Abstract
Epilithic biofilms are ubiquitous in large river environments and are crucial for biogeochemical processes, but their community structures and functions remain poorly understood. In this paper, the seasonal succession in the morphological structure and the taxonomic composition of an epilithic bacterial biofilm community at a polluted site of the Danube River were followed using electron microscopy, high-throughput 16S rRNA gene amplicon sequencing and multiplex/taxon-specific PCRs. The biofilm samples were collected from the same submerged stone and carried out bimonthly in the littoral zone of the Danube River, downstream of a large urban area. Scanning electron microscopy showed that the biofilm was composed of diatoms and a variety of bacteria with different morphologies. Based on amplicon sequencing, the bacterial communities were dominated by the phyla Pseudomonadota and Bacteroidota, while the most abundant archaea belonged to the phyla Nitrososphaerota and Nanoarchaeota. The changing environmental factors had an effect on the composition of the epilithic microbial community. Critical levels of faecal pollution in the water were associated with increased relative abundance of Sphaerotilus, a typical indicator of “sewage fungus”, but the composition and diversity of the epilithic biofilms were also influenced by several other environmental factors such as temperature, water discharge and total suspended solids (TSS). The specific PCRs showed opportunistic pathogenic bacteria (e.g. Pseudomonas spp., Legionella spp., P. aeruginosa, L. pneumophila, Stenotrophomonas maltophilia) in some biofilm samples, but extended spectrum β-lactamase (ESBL) genes and macrolide resistance genes could not be detected.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The drinking water supply of Budapest with a population of almost 3 million people (including the agglomeration) is predominantly based on wells with coastal filtration. Beyond drinking water, the Danube River plays an important role in water transport (riverine transport route), tourism, recreation and even in food supply (fishing). The river covers approximately 801, 500 km2 and is connected to 19 countries. Therefore, the importance of the Danube River is reflected in the many international conventions, research and monitoring programmes established to protect it and its catchment area (e.g. Joint Danube Survey [JDS], the TransNational Monitoring Network, Modelling Nutrient Emissions in River Systems), which are coordinated/organised by the ICPDR (International Commission for the Protection of the Danube River, https://www.icpdr.org/main/).
Thanks to their metabolic activity, biofilms play a key role in the self-purification and detoxification of our natural waters and in the processes of degradation and mineralisation in aquatic environments as it happens also during the gravel-filtration processes for drinking water production along the Danube River (Makk et al. 2003a; Vargha et al. 2005; Sharma 2022).
Research to understand the bacterial communities in the rolling gravel samples of the Danube sediments has been already studied between 1994 and 95. With cultivation methods, Pseudomonas spp., facultative methylotrophic and H2 autotrophic Hydrogenomonas, Xanthomonas, Xanthobacter spp. were detected as the dominant members of heterotrophic biofilms, and in hypertrophic conditions (at high number of algae) members of the genus Aeromonas appeared to be significant (Márialigeti et al. 1997; Zalmum 1997; Salem 1997). It was found that characteristic bacterial communities were associated also to the surface of diatoms in different biofilms. Electron micrographs showed that numerous rods, curved rods, cocci, filamentous, sometimes elongated bacteria attached to the surface of the diatoms and filamentous algae (Makk and Ács 1997; Makk et al. 2003a).
The structure, spatial and year-round temporal variations of the planktonic and epilithic prokaryotic communities of Danube River were studied recently by next-generation sequencing up- and downstream of Budapest. Samples were taken between February of 2019 and January of 2020. It was found that the composition of planktonic communities clearly followed the prolongation of the summer resulting from climate change, while epilithic communities appeared to be less responsive to the changes (Engloner et al. 2023).
Recently, research on the water quality of the Danube has been highlighting new threats in addition to the already known pollutants (e.g. natural organic matter, nitrite/nitrate, heavy metals and thermal pollution). Effluents of wastewater treatment plants play a role in the spread of chemical contaminants, pathogenic bacteria, antibiotics and the emergence of antibiotic resistance in urban rivers (Alygizakis et al. 2019; Sabri et al. 2020; Marutescu et al. 2023). As a result of increasing drug consumption, many pharmacologically active compounds enter our natural waters as organic micropollutants (e.g. antibiotics and their derivatives). The spread of antibiotic resistance among microbes is increasing worldwide, posing a growing health risk to humans and animals, too (Ferri et al. 2017). The research is often focused to determine the antibiotic-resistant organisms and their antibiotic resistance genes (ARGs) (Banciu et al. 2021; Kittinger et al. 2016a, 2018; Schachner-Groehs et al. 2024) in different aquatic habitats.
Biofilms formed on natural substrates (e.g. gravel, stones, leaves or feathers) can gather ARGs or pathogens and facilitate their spread in the rivers (Kneis et al. 2022; Liu et al. 2023). ARGs can be rapidly transmitted by horizontal gene transfer, which has been shown to occur more frequently in biofilms than in planktonic states (Madsen et al. 2012). Several studies have shown that biofilms can facilitate the spread of antibiotic resistance in the environment (Michaelis and Grohmann 2023).
The increasing use of next-generation sequencing (NGS) techniques has led to a growing knowledge of the diversity of epilithic biofilm communities (Zancarini et al. 2017; Lin et al. 2019; Mao et al. 2021; Pin et al. 2021; Engloner et al. 2023). Indeed, in large rivers such as the Danube, our knowledge of the temporal dynamics of microbial communities forming on the surface of the same site (e.g. on same submerged stones) is limited and mostly based on cultivation-based studies. The occurrence of potential pathogens or ARGs in such types of biofilms in the Danube River has also been little investigated so far. The aim of the present study was to use an epilithic biofilm community on the same stone located at a polluted site as bio-indicator of anthropogenic pollution. This was done by (i) assessing seasonal changes in the bacterial and archaeal community composition using amplicon sequencing, (ii) visualising the structure of the biofilms using scanning electron microscopy, (iii) detecting the presence of representative facultative pathogenic bacteria and antibiotic resistance genes and (iv) relating these data with a set of environmental parameters including information on anthropogenic pollution.
Materials and methods
Sampling
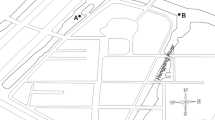
The epilithic biofilms were originated from a Danube River site downstream of the city of Budapest, at 1632 river km and right bank (GPS coordinates: 47.390717N, 19.013607E). This sampling point is characterised by significant anthropogenic influence as it is located downstream of a large city with > 2 million inhabitants and upstream inflow of a small polluted stream (Rákos). This site also coincided with one of the sampling points of JSD4 (Joint Danube Survey 4) (H3R sampling point, Schachner-Groehs et al. 2024) series downstream of Budapest. The regular sampling was conducted bimonthly during the study: 13 October 2020, 8 December 2020, 16 February 2021, 08 April 2021 and 15 June 2021 (referred to as Oct-B, Dec-B, Feb-B, Apr-B and Jun-B samples, respectively). Biofilm samples were collected from upper surface of the same submerged large stone each time. After the stone was carefully removed from the water until the time of sampling, the biofilm biomass was scraped off a c.a. 3 cm2 surface using a sterile knife and tweezers in four replicates. After sampling, the stone was placed back each time to the same position. Half of the biofilm samples were placed in a sterile Falcon tube for molecular studies, while the other half were placed in 5-ml plastic tube containing a 5% (v/v) glutaraldehyde solution, a fixative for electron microscopy studies. One litre of water samples (referred to as Oct-W, Dec-W, Feb-W, Apr-W and Jun-W samples) was collected aseptically in sterilised screw-capped flasks from 10 cm below the surface according to ISO 19458:(2006). Samples were transported to the laboratory in a cool bag (at 10 °C) and processed within 4 h after sampling.
