Abstract
Understanding why sexual reproduction—which involves syngamy (union of gametes) and meiosis—emerged and how it has subsisted for millions of years remains a fundamental problem in biology. Considered as the essence of sex, meiotic recombination is initiated by a DNA double-strand break (DSB) that forms on one of the pairing homologous chromosomes. This DNA lesion is subsequently repaired by gene conversion, the non-reciprocal transfer of genetic information from the intact homolog. A major issue is which of the pairing homologs undergoes DSB formation. Accumulating evidence shows that chromosomal sites where the pairing homologs locally differ in size, i.e., are heterozygous for an insertion or deletion, often display disparity in gene conversion. Biased conversion tends to duplicate insertions and lose deletions. This suggests that DSB is preferentially formed on the “shorter” homologous region, which thereby acts as the recipient for DNA transfer. Thus, sex primarily functions as a genome (re)loading mechanism. It ensures the restoration of formerly lost DNA sequences (deletions) and allows the efficient copying and, mainly in eukaryotes, subsequent spreading of newly emerged sequences (insertions) arising initially in an individual genome, even if they confer no advantage to the host. In this way, sex simultaneously repairs deletions and increases genetic variability underlying adaptation. The model explains a remarkable increase in DNA content during the evolution of eukaryotic genomes.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Mendel’s second law, the principle of segregation, states that when a diploid cell undergoes meiosis, one half of the genes in the four haploid products (gametes) should be maternal while the other half paternal (Morgan 1919). If this were exactly the case, one would expect that the proportion of DNA sequences with no contribution to the fitness of the organism should be nearly constant in the subsequent generations, that is, only a few individuals of the population at each generation contained such a sequence—according to the Hardy–Weinberg principle (Stern 1943). Thus, non-functional sequences such as microsatellites (short tandem DNA repeats), non-transcribed long inverted repeats, or inactive mobile genetic elements should only be sporadically present among the individuals of an eukaryotic species (note that each of these sequences primarily arose in an individual genome). Contrary to this expectation, considerable parts of eukaryotic genomes appear to have no adaptive role (Dawkins 1976; Doolittle and Sapienza 1980; Venter et al. 2001; Nobrega et al. 2004; Ponting and Hardison 2011; Doolittle 2013), and these “wicked” or even harmful sequences are located at identical genomic loci in almost all members of a given species. Uncovering the molecular mechanisms by which newly emerged sequences providing no advantage to the host can spread within a species is of major interest to science (Lynch 2007). Recent findings concerning the mechanisms of meiotic homologous recombination (HR) may help understand why eukaryotic genomes tend to accumulate non-functional sequences, with important implications for the evolution of sex, which still represents a central, long-standing issue in biology (Maynard-Smith 1978; Kondrashov 1993; Barton and Charlesworth 1998; Partridge and Hurst 1998; Butlin 2002; Otto and Lenormand 2002; Webster and Hurst 2012).
Sex includes syngamy, the formation of a diploid zygote by the union of a haploid maternal gamete and a haploid paternal gamete, and meiosis, which enables the exchange of genetic information between homologous chromosomes (the process is called homologous recombination; HR) and generates haploid gametes. Meiosis has two major effects: the segregation of alleles at each locus and recombination of alleles at several loci. Most evolutionary models have assumed that HR is the evolutionary value of sex and that HR is driven by the reciprocal exchange of allelic sequences between the two pairing homologous chromosomes, the process called crossing over (reviewed in Ref. Kondrashov 1993). However, meiotic HR is initiated by a DNA double-strand break (DSB) that forms on one of the pairing homologs, and this endogenously (enzymatically) generated DNA “lesion” is primarily repaired via gene conversion, which mediates the non-reciprocal copying of genetic information from the intact homolog to the broken one (unidirectional gene exchange). Gene conversion is often but not always accompanied by the reciprocal exchange of flanking homologous sequences (crossing over). Thus, gene conversion, but not crossing over, is indispensable for HR. It is intriguing that the effect of gene conversion has rarely been taken into account in evolutionary models for the origin and short-term maintenance of sexual reproduction.
