criteria in Table 2 for this model showed its weak ability to predict with relatively low R and R2 values (under the accepted value of R2 < 0.6). The table indicates that the MLR model failed to pass the threshold value for r¯m2 (> 0.5) and ∆rm2 (< 0.2) (Ojha et al. 2011; Euldji et al. 2022). Fig.S-1 shows the regression plot for the train and test sets, where the predicted values do not corroborate with the experimental values. The highest AIC value of the model shown in Table 3 (− 64.9684) indicates that it fitted the experimental data very weakly.
Support vector regression model
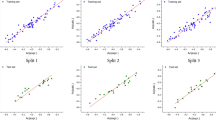
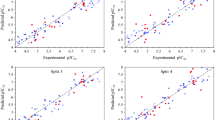
To determine the optimal values of the SVR hyperparameters, a loop was constructed to evaluate the cost function values each time based on the hyperparameters’ incrimination. The chosen Kernel function and the best combination of the hyperparameters were found to be C = 500, γ = 1, ε = 0.0485, and the Radial Basis Function RBF as the kernel function. The predicted values of pIC50 by the SVR method are listed in Table S-II in the supplementary file. The evaluation criteria for the training set and the test set performance of the SVR model are summarized in Table 2. The predicted values of pIC50 of the inhibitors for the training set test set versus their observed values are represented in Fig. 3. The developed model showed a good correlation between the predicted and observed values with low RMSE (0.1367), high R2 (0.9665), and good robustness indicating by the Q2, r¯m2, and ∆rm2 values that satisfy the threshold value. The AIC value of this model (− 249.1234) reveals that the SVR model fitted the experimental data better than the MLR model.
SVR-particle swarm optimization model
The combination of the PSO and SVR algorithms aims to find the optimal variety of the hyperparameters that yields the smallest error values by testing a huge number of possible combinations. The best combination consisted of C = 131.7001, γ = 1, and ε = 0.0350. It is thus a way to avoid the overfitting of the model with a higher R2 coefficient (0.9720)and lower RMSE value (0.1243). The values of Q2, r¯m2 and ∆rm2 of this model also satisfy the threshold value. The value of R2-Q2 for the whole set (both training and test) 0.9650–0.9649 is 0.0001 (accepted value < 0.3), indicating that the model is not overfitted (Euldji et al. 2022). Table 2 shows the statistical parameters for the PSO-SVR model. The AIC value of this model (− 264.5433) indicates that the model could fit the experimental data very well. Figure 4 illustrates the correlation between the predicted pIC50 and the observed values for the training and test sets.
External validation
The calculation of the external validation criteria on the test set of each model shows the performance for SVR and PSO-SVR followed by MLR (R2: 0.9310, 0.8932, and 0.2851respectively) and low RMSEP values (0.1870, 0.2203, and 0.6465 respectively. The external validation criteria evaluated the models’ accuracy and ability to predict unseen data, and the results are demonstrated in Table 4. The results show that the MLR model could not perform well on the test data. This is shown by the low CCC value, the high root mean square error RMSE and the mean absolute error MAE. The PSO-SVR model outperformed the SVR model regarding generalization and prediction ability regarding correlation metrics and error metrics, indicated by higher CCC and Q2F1 values and lower RMSE and MAE values. Generally, the SVR and PSO-SVR models showed good statistical significance and robustness.
Applicability domain
Multiple linear regression
The domain of applicability of the MLR model was analyzed by the William plot (FigS-2). It shows a total distribution of the predicted points in the range of the.applicability domain [− 3, 3], with one influential point exceeding the critical leverage point of 0.35. All the predicted values did not align close, but all fell inside the domain, indicating that the model found no response outliers
Support vector regression
The applicability domain was analyzed using the William Plot, as shown in Fig. 5. a. This shows that about 98.59% of the values are within the applicability domain's critical leverage point range. Also, it was noticed that the whole test set and 96.5% of the training set are included in the domain. Only two points (molecules 1 and 14) are outside the domain with relatively high standardized residual values.
SVR-particle swarm optimization
William’s plot for the PSO-SVR model (Fig. 5 b) shows that one molecule (molecule 55) did exceed the critical leverage value of 0.35, and two other training molecules (molecules 1 and 17) lie outside the standard residual’s boundaries [− 3, 3]. However, most points aligned with the 0 standard residuals, and only a few were distributed in the space domain.