Physical and chemical parameters, total suspended solids, water level and discharge data
At the samplings, the water temperature, pH, electrical conductivity and oxygen concentration values were measured on site with the handheld multi-parameter meter Multi 3430 (WTW, Xylem Analytics, Germany). Total suspended solids (TSS) and inorganic particulate matter were determined during the annual cycle according to Eiler et al. (2003). Water discharge and level data were obtained from national monitoring systems for the sampling site (https://www.hydroinfo.hu) (Fig. 1). For quantification of chemical elements, a double-focusing sector field high-resolution inductively coupled plasma mass spectrometer (HR-ICP-MS), ELEMENT2 from Thermo Finnigan (Bremen, Germany) was used. Detailed information on the standards and HR-ICP-MS measurements was detailed in the article by Schachner-Groehs et al. (2024).
The water level and water discharge of the Danube River near Budapest at the sampling times (https://www.hydroinfo.hu)
Enumeration of the faecal indicator Escherichia coli (E. coli) and bacterial total cell count
Number of E. coli of the water samples was determined according to ISO 9308-2:2012 (International Organisation for Standardisation 2012) using the Colilert-18 (97-well Quanty Tray 2000; IDEXX, Ludwigsburg, Germany). Samples were incubated at 36 ± 1 °C for 18–22 h and analysed in a UV cabinet. Quantitative values were obtained by comparing the results with the Most Probable Number (MPN) table provided by the manufacturer.
In order to assess the degree of faecal pollution of the water samples, faecal indicators were classified according to a system of 5 microbiological water quality categories (little, moderate, critical, strong, and excessive pollution) as described by Kirschner et al. (2009).
To determine the total bacterial cell count (TCC), one mL of water sample was fixed with sterile paraformaldehyde (PFA) at a final concentration of 0.8% and filtered through an Anodisc filter (pore size 0.2 µm, Whatman, Germany) and stained with 1:400 diluted SYBR gold solution (Van Driezum et al. 2018). TCC was determined using a Nikon Eclipse 8000 epifluorescence microscope by counting at least 200 bacterial cells in 10 microscopic fields per filter (Velimirov et al. 2011).
Sample preparation for electron microscopy
Scanning electron microscopy (SEM) was used to observe the morphological characteristics of the biofilm samples. The collected samples were immediately fixed in situ in 5% glutaraldehyde solution (0.2 M Na–K phosphate buffer, ~ pH = 7) for 3 h. The samples were then rinsed twice for 10 min in 0.1 M Na–K phosphate buffer (~ pH = 7), then frozen at –85 °C. The samples were freeze dried in an Edward freeze-dryer for 5–10 h at 2 × 10−2 mbar, at –60 °C. The dried and gold sputter-coated samples were examined using an EVO MA 10 Zeiss scanning electron microscope at 10 kV.
Extraction of environmental DNA
The community DNA was isolated from the biofilm samples using the DNeasy PowerSoil® Kit (Qiagen, Hilde, Germany). The biofilm samples were vortexed and centrifuged at 4500 × g for 25 min. The resulting pellet was loaded onto the PowerBead Tube of the DNA isolation kit. The DNA isolation was performed according to the manufacturer’s descriptions. The concentration of 5 µL of the isolated DNA was measured using the Qubit dsDNA HS Assay Kit (Thermo Scientific) and the PCR (polymerase chain reaction) master mix was adjusted according to the measured DNA concentrations.
Next-generation DNA sequencing
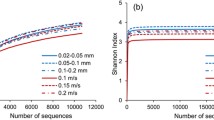
The V3–V4 region of the 16S rRNA gene was used for prokaryotic taxonomic diversity analysis. PCR amplification was performed in triplicate using primers with the following target-specific sequences: Bact-341F (5’-CCT ACG GGN GGC WGC AG-3’) and Bact-805NR (5’-GAC TAC NVG GGT ATC TAA TCC-3’) for Bacteria (Herlemann et al. 2011) and Arch-519F (5’-CAG CMG CCG CGG TAA-3’) (Øvreas et al. 1997) and Arch-855R (5’-TCC CCC GCC AAT TCC TTT AA-3’) (Gray et al. 2002) for Archaea. In both cases, the first section of the primers 5’ end contained Fluidigm CS1 and CS2 adapter sequences. DNA sequencing was performed on an Illumina MiSeq platform (Illumina Inc., San Diego, USA) at the Genomics Core, Research Technology Support Facility (Michigan State University, Trowbridge, USA) using a MiSeq Reagent Kit v2 (500 cycle, 2 × 250 bp paired end). Sequencing and the description of bioinformatic analysis were done according to Lange-Enyedi et al. (2023). Operational taxonomic units (OTUs) were determined at the 97% similarity threshold levels according to Tindall et al. (2010). Raw sequence data can be accessed under the NCBI BioProject ID PRJNA1067519 in the Sequence Read Archive (SRA) database (https://submit.ncbi.nlm.nih.gov/subs/sra/). Good’s coverage, Shannon and inverse Simpson indices were calculated using mothur programme (Schloss et al. 2009), and reads were subsampled to the read number of the sample having the lowest sequence count. The relative abundances of different genus level taxa of the samples were compared using a Bray–Curtis similarity index; UPGMA (unweighted pair group method using arithmetic averages) dendrogram was created using the PAST3 (PAleontological STatistics) software (Hammer et al. 2001). To analyse the effect of environmental properties on the bacterial community composition, NMDS (Non-metric Multidimensional Scaling) analysis was used. The study was performed at OTU-level based on Bray–Curtis dissimilarities using functions of the vegan (Oksanen et al. 2020) and ggplot2 packages in R (R Core Team 2018).
Taxon-specific PCRs
Taxon-specific PCRs were applied for the following bacterial groups: coliforms according to Bej et al. (1990), Pseudomonas spp. according to Spilker et al. (2004), Pseudomonas aeruginosa according to Lavenir et al. (2007), Legionella spp. according to Cloud et al. (2000), Legionella pneumophila according to Fiume et al. (2005) and Stenotrophomonas maltophilia according to da Silva Filho et al. (2004).
Multiplex PCRs
The presence of extended-spectrum β-lactamase (ESBL) genes (blaSHV, blaTEM and blaCTX-M) and macrolide resistance genes (ermA, ermB, ermC, msrA and mef) was tested in all biofilm samples using multiplex PCR according to the protocol of Trung et al. (2015) and Zmantar et al. (2011).