Models provided to date for the origin of sex can be divided into two major classes: immediate benefit hypotheses and variation/selection hypotheses (Kondrashov 1993). According to the former models, sex is advantageous regardless of reciprocal gene exchange, while the latter models assume that sex allows reciprocal gene exchange (crossing over) promoting genetic diversity and response to selection among the progeny. Briefly, immediate benefit hypotheses have proposed that sex is advantageous as (1) it increases the fitness of progeny (Dougherty 1955; Lloyd 1980; Bernstein and Bernstein 1991), (2) reduces the deleterious mutation rate (Bengtsson 1986; Ettinger 1986; Holliday 1988), or 3) increases the efficiency of selection (Geodakyan 1965; Trivers and Hare 1976). Variation and selection hypotheses include the (1) environmental stochastic (Morgan 1913; Fisher 1930; Muller 1932; Manning and Jenkins 1980; Manning 1982), (2) environmental deterministic (Sturtevant and Mather 1938; Mather 1943; Eshel and Feldman 1970; Treisman 1976; Charlesworth 1976), (3) mutational stochastic (Muller 1964), (4) drift (e.g., by generating negative linkage disequilibrium—Otto and Lenormand 2002), and (5) mutational deterministic models (Kondrashov 1982; Crow and Simmons 1983). In general, the mutational deterministic hypothesis is actually the most preferred model for the origin of sex (Szathmáry 2015) and presumes an increased selection efficiency against synergistically interacting deleterious mutations (mutations are purged from populations through the loss of individuals in which they accumulate by sex). To summarize these models, sex provides the advantage of preventing the accumulation of deleterious mutations or/and generating novel (adaptive) gene combinations. These advantages rely on long-acting group selection among populations to maintain sex. However, one can question the short-term advantage of sex to individuals which is necessary if sex is to be maintained long enough for such group selection to operate (Prahlad et al. 2003). To address this subject, it is worth considering what is known about the mechanisms of meiotic HR.
The mechanism of homologous recombination: an essential role for gene conversion
The current consensus model of meiotic recombination is a composite of the DSBR (DNA double-strand break repair) and SDSA (synthesis-dependent strand annealing) mechanisms (Fig. 1) (Sostak et al. 1983; Kusano et al. 1994; Paques and Haber 1999; Zickler and Kleckner 1999; Petes 2001; de Massy 2003; Haber et al. 2004; Chen et al. 2007; Serrentino and Borde 2012). The process begins with a homology search between the homologous non-sister chromatids, after the chromosomes become duplicated in the S phase of the cell cycle. Homologous pairing of DNA strands during the pre-leptotene phase precedes DSB formation that actually induces recombination (Danilowicz et al. 2009; Gladyshev and Kleckner 2014). Homology recognition is based on electrostatic interactions between the two homologous DNA helical structures, which have complementary charge distribution, and can occur at distances of 2 nm (Kornyshev and Leikin 2001). Similarities between the structures of 30 nm chromatin fibers could account for distant recognition of homology. As meiotic prophase I progresses, the assembly of the synaptonemal complex commences in the zygotene stage and becomes completed by the pachytene stage (Zickler and Kleckner 1999). In the yeast Saccharomyces cerevisiae, the chromosomal axis proteins Red1, Hop1, and Rec8 are necessary for both homologous pairing of non-sister chromatids and DSB formation (Zickler and Kleckner 2015).
Mechanisms of meiotic recombination. The model combines the DNA double-strand break (DSBR) repair and synthesis-dependent strand annealing (SDSA) pathways. Proteins involved in this process are indicated as colored globes. For details, see the text. Briefly, the pairing of homologous chromosomal regions precedes the recombination-inducing DSB formation that occurs on one of the homologs. DSB is formed endogenously by Spo11. The local chromatin structure largely affects the site where DSB forms (Spo11 interacts simultaneously with the two DNA duplexes). After nucleolytic processing of the free DNA ends, DSB is essentially repaired by gene conversion, the non-reciprocal transfer of genetic information from the intact DNA helix. Thus, the homolog on which DSB is formed becomes the recipient of gene conversion-meditated sequence copying. After mismatch or large loop repair, gene conversion is infrequently associated with the reciprocal exchange of flanking sequences (crossing over). Thus, gene conversion serves as an essential process of homologous recombination to generate recombinant gametes. H3K4me3: histone 3 lysine 4 trimethylation; γ-H2AX: H2AX histone phosphorylation; dHJ: double Holliday junction; CO: crossing over
The DSB is formed on one of the pairing homologs, and this site is generally trimethylated at the lysine-4 residue on histone 3 (H3K4me3) protein (note that in yeast DSB formation generally occurs at nucleosome depleted regions) (Fig. 1) (Borde et al. 2009). While the widespread mark of DSB formation in eukaryotes is H3K4 di- or trimethylation, in the nematode Caenorhabditis elegans DSB formation correlates with H2AK5ac (histone H2A lysine 5 acetylation) (Wagner et al. 2010). The accessibility of chromatin structure and nucleosome positions thus greatly affects where DSB formation occurs (Pan et al. 2011; Brachet et al. 2012). The distribution of DSBs is controlled by a multi-level regulatory system. In S. cerevisiae, the Set1 catalytic subunit of the Complex Proteins Associated with Set1 (COMPASS) histone methylase complex carries out all H3K4 di- and trimethylation (Fig. 1) (Acquaviva et al. 2013). In mammals, Prdm9, a meiosis-specific histone trimethyltransferase, is responsible for H3K4 trimethylation (Baudat et al. 2010). While H3K4me3 is not the sole determinant of the site where the DSB forms, it promotes DSB formation through interactions with Spp1 (Acquaviva et al. 2013). Actually, the PHD (Plant Homeo Domain) finger domain of Spp1 binds to H3K4me3 sites and then anchors hot spots to DSB proteins (Mer2). The PHD finger-containing protein Spp1, another component of the COMPASS complex, connects histone modification to DSB formation by binding to H3K4me3 sites and interacting with Mer2, a part of the recombination initiation complex. Spp1-Mer2 interaction may allow the tethering of the H3K4me3 rich region to the chromosomal axis and help DNA cleavage at a nearby nucleosome-depleted region (Panizza et al. 2011).