Models’ comparison
The analysis of the applicability domains indicates that despite the existence of a few outliers in the SVR and PSO-SVR models, they remain reliable as the standard residual values of the influential molecules are low, and the majority of the points align with the zero standard residual rather than covering the whole domain. A closer look at the SVR and PSO-SVR models shows that the predicted values by the PSO-SVR align better with zero standard residual value than those predicted by the SVR, and both models had an X outlier. These observations suggest that molecules 55 and 34 for PSO-SVR and SVR models are descriptors’ outliers and, thus, an influential point. Although the presence of outliers is a normal phenomenon, it may be due to the model errors and/or errors associated with experiments. In conclusion, the analysis of the applicability domain confirmed the robustness and the predictivity of the PSO-SVR model.
According to the statistical criteria above, the PSO-SVR model showed the best performance, followed closely by the SVR model with good predictivity. In contrast, the MLR model had relatively weak metrics, resulting in the lowest performance and predictability. While the MLR method aims to build a predictive model, it is also, more importantly, to verify the linearity of the studied phenomenon. It confirms that the studied phenomenon is nonlinear and that the MLR method cannot be applied to this database. Henceforth, using nonlinear methods is a justified need, which explains the better results obtained by the PSO-SVR and SVR algorithms. Using a metaheuristic algorithm to tune the SVR parameters rather than the trial-and-error method is one reason the PSO-SVR method performed better. Moreover, it avoids the overfitting of the model as it allows the model to reach a degree of complexity suitable to that of the studied phenomenon.
Moreover, the SVR model showed a close performance on the training set with the PSO-SVR model with R2SVR = 0.96, R2PSO-SVR = 0.97, RMSESVR = 0.12, and RMSEPSO-SVR = 0.13. This renders it complex to conclude the outperformance of PSO-SVR. However, a closer look at the external validation criteria shows the significant outperformance of PSO-SVR with R2 = 0.93, Q2 = 0.97, and RMSE = 0.18 in contrast to R2 = 0.89, Q2 = 0.91 and RMSE = 0.22 of the SVR model, indicating the superiority of PSO-SVR in generalization and predictive ability of unknown datasets, endorsing the importance of parameters tuning by metaheuristic algorithms. These findings are confirmed by the analysis of the AIC, where the PSO-SVR model had the lowest value, followed by SVR and MLR, indicating that the PSO-SVR performed the best. Furthermore, since the difference in the AIC values between the PSO-SVR and SVR models is greater than 2, the PSO-SVR model can safely be considered significantly better than the SVR model, indicating that having a metaheuristic algorithm improved the prediction results.
Comparison with literature
Many papers were published to provide a predictive model for the 3CLpro inhibitors, using classical algorithms, with acceptable performance. The detailed comparison is demonstrated in Table 5. However, the hybrid PSO-SVR algorithm improved the models’ precision and accuracy and outperformed all the listed models. This confirms the significance of the proposed model in predicting the inhibiting activity against the 3CLpro.This outperformance can be algorithmically explained as follows:
-
Nonlinearity aspect: Unlike MLR, PLS and their hybridization (GA-MLR and PLS-MLR), which apply only to linear problems, SVR can handle complex nonlinear issues with excellent efficiency. Notably, the prediction of the inhibitory activity of molecules is a complex phenomenon, most often nonlinear, and thus requires a strong nonlinearity potential and the ability to adapt to its level of complexity, which linear algorithms fail to have. This is why SVR outperformed linear models such as MLR and PLS.
-
Generalization and overfitting: SVR and ANN have their strengths and weaknesses; SVR can be considered better than ANN when applied to small datasets. ANN tends to overfit when trained on a small dataset since it has many parameters. When the number of parameters is large enough compared to the number of observations, ANN tends to learn the noise instead of the underlying patterns, which affects its performance on the test/external datasets. On the contrary, SVR has fewer parameters to tune and regularization parameters, ensuring avoiding overfitting problems and a good performance on unseen datasets (Byvatov et al. 2003). In Computer-Aided Drug Design (CADD) and QSAR modelling for identifying SARS-CoV-2 inhibitors (or a novel pandemic or disease), little is known about potential molecules, and derivatives and analogues are designed in a limited number. Therefore, datasets of molecules against novel diseases like COVID-19 tend to be small and risk being overfitted if modelled by ANN. This explains why the proposed model outperformed the ANN-reported model.