Results
Physico‑chemical and biological parameters
The physico-chemical parameters together with the E. coli MPN and total cell count values of the water samples from the Danube River are given in Tables 1 and 2. At the time of sampling, the main differences in the physical and chemical properties of the water at the sampling site occurred in water discharge, temperature and the amount of total suspended solids (TSS). The Danube water was aerobic at all sampling times, and the majority of the TSS appeared as particulate inorganic matter (PIM). According to the Colilert-18 study, the Danube water was critically polluted in October 2020 (E. coli 3500 MPN/100 ml), while low levels of pollution were detected at other sampling times (E. coli 344–55 MPN/100 ml). Some of the chemical elements (e.g. P, Al, Co, Cr, Cu, Fe, Ni, Pb, V, Zn, Gd) showed higher concentrations in the Oct-W, Feb-W and Jun-W samples.
Electron microscopic analysis
Scanning electron microscopic images of the biofilm samples revealed complex bacterial communities and diatoms with a wide variety of morphologies (Fig. 2a–j). At different times, the analysis showed a multitude of filamentary formations of different diameters and lengths. The images have revealed also network structures formed by filamentous microbes (bacteria, diatoms, e.g. Melosira varians), which could be the basis for the developing of large slimy coatings on the surfaces (up to several mm in size) by the microorganisms. Higher-resolution electron micrographs showed that the surfaces of the diatoms colonising the stone were a suitable habitat for a variety of bacteria (e.g. Figure 2d, e, h). On some sampling dates, the biofilms were less visible because they were covered by large amounts of small debris (Oct-B, Feb-B and Jun-B samples).
Next‑generation sequencing
The bioinformatic analysis identified a total of 79,633 sequences from the Bacteria domain, which could be classified into 5942 OTUs. In comparison, only 154 OTUs and a much smaller number of 5672 sequences from the Archaea domain were detected.
In the case of bacteria, the number of observed OTUs showed that the Oct-B, Feb-B and Jun-B samples had a higher species richness (1520–1733) than the Dec-B and Apr-B samples (434 and 661). According to different diversity indices, Dec-B and Apr-B samples had the least diverse bacterial community (Table 3). Regarding Archaea, the samples were less species-rich (number of OTUs: 22–39). As the low obtained archaeal sequence number gave a brief overview on the archaeal community structure, the diversity indices of archaea were disregarded.
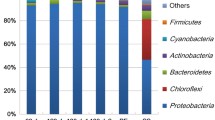
The members of the bacterial OTUs with a relative abundance of more than 1% could be classified into 9 phyla, while the OTUs belonging to the Archaea domain could be classified into 6 phyla (Fig. 3). The composition of the bacterial and archaeal communities was dominated by previously uncultured and unidentified microbes at the genus, family, order, and class level (Figs. 3, 5). In addition, a high percentage (43.3‒50.5%) of the communities were represented by genera of the bacteria with a relative abundance of less than 1%. This was especially typical for samples with high diversity based on the diversity indices (Oct-B, Feb-B, Jun-B samples) (Fig. 5a). Cluster analysis based only on the bacterial taxa showed that samples from February, October and June are grouped closely together while Dec-B is clearly separated (Fig. 4).
Overall, based on the number of the sequences detected, the phyla Pseudomonadota (synonym Proteobacteria) (47.8–82.1%) and Bacteroidota (15.6–26.3%) were the most abundant at all sampling times. However, differences in relative abundances were observed already at this level. The phylum Pseudomonadota (with 82.1% relative abundance) was the absolute dominant in the Dec-B sample, while the phylum Bacteroidota had the highest relative abundance (26.3%) in the Feb-B sample. The phylum Cyanobacteria could be detected in higher numbers in the Oct-B and Jun-B samples (8.1% and 4.5%, respectively). The representatives of the phyla Acidobacteriota, Actinomycetota, Gemmatimonadota, Patescibacteria, Planctomycetota and Verrucomicrobia were detected in higher proportion in the Oct-B, Feb-B and Jun-B samples than in the Dec-B and Apr-B samples. Note that, with the exception of Actinomycetota and Verrucomicrobia, the representatives of the latter phyla showed a relative abundance of less than 1% in the Dec-B and Apr-B biofilm samples.
Regarding the lower taxonomic levels, from the phylum Pseudomonadota, the following 5 taxonomic groups were prominent (˃ 4%) in all samples, but with different relative abundance values: unc. Burkholderiaceae (7.7–47.0%), unc. Burkholderiaceae CM1G08 (0.0–4.7%), genera Hydrogenophaga (0.3–4.4%), Rhodoferax (0.8–8.0%) and Sphaerotilus (0.6–60.4%). Within this phylum, the occurrence of taxonomic groups with a lower relative abundance (between 1 and 4%) showed a large variation between the sampling times. The highest % abundance was achieved by the genera Sphingomonas (2.9%), Massilia (2.1%), Pseudomonas (1.0%) and unc. Gammaproteobacteria (1.7%) in the Oct-B samples; unc. Sphingomonadaceae (2.0%) and genus Rhizobacter (2.5%) in the Feb-B sample; genera Cellvibrio (1.9%), Halioglobus (1.1%) and Halieaceae OM60 (NOR5) clade (2.0%) in the Jun-B sample. From the phylum Bacteroidota, only the genus Flavobacterium was present in high proportions at different sampling times (3.9–13.8%), the other representatives having low relative abundances: genus Hymenobacter (1.7%) in the Oct-B sample; family Spirosomaceae (1.0%) in the Dec-B sample; genus Pedobacter (1.5%) in the Feb-B sample; unc. Bacteroidia (1.2%) in the Apr-B sample; unc. Saprospiraceae (1.4%) and family Microscillaceae OLB12 (1.4%) in the Jun-B sample. One genus and one order from the phylum Cyanobacteria were detected mainly in the Oct-B and Jun-B samples. These taxa belonged to the order Nostocales (unc., 0.3–1.4%) and the genus Potamolinea 1PC (3.4–3.8%). From the phylum Verrucomicrobia, the genus Luteolibacter (0.04–4.4%) was present in high proportion. Other Bacteroidota taxa of the biofilm sample were also identified as unc. Pedosphaeraceae (1.5%) in the Jun-B sample and unc. Verrucomicrobiaceae (1.1%) in the Oct-B sample (Fig. 5a).
A complex NMDS analysis (based on chemical data and bacterial OTUs) clearly separates Dec-B sample from all the others by the high abundance of genera Sphaerotilus and Hydrogenophaga. Higher temperature seems to move together with the higher ratio of Cyanobacteria (Fig. 6).
NMDS plot of bacterial communities from the biofilm samples of Danube based on Bray–Curtis distance of the bacterial OTUs (Only genera that occurred in at least 1% of at least one sample are represented in the figure) (Temp, temperature; Cond, electrical conductivity; TSS, total suspended solids; POM, particulate organic matter; PIM, particulate inorganic matter; Q, discharge; TCC, total cell counts; P, phosphorus; E. coli, E. coli count)
The most abundant archaeal phylum was Nitrososphaerota (previously Thaumarchaeota) in almost all samples (31.6–62.8%). Among the members of this phylum, Candidatus Nitrosoarchaeum (16.9–32.0%) and unc. Nitrososphaeraceae (3.2–22.0%) were the most frequent taxa in all biofilm samples. Representatives of the phylum Nanoarchaeota (mainly belonging to the class Woesearchaeia) also reached the high abundance in all samples (16.9–38.2%). Euryarchaeota appeared in 3.8–23.8% in the samples. From this phylum, the members of the genus Methanoregula were detected in each sample with a variable distribution. Its maximum was detected in the Apr-B sample (13.4%). The phylum Thermoproteota (previously Crenarchaeota) (especially class Bathyarchaeia) had a high relative abundance in the Dec-B and Apr-B samples (8.9 and 11.7%) (Fig. 5b).