In eukaryotes, DSBs are generated by the highly conserved topoisomerase II-like transesterase Spo11 in the leptotene phase (prophase I) (Atcheson et al. 1987). Spo11 shows clear homology with the subunit A of the Archaeal Type-IIB topoisomerase TopVI (Bergerat et al. 1997). The catalytic residue on Spo11p subunit, Tyr135, attacks the DNA backbone to form a link between the enzyme and the DNA strand, thereby cleaving it via a transesterification reaction. Protein–protein interaction studies have shown that Spo11 is part of a recombination initiation complex, which consists of the Mer2/Rec107-Mei4-Rec114, Ski8/Rec103-Spo11, Rec102-Rec104, and MRX/MRN (Mre11-Rad50-Xrs2/NBS1) subcomplexes (Fig. 1). Mer2 acts as a scaffold and regulator of Spo11, while Mei4 controls the place of DSB formation (Kumar et al. 2010). Ski8 stabilizes association of Spo11 with the chromosomal axis, while Rec102 and Rec104 aid the Spo11-catalyzed DNA cleavage. The presence of MRX/MRN subcomplex is necessary for both DSB formation and subsequent gene conversion-mediated repair. The basic mechanism and regulation of DSB formation are evolutionarily conserved (Keeney et al. 2014).
Mer2 is phosphorylated by the conserved CDK-S (cyclin-dependent kinase Cdc28) and DDK (the Dbf4-dependent Cdc7 kinase) proteins (Benjamin et al. 2003). This mechanism ensures that DSB formation occurs after DNA replication only (Murakami and Keeney 2014). The phosphorylation of Mer2 is necessary for DSB formation, because only phosphorylated Mer2 is able to recruit the recombination initiation complex. Therefore, CDK-S and DDK link the regulation of DSB formation to cell cycle progression.
DSB repair is initiated by H2AX histone phosphorylation at Ser139 (γ-H2AX) by Tel1/ATM and Mec1/ATR, two highly conserved DNA damage serine/threonine protein kinases (Grabarz et al. 2012). Tel1 and Mec1 regulate the number of DSBs on a chromosome via a negative feedback mechanism (Lange et al. 2011). The ATP-dependent chromatin remodeler Fun30 is also directly involved in DSB response. It counteracts the inhibitory effect of Rad9 (a DNA damage-dependent checkpoint protein) on DNA resection (Chen et al. 2012). These chromatin modifications weaken histone-DNA interactions in nearby nucleosomes, which promote 5’ DNA-end resection, by helping the resection machinery to access the DNA strand.
Meiotic DSB formation is catalyzed by a complex involving SPO11 and TOPOVIBL (Robert et al. 2016). Contrary to earlier views, Spo11 proteins remain attached to 5’ DNA ends after the DSB is formed (Keeney and Kleckner 1995). The endo-exonuclease Mre11 and the endonuclease Sae2/CtIP have to nick the dsDNA to be resected up to 300 nt from the 5’ ends of the break. Mre11 facilitates resection in 3’ to 5’ direction (toward the break), while Exo1 or the Sgs1 (RecQ helicase)-Dna2 nuclease complex acts in 5’ to 3’ direction (away from the break). Thus, resection is bidirectional. As a result of 5’-end resection, two 3’ single strands form on opposite sides of the break. Then, RPA (Replication Protein A) is loaded on both sides of the DSB to the two 3’ DNA ends (Fig. 1) (Huertas 2010).