-
Flexibility and ability to handle outliers: Hologram QSAR (HQSAR), Comparative Molecular Field Analysis (CoMFA) 3D QSAR and Monte Carlo techniques are based on different principles (images of electronic structures and steric properties of molecules for HQSAR (Adhikari et al. 2022), deriving quantitative relationships between the activity and the structure by comparing the electrostatic and steric interactions of molecules and a reference drug for CoMFA (Chedadi et al. 2021), and evolving randomly to generate several models and selecting the best one for Monte Carlo (Toropov et al. 2022)). However, they are similar in how they differ from SVR: they use images and interactions as input instead of comparison or simulation instead of numerical values of molecular descriptors and rely on comparing molecular similarity or fitting a mathematical function to data instead of supervised machine learning. SVR uses less complex inputs and can adapt to different levels of complexity thanks to the kernel functions and perform better on nonlinear phenomena. Also, thanks to the support vectors and the boundaries in SVR, it handles outliers more effectively than the methods above, which can be very sensitive to outliers. This is useful, especially when handling heterogeneous datasets.
-
PSO robustness: Particle Swarm Optimization (PSO) is used to tune the parameters of machine learning algorithms. It is advantageous over other metaheuristic algorithms such as Genetic Algorithm GA since it is less sensitive to the initialization parameters, unlike GA, which is highly influenced by them and, therefore, less robust than PSO. PSO allows a large search space leading to a fine-tuning of parameters and, thus, a better convergence, in addition to a short computational time. Algorithmically, this is the reason why the proposed PSO-SRV model outperformed the reported GA-SVR model (Siddique et al. 2018).
While this outperformance has algorithmic reasons, it is worth clarifying that comparing different algorithms is very complex and requires a thorough, comprehensive analysis of various factors, mainly database similarity, size and quality. The comparison was elaborated for indicative purposes and was based on the similarity of the target 3CLpro solely to position the proposed SVR-PSO in the range of errors. Therefore, it does not imply the absolute superiority of SVR-PSO over other algorithms but rather in this proposed context.
Conclusion
In this work, three models were built to predict the inhibitory activity of potentially therapeutic molecules to contribute to the acceleration of drug discovery by the mean of computational chemistry. The results indicate that the relation between the IC50 and the selected descriptors is nonlinear and thus needs a nonlinear approach to model it. Among the three algorithms, the PSO-SVR model performed best with the lowest errors and highest regression coefficients. The PSO-SVR model obtained was demonstrated to be a robust, statistically and biologically meaningful model capable of predicting the activity of unknown potential inhibitors. This paper’s results also revealed the key descriptors to maximize the activity of the selected molecules, among which the structural complementarity characteristics in terms of the moment of inertia, rotatable bonds, and weighted paths were found proportional to the activity.
On the other hand, a very complex hypervolume of reactivity may hinder the binding of the molecule and the active site. Further docking and virtual screening studies are being investigated based on the findings of this article. The results obtained in this article have promising aspects in designing a potential drug against COVID-19.
References
Abdulfatai U, Uzairu A, Shallangwa GA, Uba S (2020) Designing and estimating antioxidant properties of some lubricant additives via QSPR and MD methodologies. Sci African. https://doi.org/10.1016/j.sciaf.2020.e00451
Adhikari N, Banerjee S, Baidya SK et al (2022) Ligand-based quantitative structural assessments of SARS-CoV-2 3CLpro inhibitors: an analysis in light of structure-based multi-molecular modeling evidences. J Mol Struct. https://doi.org/10.1016/j.molstruc.2021.132041