Specific PCRs
Coliform bacteria were not detected in any biofilm sample using taxon-specific PCR. Pseudomonas spp. and Legionella spp. were present in several samples, but opportunistic pathogenic P. aeruginosa was detected only in the Dec-B sample, L. pneumophila in the Oct-B, Dec-B, and Apr-B samples. Stenotrophomonas maltophilia was present in all samples. Using multiplex PCR, ESBL genes and macrolide resistance genes were not detected in any of the biofilm samples (Table 4).
Discussion
Filamentous microorganisms dominate the biofilm throughout the year
Electron micrographs of the biofilms collected from the surface of the submerged stone at the sampling point downstream Budapest revealed different morphological features, taken at different times showing a multitude of filamentary formations (Fig. 2a–j). Chain-forming diatoms (e.g. Melosira varians, Fig. 2b, d, h) and gelatinous, long, stalk-forming diatoms (e.g. Gomphonema spp. Figure 2f) also contributed to the development of thick biofilms on the surface of the studied stone, as previously observed in Danube biofilm studies. During colonisation experiments on the artificial substrates at the main branch of the Danube at Göd (1669 riv. km), these diatoms played a major role in the formation of large “beard-like” biofilms (Ács et al. 2000; Makk et al. 2003b). Just as in those experiments, the higher magnification images showed that the surfaces of the diatoms are good colonisation sites for bacteria. Different morphologies (rod, coccus, filamentous, prosthaeca type) (e.g. Figure 2e, h) could be observed. Bacteria attached to the diatoms most probably not only consume the nutrients that drift to the surface, but the algae can also release organic substrates that are suitable nutrient sources for the attached bacteria (Makk et al. 2003a). Considering the results of amplicon sequencing, the abundance of long, thin filamentous structures in the Oct-B, Apr-B and Jun-B samples showed an enrichment of cyanobacteria, visible also in the microscopic images (Fig. 2b, h, j). In summer, during warm water periods, cyanobacteria were detected in high abundance (Ács and Kiss 1991; Ács et al. 2003) also by other authors. On the other hand, the Dec-B sample contained the most prokaryotic-size filaments: based on amplicon sequencing, these bacteria belonged to the genus (Fig. c, d) Sphaerotilus (with 60% relative abundance, Fig. 5a). Based on literature data, Sphaerotilus spp. are characterised by a long filamentous morphology, consisting of long chains of cells attached end-to-end (Spring 2006). These filaments often form dense, network like biofilms on surfaces that can be seen with the naked eye, just as in case of our December sample. In a previous biofilm study, changes in the bacterial community composition were most influenced by the temperature, water discharge element content and biological water quality of the Danube.
Increased water discharge promotes microbial diversity in the biofilm
At sampling times with higher water discharge in our case, the bacterial diversity of biofilms (Oct-B, Feb-B, Jun-B samples) was higher and the relative abundance of “rare” taxa (relative abundance is below 1%) was much higher. During periods of high waterflows, the Danube is rich in suspended, fine-grained sediment, it is rather turbid and carries more nutrients and trace elements. In the Danube, the heavy metal content (e.g. Pb, Cu, Zn, Cr, Mn, Fe and Ni) is related to the clay and silt fraction, and sediment metal content is thought to be controlled mainly by sorption–desorption processes associated with fine grain particles (Oreščanin et al. 2005). This may explain that at higher waterflows elevated concentrations of certain chemical elements could be clearly detected (Table 2).
Furthermore, the higher total suspended solids (TSS) values detected during such periods may explain the higher diversity of microbes in the biofilms as microbes may be transported associated with the suspended solids (SS). Our electron micrographs also support this finding; biofilm samples taken at higher water discharge periods were covered with a lot of tiny debris (Oct-B, Feb-B, Jun-B samples; Fig. 2a, b, f, g, i, j). The fraction of TSS in the water samples was mainly composed of particulate inorganic matter (PIM) (Table 2). These inorganic components may shrink less than the organic fractions at the drying procedure during the sample preparation for electron microscopy, so they are more observable on the images of the samples than their organic parts.
Influence of the local water pollution situation on biofilm diversity
The water quality of the Danube River has improved in the recent decades, most probably due to the installation of a number of wastewater treatment plants. This is reflected in the fact that no significant differences in measured chemical variables were detected between upstream and downstream sampling points at Budapest (Trábert et al. 2020). Nevertheless, contamination by faecal pollution from anthropogenic sources is still considered a critical problem throughout the Danube River Basin (Kirschner et al. 2009, 2017). This was observed in October 2020 when, according to the microbiological classification system (Kirschner et al. 2017), Colilert-18’s study calculations showed “critical” faecal pollution in the Danube water sample (E. coli 3500 MPN/100 ml, Table 1). The one-year study series of Schachner-Groehs et al. (2024) also showed the highest concentrations of all ARGs and human faecal pollution markers in the Danube water samples close to our sampling site in October 2020. A possible source of pollution in this stretch of the Danube could be the Rákos stream, which is located upstream of the sampling point. The water quality of the stream is strongly influenced by the sporadically occurring pollution of illegal point sources (local news). When the river was exposed to significant faecal loading, the bacterial community composition of the biofilms was also significantly affected. This was observed in the December sample: Sphaerotilus, a typical bacterium of high organic matter waters dominated our biofilm sample (Fig. 5a).
Despite the detection of varying amounts of E. coli bacteria in the water samples, none of the biofilm samples contained faecal indicator sequences by NGS or taxon-specific PCRs in our case. Very few sequences belonging to the Enterobacteriaceae family were identified by NGS with a relative abundance of < 1% (e.g. Citrobacter and Yersinia). However, the results for the filamentous Sphaerotilus bacteria were remarkable, as they were found to be unexpectedly abundant in the biofilm (Dec-B sample) after high levels of faecal pollution of the Danube water. Sphaerotilus spp. are known to be most frequently associated with “sewage fungus” (Curtis 1969). “Sewage fungus” is a “fungus-like” form in water, consisting mainly of a polymicrobial biofilm of filamentous bacteria. These are most likely appear where untreated or inadequately treated wastewater is discharged into the river (Curtis and Harrington 1971). “Sewage fungus” proliferates in saprobic rivers, and their presence is often a sign of severe pollution (Albini et al. 2023). In our case, the high abundance of Sphaerotilus in the biofilm samples (60% relative abundance) clearly indicated that the Danube might have been affected by some kind of faecal pollution. The detection of “sewage fungus” indicates longer-term changes in water quality according to microbiological-based classification, whereas the counts of the faecal indicator bacteria reflect an instantaneous status. Studies on river water quality have also proven that the algae living in biofilms are sensitive to the changing physical and chemical parameters of the environment (such as organic pollution, changes in salinity, and acidification), and due to their lifestyle, they are able to indicate environmental changes in the long term, in contrast to physical and chemical measurements (Makovinská et al. 2014; Trábert et al. 2020).