Dmc1, a homolog of the RecA bacterial/RadA archaeal strand exchange protein, forms a filament on one of the 3′ ssDNA tails (Neale and Keeney 2006). This strand then invades the opposing homologous DNA duplex to form a displacement loop (D-loop) during a process called strand invasion. Dmc1 interacts with Rad51, Mei5, Sae3, Rad52, Rad54, Rad55, Rad57, BRCA1, and BRCA2 proteins and with the Hop2-Mnd1 complex in aiding Dmc1-mediated strand exchange (Neale and Keeney 2006; Cloud et al. 2012). If mismatches are formed during strand exchange, they are corrected later by repair mechanisms (Jiricny 2006). After strand invasion, the D-loop is lengthened during branch migration by DNA synthesis (Chen et al. 2007).
Branch migration without the displacement of the newly synthesized strand from its template leads to double Holliday junction formation (dHJ). dHJ formation can lead to crossover and non-crossover events (Serrentino and Borde 2012). In the DSBR pathway, dHJ can be either resolved or dissolved. dHJs can be resolved by the eukaryotic HJ resolvase Yen1/Gen1, the Slx1-Slx4 complex, or the Mus81-Mms4 complex (Youds and Boulton 2011). dHJ resolution is promoted by the ZMM proteins, which leads to both crossover and non-crossover products. dHJ dissolution is promoted by the STR (Sgs1 (RecQ) helicase-TopIII topoisomerase-RIM1-RIM2) complex, and it leads to only non-crossover products (Fig. 1).
The displacement of the newly synthesized strand during D-loop extension and then its reconnection to its partner strand lead to the SDSA pathway (McMahill et al. 2007). The pathway is promoted by the Srs2 DNA helicase and the Sgs1/BLM RecQ helicase, and it produces non-crossover events.
The highly conserved mismatch repair (MMR) system recognizes mismatches in the resulting recombination intermediates (heteroduplexes) and then repairs them (Jiricny 2006).
Repair of loops larger than 16 nucleotides (nt) is facilitated largely independently of the MMR system by the large loop repair (LLR) system (Jensen et al. 2005). LLR in yeast requires DNA Polδ, the RFC (replication factor C) protein complex, PCNA (proliferating cell nuclear antigen), and FEN1/Rad27, which can repair mismatches up to about 5.6 kb, although repair efficiency decreases as the loop gets larger (Corrette-Bennett et al. 2001).
Depending on which of the pairing homologous regions (the shorter or the longer) acts as a template for DNA transfer, MMR and LLR can restore the original genetic information (a deletional allele is restored to the wild-type allele) or duplicate the donor sequence (the wild-type allele is transformed to an insertional allele) (Kirkpatrick 1999). This non-reciprocal sequence copying by gene conversion is largely the result of LLR and MMR of recombination intermediates that are formed at the end of the DSBR and SDSA pathways (Fig. 1). The vast majority of recombination events result in gene conversion without crossing over.
Below, we summarize those molecular events of meiotic HR that appear particularly important in understanding the evolutionary function of sex:
-
homology search between the homologs (DNA pairing) precedes DSB formation; HR is initiated by the formation of a DSB on one of the pairing homologous chromosomes
-
the opposite (allelic) homologous site remains intact throughout the recombination process
-
chromatin structure greatly affects the site where the DSB forms (a local chromatin opening allows recombination proteins to access the DNA strand): the accessibility of chromatin structure and nucleosome positions largely influence where DSB formation happens
-
DSB is endogenously generated by Spo11
-
Spo11 simultaneously interacts with both DNA helixes (homologous regions)
-
sequences at allelic position greatly influence DSB formation
-
DSB is repaired by gene conversion, the non-reciprocal transfer of genetic information from the intact homologue
-
gene conversion is indispensable for generating recombinant chromosomes
-
gene conversion is not always associated with the reciprocal exchange of the flanking sequences (crossing over)
-
gene conversion promotes the transmission of alleles even without conferring advance; thus, HR may be induced by self-promoting genetic elements (“selfish genes”)
Disparity in gene conversion
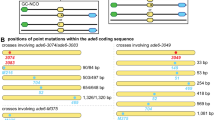
Meiotic HR is initiated by the formation of a DSB on one of the pairing homologs. DSB is then repaired by unidirectional sequence copying from the intact homolog (gene conversion). An important aspect of the process that requires further clarification is the homolog on which DSB is formed, in other words which of the pairing homologous chromosomal regions becomes the recipient of DNA transfer during gene conversion (Garcia et al. 2015). Experimental data show that in yeast heterozygous insertions often display disparity in gene conversion that duplicates, rather than eliminates, the insertion (Kearney et al. 2001). The segregation pattern of spore colonies is 6:2 when conversion transmits the wild-type allele, whereas copying of the mutant allele (insertion) results in a 2:6 gene conversion tetrad (note that without recombination the spore segregation pattern is 4:4). In meiotic recombination including heterozygous insertions, 2:6 conversions are more common than 6:2 segregations (Fig. 2) (Kearney et al. 2001; Johnson-Schlitz and Engels 1993). Kearney et al. observed (2001) that heterozygous very large (up to 5.6 kb) insertions show 11-fold 2:6 conversion bias (toward duplication of the insertion). This conversion bias results from the preference at which DSB forms on one homolog. Biased conversion has been accounted to be primarily for responsible for spreading novel DNA sequences in mammals (Chen et al. 2006). For example, conversion bias is widely implicated in the evolution of human microsatellites (Xu et al. 2000).