Alaran AJ, Adebisi YA, Badmos A et al (2021) Uneven power dynamics must be levelled in COVID-19 vaccines access and distribution. Public Heal Pract. https://doi.org/10.1016/j.puhip.2021.100096
Aljoundi A, Bjij I, Rashedy A, Soliman M (2020) Covalent versus noncovalent enzyme inhibition: which route should we take? A justification of the good and bad from molecular modelling perspective. Protein J. https://doi.org/10.1007/s10930-020-09884-2
Alkabbani H, Ahmadian A, Zhu Q, Elkamel A (2021) Machine learning and metaheuristic methods for renewable power forecasting: a recent review. Front Chem Eng. https://doi.org/10.3389/fceng.2021.665415
Attenborough M (2003) 19—Graph theory. In: Attenborough MBT-M for EE and C (ed). Newnes, Oxford. 461–478
Awad M, Khanna R (2015) Support Vector Regression. Efficient learning machines: theories, concepts, and applications for engineers and system designers. Apress, Berkeley, CA, pp 67–80
Byvatov E, Fechner U, Sadowski J, Schneider G (2003) Comparison of support vector machine and artificial neural network systems for drug/nondrug classification. J Chem Inf Comput Sci. https://doi.org/10.1021/ci0341161
Chedadi O, Aissouq A, El Ouardi Y et al (2021) In silico prediction of novel SARS-CoV 3CL pro inhibitors: a combination of 3D-QSAR, molecular docking, ADMET prediction, and molecular dynamics simulation. Biointer Res Appl Chem. https://doi.org/10.33263/BRIAC124.51005115
Ciotti M, Ciccozzi M, Terrinoni A, et al (2020) The COVID-19 pandemic. Crit. Rev. Clin. Lab. Sci. 365–388
Calvet L, de Armas J, Masip D, Juan AA (2017) Learnheuristics: hybridizing metaheuristics with machine learning for optimization with dynamic inputs. Open Math. https://doi.org/10.1515/math-2017-0029
Almeida BSG de (2019) In: Osaba VCLE-JDSE-EVE-E (ed) Particle Swarm Optimization: A Powerful Technique for Solving Engineering Problems. IntechOpen, Rijeka
De P, Bhayye S, Kumar V, Roy K (2020) In silico modeling for quick prediction of inhibitory activity against 3CLpro enzyme in SARS CoV diseases. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2020.1821779
Dong J, Cao DS, Miao HY et al (2015) ChemDes: an integrated web-based platform for molecular descriptor and fingerprint computation. J Cheminform. https://doi.org/10.1186/s13321-015-0109-z
Dongliang Y, Ejaz S, Aziz M et al (2022) Benzene-1,3-diol derivatives as the inhibitors of butyrylcholinesterase: an emergent target of Alzheimer’s disease. J Serbian Chem Soc. https://doi.org/10.2298/jsc210416073d
Edache EI, Uzairu A, Mamza PA, Shallangwa GA (2022) QSAR, homology modeling, and docking simulation on SARS-CoV-2 and pseudomonas aeruginosa inhibitors, ADMET, and molecular dynamic simulations to find a possible oral lead candidate. J Genet Eng Biotechnol. https://doi.org/10.1186/s43141-022-00362-z
Elekofehinti OO, Iwaloye O, Molehin OR, Famusiwa CD (2021) Identification of lead compounds from large natural product library targeting 3C-like protease of SARS-CoV-2 using E-pharmacophore modelling, QSAR and molecular dynamics simulation. Silico Pharm. https://doi.org/10.1007/s40203-021-00109-7
Estrada E (2002) Physicochemical interpretation of molecular connectivity indices. J Phys Chem A. https://doi.org/10.1021/jp026238m
Euldji I, Si-Moussa C, Hamadache M, Benkortbi O (2022) QSPR modelling of the solubility of drug and drug-like compounds in supercritical carbon dioxide. Mol Inform. https://doi.org/10.1002/minf.202200026
Falyouna O, Eljamal O, Maamoun I et al (2020) Magnetic zeolite synthesis for efficient removal of cesium in a lab-scale continuous treatment system. J Colloid Interface Sci. https://doi.org/10.1016/j.jcis.2020.03.028
Feinmann J (2021) How the world is (not) handling surplus doses and expiring vaccines. BMJ. https://doi.org/10.1136/bmj.n2062
Fu L, Ye F, Feng Y et al (2020) Both Boceprevir and GC376 efficaciously inhibit SARS-CoV-2 by targeting its main protease. Nat Commun. https://doi.org/10.1038/s41467-020-18233-x
Guha R, Jurs PC (2005) Interpreting computational neural network QSAR Models: a measure of descriptor importance. J Chem Inf Model. https://doi.org/10.1021/ci050022a