The bacterial phyla Pseudomonadota and Bacteroidota dominated the epilithic biofilms
Next-generation DNA sequencing allowed a detailed insight into the composition of the bacterial communities harboured at the sites studied. Pseudomonadota and Bacteroidota were the most common phyla in all the epilithic biofilms, similar to many previous studies of freshwater biofilm communities (Battin et al. 2016; Pineda-Mora et al. 2020; Engloner et al. 2023). The class Betaproteobacteria (e.g. Sphaerotilus, unc. Burkholderiaceae, Rhodoferax) were present in high quantities in all samples. These taxa are also known to occur with high frequency in nutrient-rich environments (Fierer et al. 2007). Despite the large input of allochthonous material and the variability of physicochemical parameters during the seasons, the composition of the dominant representatives of Betaproteobacteria (e.g. Rhodoferax, Hydrogenophaga, Sphaerotilus natans) in biofilm communities was found to be amazingly stable in previous studies of polluted rivers (Brümmer et al. 2003). Genomic analyses have shown that members of the Burkholderiaceae family harbour a variety of aromatic catabolic pathways, making them potential organisms for bioremediation of sites contaminated with aromatic pollutants (Pérez-Pantoja et al. 2012). Rhodoferax species have been isolated from different environments, e.g. sewage or an Antarctic microbial mat (Imhoff 2006). Recent research has identified Rhodoferax bacteria in “sewage fungus”. This genus was dominant, followed by Sphaerotilus, in airport de-icing events when the composition of “sewage fungus” was investigated by amplicon sequencing (Exton et al. 2023).
From the phylum Bacteroidota, the genus Flavobacterium (3.9–13% relative abundance) was present in high quantities in the samples. This taxon with characteristic surface-dependent gliding motility has been detected in previous studies with Danube gravel biofilm (Makk et al. 2003a, b). Their presence in biofilms is important due to their ability to degrade proteins and difficult-to-degrade polysaccharides such as cellulose and chitin from dead plant material (Kirchman 2002). Furthermore, Flavobacterium was found to be the dominant member of diatom-associated communities (Grossart et al. 2005). Phylogenetic analysis and co-culture studies indicated an adaptation of Bacteroidetes and Pseudomonadota to the microenvironment generated by the diatom biofilm (Bruckner et al. 2008). A mutualistic relationship between a Flavobacterium sp. and the diatom Navicula muralis has also been described (Jolley and Jones 1977).
The archaeal phyla Nitrososphaerota and Nanoarchaeota are also permanent members of the biofilm community
Representatives of ammonia-oxidising archaea such as Candidatus Nitrosoarchaeum and the family Nitrososphaeraceae belonging to the phylum Nitrososphaerota were found in high proportion in all biofilm samples. These taxa are ubiquitous microbes in marine, freshwater, and terrestrial environments, and their occurrence suggests that nitrification also plays an important role in primary production (Kim et al. 2011; Tourna et al. 2011) also in Danube biofilms. Despite their potential role in global carbon and nitrogen cycles, many questions about their physiology, metabolism and ecological niche remain unanswered due to the isolation difficulties.
The other dominant archaeal group in our biofilm samples belonged to Woesearchaeia (phylum Nanoarchaeota). They are widespread in different environments and enable carbon and hydrogen metabolism under anoxic conditions, which could be associated with symbiotic and/or fermentative lifestyles. This syntrophic and/or mutualistic partnership arose from deficiencies in metabolic pathways within the Woesearchaeia class. Authors suggested Woesearchaeia develop a partnership with certain bacterial methanogens and proposed a Woesearchaeia-methanogens consortium that likely influences methane formation (Liu et al. 2018). Methanogenic archaea were also detected in our samples (the third most abundant bacterial phyla were the Euryarchaeota, mainly methanogens with variable abundance, such as, e.g. members of the genus Methanoregula), so the appearance of this kind of association is assumed also in Danube biofilms.
Occurrence of facultative pathogens and antimicrobial resistance genes in the epilithic biofilm
No coliform bacteria were detected in our biofilm samples, but some water-related facultative pathogenic bacteria were found, such as Pseudomonas aeruginosa, Legionella pneumophila and Stenotrophomonas maltophilia. Sequences belonging to the latter genera were only detected in low quantities (Pseudomonas 1.01–0.07% and others with relative abundance < 1%).
The most widely used antibiotics in Hungary are beta-lactams and macrolides—therefore these antibiotics were tested in our assay (Antimicrobial consumption—Annual Epidemiological Report for 2021). Recently, the extended-spectrum beta-lactamase (ESBL) producing Gram-negative bacteria have become widespread in environmental samples, especially members of the family Enterobacteriaceae (Cho et al. 2023). ESBL-producing bacteria have been studied in the Austria, Germany in different studies, and a number of Danube countries during Joint Danube Survey 3 and 4 (Stoll et al. 2012; Grenni 2022; Kittinger et al. 2016b; Schachner-Groehs et al. 2024). In surface water samples taken over a total course of the Danube during JDS3, Escherichia coli, Klebsiella pneumoniae, Enterobacter cancerogenus, and Enterobacter cloacae were tested as positive for ESBL-harbouring genes (Kittinger et al. 2016b). The blaTEM which is one of the three target genes of ESBL-encoding genes was detected in more than half of the Danube water samples from upstream and downstream of Budapest (Phuong 2020). In a year-long temporal monitoring experiment, Schachner-Groehs et al. (2024) described that human faecal pollution from municipal wastewater discharges was the dominant factor shaping antibiotic resistance gene patterns in Danube water samples.
Macrolide resistance in human medicine usually develops during antibiotic therapy of bacteria that cause respiratory tract and soft tissue infections. Macrolide resistance (encoded by erm genes) can be easily transferred among bacteria as these genes are located on plasmids and transposons. In one study, macrolide resistance genes were studied in case of the Danube (Stoll et al. 2012). The water samples were collected from Rivers Rhine and Danube and their tributaries and were screened for the presence of ermA and ermB. The percentage of ermB positive samples from the Rhine River water was higher (69%) than from the Danube River (47%), while ermA was not detected in any of the samples. Based on Phoung’s study (2020), the ermB, msrA and mef, three out of the five target genes of macrolide resistance genes, were present in more than half of the Danube water samples.
However, in the current study, ESBL producing and macrolide resistance genes in our Danube biofilm samples could not be detected. A more detailed study of river biofilms (e.g. gravel coatings, sediments) would be important to understand the spread and dynamics of antibiotic resistance, which may act as long-term reservoirs (Schachner-Groehs et al. 2024).