DSB formation preferentially occurs on the “shorter” homologous region. Left column: heterozygous point mutations (at this site the two pairing chromosomal regions do not differ in size) display an equal frequency of DSB formation. Both alleles (wild-type and substitution) can undergo DSB formation and thereby become the recipient for sequence copying, with a similar probability (~ 50 to 50%). Middle column: At a chromosomal locus being heterozygous for an insertion, disparity in gene conversion preferentially duplicates (> 50%) the insertion rather than lose (< 50%) the extra genetic information. In this case, DSB is preferentially formed on the shorter (wild-type) homologous region, thereby becoming the recipient of sequence copying. Right column: Biased conversion involving a heterozygous deletion duplicates the wild-type allele, thereby restoring the original genetic information at the site of deletion. Here, too, the shorter (deletion) allele becomes the homolog that preferentially undergoes DSB formation. These mechanisms imply that meiotic recombination is often initiated by self-promoting elements: a wild-type allele transmits itself into its deletion derivative, or an insertion copies itself into its wild-type allele. Sequence transmission in both cases occurs by biased gene conversion (non-reciprocal transfer of genetic information). DNA helixes are indicated by semi-arrowed colored lines. Black lines between the homologs represent homology pairing. wt: wild-type allele; DSB: DNA double strand break
Conversion bias is also evident in meiotic recombination events involving heterozygous deletions, however, in favor of 6:2 segregations. This bias duplicates the wild-type allele and eliminates deletion (McNight et al. 1981). Thus, deletions (formerly lost DNA sequences) can be repaired by copying the corresponding allelic information from the intact homolog (Fig. 2). For example, deletion in a paternal genetic lineage can be restored to the original DNA content by using genomic information from a non-relative maternal genetic lineage, and vice versa (closely related genomes often share deletions at identical genomic loci and thus cannot repair each other). In other words, lost genetic information on a paternal chromosome can be repaired by copying the corresponding wild-type sequence from the maternal homolog.
In contrast with heterozygous insertions and deletions (at these loci the two homolog differ in DNA content), gene conversion involving a heterozygous point mutation (substitution) generally shows parity, i.e., a nearly equal frequency of 6:2 and 2:6 segregations (Fig. 2) (Nagylaki and Petes 1982). At such a locus, the two homologous chromosomal regions do not diverge in size. Thus, the molecular machinery underlying HR cannot “recognize” a gap in the original DNA content relative to its allelic sequence. In such case, DSB formation can occur with an equal probability on either of the homologs.
Biased gene conversion: recombination is often initiated by self-promoting (“selfish”) DNA elements
During meiotic recombination, the two homologous DNA strands interact before DSB formation occurs. Thus, the initial matching of intact homologs (homology search) precedes and somehow influences DSB formation through a sequence-specific chromatin structure. Indeed, Spo11, the protein that generates the DSB, interacts simultaneously with the two pairing DNA duplexes and produces a non-reversible break on one of the pairing homologs only in the presence of the allelic sequence. We suggest that at chromosomal sites where the pairing homologs differ in size, i.e., heterozygous for a deletion or insertion, an unpaired DNA double-strand loop can be formed on the longer homologous region. This structural change that we call C-loop formation ensures the proper pairing of the flanking homologous sequences (homology is represented by horizontal black lines between the pairing DNA helixes in Fig. 3). Presumably, this gene conversion-directed repair mechanism can only recognize the presence or absence of a DNA sequence relative to its allelic information. When the alleles do not differ in DNA content (size), the mechanism does not show preference to any of the alleles (an equal chance of DSB formation on either homologs).