Hamadache M, Hanini S, Benkortbi O et al (2016) Artificial neural network-based equation to predict the toxicity of herbicides on rats. Chemom Intell Lab Syst. https://doi.org/10.1016/j.chemolab.2016.03.007
Hamadache M, Benkortbi O, Hanini S, Amrane A (2017) Application of multilayer perceptron for prediction of the rat acute toxicity of insecticides. Energy Procedia. https://doi.org/10.1016/j.egypro.2017.11.169
Hammoudan I, Matchi S, Bakhouch M, et al (2021) QSAR Modelling of Peptidomimetic Derivatives towards HKU4-CoV 3CL pro Inhibitors against MERS-CoV. 391–401
Han SH, Goins CM, Arya T et al (2021) Structure-based optimization of ml300-derived, noncovalent inhibitors targeting the severe acute respiratory syndrome coronavirus 3CL protease (SARS-CoV-2 3CLpro). J Med Chem. https://doi.org/10.1021/acs.jmedchem.1c00598
Harapan H, Itoh N, Yufika A et al (2020) Coronavirus disease 2019 (COVID-19): a literature review. J Infect Public Health 13:667–673
Ishola AA, Adedirin O, Joshi T, Chandra S (2021) QSAR modeling and pharmacoinformatics of SARS coronavirus 3C-like protease inhibitors. Comput Biol Med. https://doi.org/10.1016/j.compbiomed.2021.104483
Jain AN (2004) Ligand-based structural hypotheses for virtual screening. J Med Chem. https://doi.org/10.1021/jm030520f
Janairo GIB, Yu DEC, Janairo JIB (2021) A machine learning regression model for the screening and design of potential SARS-CoV-2 protease inhibitors. Netw Model Anal Heal Informatics Bioinforma. https://doi.org/10.1007/s13721-021-00326-2
Jawarkar RDD, Bakal RL, Zaki MEAA et al (2022) QSAR based virtual screening derived identification of a novel hit as a SARS CoV-229E 3CLpro Inhibitor: GA-MLR QSAR modeling supported by molecular docking, molecular dynamics simulation and MMGBSA calculation approaches. Arab J Chem. https://doi.org/10.1016/j.arabjc.2021.103499
John SES, Tomar S, Stauffer SR, Mesecar AD (2015) Targeting zoonotic viruses: Structure-based inhibition of the 3C-like protease from bat coronavirus HKU4—The likely reservoir host to the human coronavirus that causes Middle East Respiratory Syndrome (MERS). Bioorg Med Chem. https://doi.org/10.1016/j.bmc.2015.06.039
Khanna V, Ranganathan S (2009) Physiochemical property space distribution among human metabolites, drugs and toxins. BMC Bioinformatics. https://doi.org/10.1186/1471-2105-10-S15-S10
Kirmani SAK, Ali P, Azam F (2021) Topological indices and QSPR/QSAR analysis of some antiviral drugs being investigated for the treatment of COVID-19 patients. Int J Quantum Chem. https://doi.org/10.1002/qua.26594
Kombo DC, Tallapragada K, Jain R et al (2013) 3D molecular descriptors important for clinical success. J Chem Inf Model. https://doi.org/10.1021/ci300445e
Kortagere S, Ekins S, Welsh WJ (2008) Halogenated ligands and their interactions with amino acids: implications for structure-activity and structure-toxicity relationships. J Mol Graph Model. https://doi.org/10.1016/j.jmgm.2008.04.001
Laidi M, Abdallah el Hadj A, Si-Moussa C et al (2020) CMC of diverse Gemini surfactants modelling using a hybrid approach combining SVR-DA. Chem Ind Chem Eng Q. https://doi.org/10.2298/ciceq200907048l
Liu B, Jin C, Wan J et al (2018) Modelling and optimizing an electrochemical oxidation process using artificial neural network, genetic algorithm and particle swarm optimization. J Serbian Chem Soc. https://doi.org/10.2298/JSC170721101L
Mitikiri P, Jana G, Sural S, Chattaraj PK (2018) A machine learning technique toward generating minimum energy structures of small boron clusters. Int J Quantum Chem. https://doi.org/10.1002/qua.25672
Mozafari Z, Chamjangali MA, Arashi M, Goudarzi N (2021) Suggestion of active 3-chymotrypsin like protease (3CL Pro ) inhibitors as potential anti-SARS-CoV-2 agents using predictive QSAR model based on the combination of ALASSO with an ANN model. SAR QSAR Environ Res. https://doi.org/10.1080/1062936X.2021.1975167
Ojha PK, Mitra I, Das RN, Roy K (2011) Further exploring rm2 metrics for validation of QSPR models. Chemom Intell Lab Syst. https://doi.org/10.1016/j.chemolab.2011.03.011