Conclusion for future biology
The combination of morphological and taxonomic biofilm characterisation with environmental data allowed us to obtain deep insights on the influence of anthropogenic pollution and other environmental factors on the seasonal changes of epilithic microorganisms. Unexpected taxonomic bacterial diversity was revealed from amplicon sequencing data of epilithic biofilms formed on the same submerged stone in the Danube River. Many sequences showed low-level similarity to any known taxa. The morphology and microbial diversity of biofilm samples taken at different time points were influenced by changes in the physical, chemical and microbiological parameters of the water: higher temperature in the summer period resulted in the multiplication of Cyanobacteria. Despite the fact that the water quality of the Danube was classified as moderately polluted according to the microbiologically based classification system, in all our biofilm samples, we were able to detect a higher proportion of genera typical of organic-contaminated environments (e.g. Sphaerotilus, Rhodoferax, unc. Burkholderiaceae, Flavobacterium). After a critical level faecal pollution in the Danube, a less diverse biofilm (consisting mainly of Sphaerotilus bacteria) was detected, which can be defined as “sewage fungus”. Significant changes in the microbial composition of epilithic biofilms, such as high proliferation of Sphaerotilus, can be indicative of environmental changes over the longer term, in contrast to physical, chemical and microbiological parameters that reflect the current state of the water. To reveal the presence and spreading of antibiotic resistance in biofilms, a more detailed study would be essential.
References
Ács É, Kiss KT (1991) Investigation of periphytic algae in the Danube at Göd (1669 river km, Hungary). Archiv Für Hydrobiol Suppl Band Algol Stud 62:47–67
Ács É, Kiss KT, Szabó-Taylor K, Makk J (2000) Short-term colonization sequence of periphyton on glass slides in a large river (River Danube, near Budapest). Archiv Für Hydrobiol Suppl Band Algol Stud 100:135–156. https://doi.org/10.1127/algol_stud/100/2000/135
Ács É, Szabó K, Kiss KT, Hindák F (2003) Benthic algal investigations in the Danube river and some of its main tributaries from Germany to Hungary. Biol-Bratisl 58(4):545–554
Albini D, Lester L, Sanders P, Hughes JM, Jackson MC (2023) Early detection and environmental drivers of sewage fungus outbreaks in rivers. Ecol Solut Evid 4(3):e12277. https://doi.org/10.1002/2688-8319.12277
Alygizakis NA, Besselink H, Paulus GK, Oswald P, Hornstra LM et al (2019) Characterization of wastewater effluents in the Danube River Basin with chemical screening, in vitro bioassays and antibiotic resistant genes analysis. Environ Int 127:420–429. https://doi.org/10.1016/j.envint.2019.03.060
Antimicrobial consumption - Annual Epidemiological Report for (2021). https://www.ecdc.europa.eu/en/publications-data/downloadable-tables-antimicrobial-consumption-annual-epidemiological-report-2021
Banciu AR, Ionica DL, Vaideanu MA, Radulescu DM, Nita-Lazar M, Covaliu CI (2021) The occurrence of potentially pathogenic and antibiotic resistant Gram-negative bacteria isolated from the Danube Delta ecosystem. Sustainability 13(7):3955. https://doi.org/10.3390/su13073955
Battin TJ, Besemer K, Bengtsson MM, Romani AM, Packmann AI (2016) The ecology and biogeochemistry of stream biofilms. Nat Rev Microbiol 14(4):251–263. https://doi.org/10.1038/nrmicro.2016.15
Bej AK, Steffan RJ, DiCesare J, Haff L, Atlas RM (1990) Detection of coliform bacteria in water by polymerase chain reaction and gene probes. Appl Environ Microbiol 56(2):307–314. https://doi.org/10.1128/aem.56.2.307-314.1990
Bruckner CG, Bahulikar R, Rahalkar M, Schink B, Kroth PG (2008) Bacteria associated with benthic diatoms from Lake Constance: phylogeny and influences on diatom growth and secretion of extracellular polymeric substances. Appl Environ Microbiol 74(24):7740–7749. https://doi.org/10.1128/AEM.01399-08
Brümmer IHM, Felske A, Wagner-Dobler I (2003) Diversity and seasonal variability of β-Proteobacteria in biofilms of polluted rivers: analysis by temperature gradient gel electrophoresis and cloning. Appl Environ Microbiol 69(8):4463–4473. https://doi.org/10.1128/AEM.69.8.4463-4473.2003
Cho S, Jackson CR, Frye JG (2023) Freshwater environment as a reservoir of extended-spectrum β-lactamase-producing Enterobacteriaceae. J Appl Microbiol 134(3):lxad034-95. https://doi.org/10.1093/jambio/lxad034
Cloud JL, Carroll KC, Pixton P, Erali M, Hillyard DR (2000) Detection of Legionella species in respiratory specimens using PCR with sequencing confirmation. J Clin Microbiol 38(5):1709–1712. https://doi.org/10.1128/jcm.38.5.1709-1712.2000
Curtis EJC (1969) Sewage fungus: its nature and effects. Water Res 3(5):289–311. https://doi.org/10.1016/0043-1354(69)90084-0
Curtis EJC, Harrington DW (1971) The occurrence of sewage fungus in rivers in the United Kingdom. Water Res 5(6):281–290
da Silva Filho LV, Tateno AF, Velloso LDF, Levi JE, Fernandes S et al (2004) Identification of Pseudomonas aeruginosa, Burkholderia cepacia complex, and Stenotrophomonas maltophilia in respiratory samples from cystic fibrosis patients using multiplex PCR. Pediatr Pulmonol 37(6):537–547. https://doi.org/10.1002/ppul.20016
Eiler A, Farnleitner AH, Zechmeister TC, Herzig A, Hurban C et al (2003) Factors controlling extremely productive heterotrophic bacterial communities in shallow soda pools. Microb Ecol 46(1):43–54. https://doi.org/10.1007/s00248-002-2041-9
Engloner AI, Vargha M, Kós P, Borsodi AK (2023) Planktonic and epilithic prokaryota community compositions in a large temperate river reflect climate change related seasonal shifts. PLoS ONE 18(9):e0292057. https://doi.org/10.1371/journal.pone.0292057
Exton B, Hassard F, Vaya AM, Grabowski RC (2023) Polybacterial shift in benthic river biofilms attributed to organic pollution–a prospect of a new biosentinel? Hydrol Res 54(3):348–359. https://doi.org/10.2166/nh.2023.114
Ferri M, Ranucci E, Romagnoli P, Giaccone V (2017) Antimicrobial resistance: a global emerging threat to public health systems. Crit Rev Food Sci Nutr 57(13):2857–2876. https://doi.org/10.1080/10408398.2015.1077192
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88(6):1354–1364. https://doi.org/10.1890/05-1839
Fiume L, Sabattini BMA, Poda G (2005) Detection of Legionella pneumophila in water samples by species-specific real-time and nested PCR assays. Lett Appl Microbiol 41(6):470–475. https://doi.org/10.1111/j.1472-765X.2005.01779.x
Gray ND, Miskin IP, Kornilova O, Curtis TP, Head IM (2002) Occurrence and activity of Archaea in aerated activated sludge wastewater treatment plants. Environ Microbiol 4(3):158–168. https://doi.org/10.1046/j.1462-2920.2002.00280.x
Grenni P (2022) Antimicrobial resistance in rivers: a review of the genes detected and new challenges. Environ Toxicol Chem 41(3):687–714. https://doi.org/10.1002/etc.5289
Grossart HP, Levold F, Allgaier M, Simon M, Brinkhoff T (2005) Marine diatom species harbour distinct bacterial communities. Environ Microbiol 7(6):860–873. https://doi.org/10.1111/j.1462-2920.2005.00759.x
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:1–9
Herlemann DP, Labrenz M, Jürgens K, Bertilsson S, Waniek JJ, Andersson AF (2011) Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J 5(10):1571–1579. https://doi.org/10.1038/ismej.2011.41
Imhoff JF (2006) The phototrophic beta-Proteobacteria. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes, 5th edn. Springer, New York, pp 593–601. https://doi.org/10.1007/0-387-30745-1_25
International Organization for Standardization (2012) ISO 9308–2:2012 Water Quality – Enumeration of Escherichia coli and Coliform Bacteria - Part 2: Most Probable Number Method, p 45
ISO 19458:2006 Water quality. Sampling for microbiological analysis.