Meiotic recombination is often induced by inequality in DNA content between the pairing homologous regions. During homology pairing (represented by horizontal black lines between the homologs), which precedes DSB formation, loci that differ in size (i.e., heterozygous for an insertion or deletion) cannot match with each other. The longer allele forms a so-called C-loop to allow the proper pairing of the flanking homologous sequences. This structural change causes a molecular tension in the shorter homolog which can be released by a DSB formation. Thus, DSB is preferentially formed on the shorter allele, thereby conferring disparity in gene conversion. Biased conversion duplicates the insertion or restores deletion to the original DNA content. In this way, “selfish” DNA sequences with no allelic counterparts can effectively copy themselves into the homologous region even if they confer no advantage to the individual or population. dsDNA: double-strand DNA; DSB: DNA double-strand break; wt: wild-type; Δ: longer allele (insertion or a wild-type allele in front of deletion)
Molecular tension formed opposite to C-loop on the shorter homologous region can be released by the formation of a DSB. This model is supported by several experimental observations. First, chromatin structure is known to affect the probability of DBS formation (Pan et al. 2011; Brachet et al. 2012; Cummings et al. 2007). Second, recombination hotspots in mammals are largely determined by the sequence-specific Prdm9 methyltransferase (Székvölgyi et al. 2015). Third, DSB formation is strongly affected by sequences at the allelic position (Xu and Kleckner 1995). These data support the idea that meiotic HR is induced by a self-promoting element (“selfish gene”) that can be effectively copied without conferring advantage to the host (Archetti 2003). We suggest that unused (non-functional) heterozygote sequences can transmit themselves into the corresponding homologous chromosomal regions by biased gene conversion. Mediated by sexual reproduction, such “selfish” sequences effectively disperse within the population and then within the given species.
During meiosis, chromosomes are highly packaged into a condensed chromatin structure to facilitate their proper segregation. At this stage, the only information the recombination protein machinery can process is the presence (“good”) or absence (“wrong”) of a given sequence relative to its allelic information. This may be the basis on which the recombination initiation machinery mediates DSB formation. The lack of a sequence that exists in the allelic region may serve as a key message for the machinery to make the choice of breaking the shorter homolog (“the lack of sequence is always wrong” decision). In this way, more than half of the gametes produced during meiosis will contain the given sequence (without biased conversion around half of the haploid gametes will contain, while the other half will miss, the given sequence if they are generated from a heterozygous diploid cell). As a consequence, more than half of the progeny will contain this particular sequence in the next generation. By passing subsequent generations, more and more individuals share this particular sequence, and eventually almost every genome of the population will possess it. Effective copying of a novel sequence thus requires no actual biological function. Taken together, we suggest that meiotic recombination is frequently induced by self-promoting DNA elements (formerly called “selfish genes”). Gene conversion at the region of DSB formation can encourage the transmission of such neutral alleles.
Sex functions as a DNA loading mechanism: it simultaneously restores formerly lost and transmits newly emerged sequences
Despite its prevalence in eukaryotic organisms, why sex evolved and has been maintained in nature remains unclear. The process has a considerable time and energy demand and may disrupt favorable gene combinations. Furthermore, sexually reproducing populations grow at only half the rate of asexual ones, because females produce males instead of other self-propagating individuals (the twofold cost of sex). Asexual populations hence should rapidly overgrow sexual ones. In contrast, sexually reproducing species are significantly overrepresented in nature, as compared with asexual ones. The advantage of sex is assumed to rely on selection between populations by increasing genetic variation (originally proposed by Weismann in 1904) or preventing the accumulation of deleterious mutations. However, the short-term advantage of sex to individuals for maintaining the process long enough for group selection to operate is still enigmatic.
During meiotic recombination, biased gene conversion involving heterozygous insertions or deletions promotes the copying of the allele that contains the extra genetic information. This suggests that DSB preferentially forms on the shorter homologous region, thereby becoming the recipient of sequence copying (Fig. 3). Transmission of the longer allele occurs even if it provides no advantage to the host, proposing a neutralist rather than a selectionist model for the ubiquitous and highly abundant distribution of non-functional sequences in eukaryotic genomes (Kimura 1968; Eyre-Walker and Hurst 2001; Ohta 2002; Lynch 2007). This is in good accordance with a negative relationship between selection efficiency and genome complexity, implying that many characteristics of genomic structures in eukaryotes may have originated via non-adaptive, stochastic processes. Together, we suggest that meiotic recombination (through biased conversion of heterozygous deletions and insertions) may act as a genome (re)loading mechanism, resulting in a significant increase in genome size during the evolution of eukaryotes. By transferring genetic information from one DNA helix to its homolog, conversion can both restore formerly deleted sequences and allow novel sequences to be integrated into allelic positions. In this way, gene conversion accomplishes two seemingly opposite tasks: maintenance of genetic stability (by restoring lost genetic information) and generation of genetic diversity (by spreading novel sequences within species). The latter can be further increased via crossing over (the reciprocal exchange of flanking sequences), which is quite infrequently associated by gene conversion.