Peng Y, Nagata MH (2020) An empirical overview of nonlinearity and overfitting in machine learning using COVID-19 data. Chaos, Solitons Fractals. https://doi.org/10.1016/j.chaos.2020.110055
Probst P, Boulesteix AL, Bischl B (2019) Tunability: importance of hyperparameters of machine learning algorithms. J Mach Learn Res. https://doi.org/10.48550/arXiv.1802.09596
Roy K, Ambure P (2016) The “double cross-validation” software tool for MLR QSAR model development. Chemom Intell Lab Syst. https://doi.org/10.1016/j.chemolab.2016.10.009
Roy K, Kar S, Das R (2015a) A primer on QSAR/QSPR modeling: fundamental concepts. Springer, Berlin, Germany
Roy K, Das RN, Ambure P, Aher RB (2016) Be aware of error measures. Further studies on validation of predictive QSAR models. Chemom Intell Lab Syst. https://doi.org/10.1016/J.CHEMOLAB.2016.01.008
Roy K, Kar S, Das RN (2015b) In: Roy K, Kar S, Das RNBT-U the B of Q for A in PS and RA (eds). Validation of QSAR Models. Academic Press. Boston
Rupp M (2015) Machine learning for quantum mechanics in a nutshell. Int J Quantum Chem. https://doi.org/10.1002/qua.24954
Sander T, Freyss J, von Korff M, Rufener C (2015) DataWarrior: an open-source program for chemistry aware data visualization and analysis. J Chem Inf Model. https://doi.org/10.1021/ci500588j
Sauer WHB, Schwarz MK (2003) Molecular shape diversity of combinatorial libraries: a prerequisite for broad bioactivity. J Chem Inf Comput Sci. https://doi.org/10.1021/ci025599w
Shen Q, Jiang JH, Jiao CX et al (2004) Modified particle swarm optimization algorithm for variable selection in MLR and PLS modeling: QSAR studies of antagonism of angiotensin II antagonists. Eur J Pharm Sci. https://doi.org/10.1016/j.ejps.2004.03.002
Siddique M, Mohanty S, Panda D (2018) A hybrid forecasting model for prediction of stock value of tata steel using support vector regression and particle swarm optimization. Int J Pure Appl Math 119:1719–1728
Tahir Ul Qamar M, Alqahtani SM, Alamri MA, Chen LL (2020) Structural basis of SARS-CoV-2 3CLpro and anti-COVID-19 drug discovery from medicinal plants. J Pharm Anal. https://doi.org/10.1016/j.jpha.2020.03.009
Titov OI, Shulga DA, Palyulin VA, Zefirov NS (2015) Perspectives of halogen bonding description in scoring functions and QSAR/QSPR: substituent effects in aromatic core. Mol Inform. https://doi.org/10.1002/minf.201400195
Torkington P (1950) The moments of inertia of molecules with internal rotation. J Chem Phys 10(1063/1):1747651
Toropov AA, Toropova AP, Veselinović AM et al (2022) SARS-CoV Mpro inhibitory activity of aromatic disulfide compounds: QSAR model. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2020.1818627
Uyanık GK, Güler N (2013) A study on multiple linear regression analysis. Procedia Soc Behav Sci. https://doi.org/10.1016/j.sbspro.2013.12.027
Veber DF, Johnson SR, Cheng HY et al (2002) Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem. https://doi.org/10.1021/jm020017n
Wihartiko FD, Wijayanti H, Virgantari F (2018) Performance comparison of genetic algorithms and particle swarm optimization for model integer programming bus timetabling problem. IOP Conf Ser Mater Sci Eng. https://doi.org/10.1088/1757-899X/332/1/012020
Yan F, Gao F (2021) An overview of potential inhibitors targeting non-structural proteins 3 (PLpro and Mac1) and 5 (3CLpro/Mpro) of SARS-CoV-2. Comput Struct Biotechnol J 19:4868–4883
Yu X (2021) Prediction of inhibitory constants of compounds against SARS-CoV 3CLpro enzyme with 2D-QSAR model. J Saudi Chem Soc. https://doi.org/10.1016/j.jscs.2021.101262
Zhai T, Zhang F, Haider S et al (2021) An integrated computational and experimental approach to identifying inhibitors for SARS-CoV-2 3CL protease. Front Mol Biosci. https://doi.org/10.3389/fmolb.2021.661424
Funding
No funding was received to assist with the preparation of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Madani, A., Benkortbi, O. & Laidi, M. In silico prediction of the inhibition of new molecules on SARS-CoV-2 3CL protease by using QSAR: PSOSVR approach. Braz. J. Chem. Eng. 41, 427–442 (2024). https://doi.org/10.1007/s43153-023-00332-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43153-023-00332-z