Jolley ET, Jones AK (1977) The interaction between Navicula muralis Grunow and an associated species of Flavobacterium. Brit Phycol J 12(4):315–328. https://doi.org/10.1080/00071617700650341
Kim BK, Jung MY, Yu DS, Park SJ, Oh TK et al (2011) Genome sequence of an ammonia-oxidizing soil archaeon, “Candidatus Nitrosoarchaeum koreensis” MY1. J Bacteriol 193(19):5539–5540. https://doi.org/10.1128/JB.05717-11
Kirchman DL (2002) The ecology of Cytophaga-Flavobacteria in aquatic environments. FEMS Microbiol Ecol 39(2):91–100. https://doi.org/10.1111/j.1574-6941.2002.tb00910.x
Kirschner AKT, Kavka GG, Velimirov B, Mach RL, Sommer R, Farnleitner AH (2009) Microbiological water quality along the Danube River: integrating data from two whole-river surveys and a transnational monitoring network. Water Res 43(15):3673–3684. https://doi.org/10.1016/j.watres.2009.05.034
Kirschner AKT, Reischer GH, Jakwerth S, Savio D, Ixenmaier S, Tóth E et al (2017) Multiparametric monitoring of microbial faecal pollution reveals the dominance of human contamination along the whole Danube River. Water Res 124:543–555. https://doi.org/10.1016/j.watres.2017.07.052
Kittinger C, Lipp M, Baumert R, Folli B, Koraimann G et al (2016) Antibiotic resistance patterns of Pseudomonas spp. isolated from the River Danube. Front Microbiol 7:586. https://doi.org/10.3389/fmicb.2016.00586
Kittinger C, Lipp M, Folli B, Kirschner A, Baumert R et al (2016b) Enterobacteriaceae isolated from the River Danube: antibiotic resistances, with a focus on the presence of ESBL and carbapenemases. PLoS ONE 11(11):e0165820. https://doi.org/10.1371/journal.pone.0165820
Kittinger C, Kirschner A, Lipp M, Baumert R, Mascher F et al (2018) Antibiotic resistance of Acinetobacter spp. isolates from the River Danube: susceptibility stays high. Int J Environ Res Public Health 15(1):52. https://doi.org/10.3390/ijerph15010052
Kneis D, Berendonk TU, Forslund SK, Hess S (2022) Antibiotic resistance genes in river biofilms: a metagenomic approach toward the identification of sources and candidate hosts. Environ Sci Technol 56(21):14913–14922. https://doi.org/10.1021/acs.est.2c00370
Lange-Enyedi NT, Borsodi AK, Németh P, Czuppon G, Kovács I et al (2023) Habitat-related variability in the morphological and taxonomic diversity of microbial communities in two Hungarian epigenic karst caves. FEMS Microbiol Ecol 99(12):fiad161. https://doi.org/10.1093/femsec/fiad161
Lavenir R, Jocktane D, Laurent F, Nazaret S, Cournoyer B (2007) Improved reliability of Pseudomonas aeruginosa PCR detection by the use of the species-specific ecfX gene target. J Microbiol Methods 70(1):20–29. https://doi.org/10.1016/j.mimet.2007.03.008
Lin Q, Sekar R, Marrs R, Zhang Y (2019) Effect of river ecological restoration on biofilm microbial community composition. Water 11:1244. https://doi.org/10.3390/w11061244
Liu X, Li M, Castelle CJ, Probst AJ, Zhou Z et al (2018) Insights into the ecology, evolution, and metabolism of the widespread Woesearchaeotal lineages. Microbiome 6:1–16. https://doi.org/10.1186/s40168-018-0488-2
Liu YJ, Li ZH, He YT, Yuan L, Sheng GP (2023) Antibiotic resistomes in face-mask biofilm along an urban river: Multiple drivers and co-occurrence with human opportunistic pathogens. J Hazard Mater 455:131587. https://doi.org/10.1016/j.jhazmat.2023.131587
Madsen JS, Burmølle M, Hansen LH, Sørensen SJ (2012) The interconnection between biofilm formation and horizontal gene transfer. FEMS Immunol Med Microbiol 65(2):183–195. https://doi.org/10.1111/j.1574-695X.2012.00960.x
Makk J, Ács E, Márialigeti K, Kovács G (2003a) Investigations on the Danube gravel-biofilm diatom-associated bacterial communities. Biol-Bratisl 58(4):729–742
Makk J, Beszteri B, Ács É, Márialigeti K, Szabó K (2003b) Investigations on diatom-associated bacterial communities colonizing an artificial substratum in the River Danube. Archiv Für Hydrobiol Suppl Large Rivers 14:249–265
Makk J, Ács É (1997) Investigation of epilithic biofilms in the River Danube. 32. Arbeitstagung der IAD, Wien, pp 199 202
Makovinská J, Hlúbiková D, Fidlerová D (2014) Joint Danube Survey 3 - Phytobenthos; International Commission for the Protection of the Danube River: Vienna, Austria https://www.danubesurvey.org/jds3/jds3-files/nodes/documents/phytobenthos.pdf
Mao G, Liang J, Wang Q, Zhao C, Bai Y et al (2021) Epilithic biofilm as a reservoir for functional virulence factors in wastewater-dominant rivers after WWTP upgrade. J Environ Sci 101:27–35. https://doi.org/10.1016/j.jes.2020.05.014
Márialigeti K, Zabnum A, Abdullach M (1997) The composition of the Danube rolling gravel biofilm bacterial communitias. In: Proceedings of the international regional seminar environment protection: modem studies in ecology and microbiology, Uzgorod, pp 158–162
Marutescu LG, Popa M, Gheorghe-Barbu I, Barbu IC, Rodríguez-Molina D et al (2023) Wastewater treatment plants, an “escape gate” for ESCAPE pathogens. Front Microbiol 14:1193907. https://doi.org/10.3389/fmicb.2023.1193907
Michaelis C, Grohmann E (2023) Horizontal gene transfer of antibiotic resistance genes in biofilms. Antibiotics 12(2):328. https://doi.org/10.3390/antibiotics12020328
Oksanen JF, Blanchet G, Friendly M, Kindt R, Legendre P et al (2020) vegan: Community Ecology Package. R package version 2.5–7. https://cran.r-project.org/package=vegan
Oreščanin V, Lulić S, Medunić G, Mikelić L (2005) Granulometric and chemical composition of the Danube River sediments, Batina Village. Croatia Geologia Croatica 58(2):185–194. https://doi.org/10.4154/GC.2005.10
Ovreås L, Forney F L, Daae V, Torsvik (1997) Distribution of bacterioplankton in meromictic Lake Saelenvannet as determined by denaturing gradient gel electrophoresis of PCR-amplified gene fragments coding for 16S rRNA Applied and Environmental Microbiology 63(9): 3367–3373 https://doi.org/10.1128/aem.63.9.3367-3373.1997
Pérez-Pantoja D, Donoso R, Agulló L, Córdova M, Seeger M et al (2012) Genomic analysis of the potential for aromatic compounds biodegradation in Burkholderiales. Environ Microbiol 14(5):1091–1117. https://doi.org/10.1111/j.1462-2920.2011.02613.x