Biased gene conversion can repair deletions, thereby reducing the deleterious mutation rate [22]. Indeed, heterozygous deletions frequently reduce the fitness even when they are inherited in a recessive way (this phenomenon is called haploinsufficiency). For example, female mice heterozygous for a recessive deletion affecting the insulin/insulin-like growth factor (Igf) receptor gene are healthy but show a long-lived phenotype associated with reduced fertility (note that homozygous Igf null mutant mice are unviable) (Holzenberger et al. 2003). Biased conversion decreases the proportion of individual genomes bearing the deletion in the subsequent generations and hence increases the number of potentially viable progeny. Thus, sex can confer a short-term individual advantage by recovering formerly lost DNA sequences, thereby reducing the mutational load even under constant conditions. This suggests that sex in eukaryotes facilitates the restoration of formerly lost genetic information, a trait that may have been inherited from bacterial ancestors (Vellai et al. 1998, 1999; Vellai and Vida 1999; Ortutay et al. 2003; Szöllősi et al. 2006). In sum, by repairing deletions, sex can confer an immediate advantage to the host, explaining why the process could have been maintained stable for millions of years. Most deletions, however, form within intragenic, non-functional sequences and thus are not harmful. This function of sex can be advantageous (by restoring formerly lost functional sequences) or neutral (by copying nonfunctional sequences).
Deletions occur commonly in eukaryotic genomes. The only way a population can purge a deletion from the genome is to eliminate individuals bearing the mutation (lethal mutations) or to enable mating between a bearing and non-bearing individual (homozygous non-lethal mutations and heterozygous lethal ones). Inbreeding has the opposite effect. Closely related genomes often contain deletions at identical genomic loci, so the pairing chromosomes during meiosis are unable to repair (restore formerly deleted sequences) each other.
In addition to eliminating deletions, biased conversion promotes the copying of heterozygous insertions (in this case the wild-type allele behaves as a shorter chromosomal region). This suggests that disparity in gene conversion does not necessarily reduce the deleterious mutations rate. Many insertions are beneficial. For example, a series of tandem duplications of a primordial Hox gene (a master regulator of early animal development) has led to the evolution of Hox gene clusters in various animal taxa (Garcia-Fernàndez 2005). The number of Hox genes in an organism correlates with its biological complexity. The initial duplication of such a (Hox) gene originally had taken place in an individual genome in an early phase of evolution of the lineage. Sexual reproduction then mediated the spreading of the given sequence among individuals of the species, and this process did necessitate no functionality to the gene (although a novel copy of a Hox gene certainly provided an advantage to the host through functional redundancy shared by the paralogs).
A large portion of newly emerged insertions are neutral (e.g., a novel microsatellite). Such a sequence could have spread by gene conversion from an individual genome in which it primarily emerged to eventually all members of a species. Thus, sex increases genetic variability by promoting the spreading of novel sequences. In this case, sex confers no immediate advantage to the host. However, certain insertions are harmful as they are integrated into coding or regulatory sequences, thereby disrupting their function. Duplication of such an insertion by gene conversion generates a homozygous mutant genotype that effectively gets eliminated from the population by selection (as the mutational deterministic hypothesis postulates it). Together, biased conversion ensures the efficient copying of heterozygous neutral and beneficial insertions into the corresponding allelic (wild-type) sequences, leading to their spreading and, eventually, ubiquitous distribution among the individual genomes of a species. These types of insertions could have accumulated during the evolution of eukaryotic genomes, whereas deleterious insertions may have been purged from the genomes by selection against their accumulation. Sex thus simultaneously provides immediate (to individuals) and long-term (to populations) advantages, explaining its origin and stable maintenance during evolution. It can reduce mutational load by restoring formerly lost genetic information and promote the formation of adaptive gene combinations by spreading novel sequences. The essence of both processes stems from disparity in gene conversion that supports the copying of a DNA fragment into the corresponding allelic position lacking the particular fragment. Crossing over, which accompanies gene conversion in a number of recombination events, can further increase genetic diversity by producing novel linkage combinations.
Here we suggest that sex evolved for two main purposes. First, it helps to restore formerly lost DNA sequences in a genetic lineage via copying the given information from another genetic lineage. Sex can repair deletions, thereby providing short-term advantages to individuals. Second, sex mediates the efficient copying and spreading of novel sequences that primarily arose in individual genomes. The process helps accumulate novel genetic innovations into a genetic lineage from another in which the innovation emerged. As a by-product, unused sequences (non-functional genetic innovations) can also be spread during the evolution of eukaryotic genomes, explaining their tendency to accumulate “junk DNA” sequences.