Phuong DT (2020) Detection of ESBL-producing and macrolide resistance genes in River Danube and water supply systems in Budapest. MSc thesis, Eötvös Loránd University, Department of Microbiology, Budapest, Hungary
Pin L, Eiler A, Fazi S, Friberg N (2021) Two different approaches of microbial community structure characterization in riverine epilithic biofilms under multiple stressors conditions: Developing molecular indicators. Mol Ecol Resour 21(4):1200–1215. https://doi.org/10.1111/1755-0998.13341
Pineda-Mora D, Juárez-López AL, Toribio-Jiménez J, Leal-Ascencio MT, Ruvalcaba-Ledezma JC et al (2020) Diversity and functions of epilithic riverine biofilms. Water Air Soil Pollution 231:1–19. https://doi.org/10.1007/s11270-020-04692-x
R Core Team (2018) R: a language and environment for statistical computing. https://www.r-project.org/
Sabri N, Schmitt H, Van der Zaan B, Gerritsen HW, Zuidema T et al (2020) Prevalence of antibiotics and antibiotic resistance genes in a wastewater effluent-receiving river in the Netherlands. J Environ Chem Eng 8(1):102245. https://doi.org/10.1016/j.jece.2018.03.004
Salem AMA (1997) N-cycle associated facultative and obligate chemolithotrophic bacteria in Danube water and sediment. PhD Dissertation, Eötvös Loránd University, Department of Microbiology, Budapest, Hungary
Schachner-Groehs I, Koller M, Leopold M, Kolm C, Linke RB et al (2024) Linking antibiotic resistance gene patterns with advanced faecal pollution assessment and environmental key parameters along 2300 km of the Danube River. Water Res 252:121244. https://doi.org/10.1016/j.watres.2024.121244
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75(23):7537–7541. https://doi.org/10.1128/AEM.01541-09
Sharma P (2022) Role and significance of biofilm-forming microbes in phytoremediation-a review. Environ Technol Innov 25:102182. https://doi.org/10.1016/j.eti.2021.102182
Spilker T, Coenye T, Vandamme P, LiPuma JJ (2004) PCR-based assay for differentiation of Pseudomonas aeruginosa from other Pseudomonas species recovered from cystic fibrosis patients. J Clin Microbiol 42(5):2074–2079. https://doi.org/10.1128/JCM.42.5.2074-2079.2004
Spring S (2006) The genera Leptothrix and Sphaerotilus. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The Prokaryotes: a Handbook on the Biology of Bacteria, New York, Springer, 3rd edn. 5: 758–777
Stoll C, Sidhu JP, Tiehm A, Toze S (2012) Prevalence of clinically relevant antibiotic resistance genes in surface water samples collected from Germany and Australia. Environ Sci Technol 46(17):9716–9726. https://doi.org/10.1021/es302020s
Tindall BJ, Rosselló-Móra R, Busse HJ, Ludwig W, Kämpfer P (2010) Notes on the characterization of prokaryote strains for taxonomic purposes. Int J Syst Evol Microbiol 60(1):249–266. https://doi.org/10.1099/ijs.0.016949-0
Tourna M, Stieglmeier M, Spang A, Könneke M, Schintlmeister A et al (2011) Nitrososphaera viennensis, an ammonia oxidizing archaeon from soil. Proc Natl Acad Sci 108(20):8420–8425. https://doi.org/10.1073/pnas.1013488108
Trábert Z, Duleba M, Bíró T, Dobosy P, Földi A et al (2020) Effect of land use on the benthic diatom community of the Danube River in the Region of Budapest. Water 12(2):479. https://doi.org/10.3390/w12020479
Trung NT, Hien TTT, Huyen TTT, Quyen DT, Binh MT et al (2015) Simple multiplex PCR assays to detect common pathogens and associated genes encoding for acquired extended spectrum betalactamases (ESBL) or carbapenemases from surgical site specimens in Vietnam. Ann Clin Microbiol Antimicrob 14(1):1–7. https://doi.org/10.1186/s12941-015-0079-z
Van Driezum IH, Chik AHS, Jakwerth S, Lindner G, Farnleitner AH et al (2018) Spatiotemporal analysis of bacterial biomass and activity to understand surface and groundwater interactions in a highly dynamic riverbank filtration system. Sci Total Environ 627:450–461. https://doi.org/10.1016/j.scitotenv.2018.01.226
Vargha M, Takáts Z, Márialigeti K (2005) Degradation of atrazine in a laboratory scale model system with Danube river sediment. Water Res 39(8):1560–1568. https://doi.org/10.1016/j.watres.2004.10.013
Velimirov B, Milosevic N, Kavka GG, Farnleitner AH, Kirschner AKT (2011) Development of the bacterial compartment along the Danube River: a continuum despite local influences. Microb Ecol 61(4):955–967. https://doi.org/10.1007/s00248-010-9768-5
Zalmum AA (1997) Microbiological bank-wall filtered well water quality as a function of Danube rolling gravel bed biofilm bacterial species composition. PhD Dissertation, Eötvös Loránd University, Department of Microbiology, Budapest, Hungary
Zancarini A, Echenique-Subiabre I, Debroas D, Taïb N, Quiblier C et al (2017) Deciphering biodiversity and interactions between bacteria and microeukaryotes within epilithic biofilms from the Loue River. Fr Sci Rep 7(1):4344. https://doi.org/10.1038/s41598-017-04016-w
Zmantar T, Kouidhi B, Miladi H, Bakhrouf A (2011) Detection of macrolide and disinfectant resistance genes in clinical Staphylococcus aureus and coagulase-negative staphylococci. BMC Res Notes 4:1–9. https://doi.org/10.1186/1756-0500-4-453
Acknowledgements
This work was supported by ELTE Thematic Excellence Programme 2020 Supported by National Research—Development and Innovation Office—TKP2020-IKA-05. The study was supported also by the Scientific Foundations of Education Research Program of the Hungarian Academy of Sciences”. Additional support was provided by the Austrian Science Fund (FWF) project P32464-B. We thank Dr. Wolfgang Kandler (Department of Agrobiotechnology, IFA-Tulln, Institute of Bioanalytics and AgroMetabolomics, Tulln, Austria) for determining the chemical elements.
Funding
Open access funding provided by Eötvös Loránd University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Makk, J., Toumi, M., Krett, G. et al. Temporal changes in the morphological and microbial diversity of biofilms on the surface of a submerged stone in the Danube River. BIOLOGIA FUTURA (2024). https://doi.org/10.1007/s42977-024-00228-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42977-024-00228-0