The evolution of genome size in eukaryotes
Through restoring formerly lost genetic information and copying novel sequences into allelic positions, meiotic recombination resulted in a considerable increase in genome size during the evolution of eukaryotes (Vellai et al. 1998; Vellai and Vida 1999). Presumably, this genome (re)loading mechanism provided the genetic basis for further increases in biological complexity (across many taxa, the number of genes a given genome codes for correlates with the morphological complexity of the organism). It can also account for the tendency of eukaryotic genomes to accumulate nonfunctional sequences. In addition, meiotic recombination, by facilitating the transfer of genetic material between different genomes with sufficient homology, ensures the cohesion of continuously diverging clonal genetic structures (populations) into a taxonomic unit (species); it eliminates heterozygous deletions, causing the corresponding wild-type alleles to become homozygous, as well as duplicate heterozygous insertions into homozygous genotypes. A previous model for the increased eukaryotic genome size also suggests that restructuring of eukaryotic genomes during evolution was initiated essentially by non-adaptive processes, which, however, were mainly driven by the strength of selection and drift (Lynch and Conery 2003).
It is worth seeing an example of how a newly emerged nonfunctional DNA sequence of germ-line origin can spread within a species by sex (Fig. 4). Such a novel sequence generally arises by a gene duplication event that occurs during replication or recombination. For example, when a short microsatellite undergoes duplication, the replicate often integrates into a chromosomal region adjacent to the original copy (tandem gene duplication). So, the original microsatellite becomes repeated. At this stage, three copies of the sequence exist in the genome: the original two copies (if we consider a homozygous karyotype for this locus) and the novel one. Initially, the new sequence (duplicate; indicated by red coloring in Fig. 4) is heterozygous as it arose by a single duplication event in an individual genome. During meiotic recombination involving this particular chromosomal region, biased conversion transmits the duplicate into the wild-type allele located on the homolog. As being “shorter,” the wild-type allele is preferentially filled by the insertion. As a result, more than half of the gametes produced by the individual in which the duplication occurred will contain the insertion. In the following generation, the duplication-bearing gametes fuse with those derived from other individuals being homozygous for the corresponding wild-type allele. In their progeny, which are heterozygous for the insertion, biased conversion increases the ratio of gametes that contain the insertion relative to those bearing the corresponding wild-type allele. Over time, the novel microsatellite can spread within the population by HR, and this process continues until the microsatellite accumulates in almost all genomes of the species at an identical genomic locus (Fig. 4).
Model showing how sex contributes to the expansion of DNA content in a eukaryotic species. For simplicity, the population consists of single-cell individuals (protozoa) containing only a single pair of homologous chromosomes (indicated by two blue lines). Occurring primarily in an individual genome, a gene duplication event leads to a novel copy (a red chromosomal region) of a certain gene. This copy can be effectively transmitted into its allelic position by biased gene conversion (indicated by a small black arrow). If the population propagates asexually, the bearing diploid genomes produce gametes, nearly the half of which contain the novel sequence, while the other half does not. The proportion of the novel sequence thus cannot be increased as the generations pass each other. However, if the population reproduces sexually, meiotic recombination (through biased gene conversion) can increase the proportion of gametes that contains the novel sequence. As a result, the proportion of individuals bearing this novel copy gradually increases in the population with the passing generations. Migration between populations allows the sharing of the new sequence between each population of the species. Eventually, this novel sequence appears at a certain genomic locus in each member of the species
In bacteria, this evolutionarily conserved mechanism (called localized sex that relies on natural genetic transformation of DNA fragments released from lysed cells, and involves only parts of the genome) continuously expands genome size. To counteract this genome loading process, bacterial genomes also tend to eliminate unused sequences through a series of (micro)deletions, a process called genome economization (Vellai et al. 1998; Szöllősi et al. 2006). This genome erosion mechanism is driven by the so-called R reproductive strategy: cell division in prokaryotes is coupled to the completion of replication. A bacterial cell divides after its genome becomes duplicated (in other words a smaller genome can be duplicated faster than a larger one). As a consequence of these two opposite processes, genome loading, and streamlining, the genome size of contemporary bacterial species fluctuates between ranges of 1 and 9 Megabp.
In contrast, eukaryotes do not rely on genome economization, most probably due to their compartmentalized energy metabolism. At a very early phase of eukaryote evolution, the compartmentalization of energy-converting metabolism (the endosymbiotic emergence of the mitochondrion from a free-living alpha-proteobacterial ancestor) largely liberated the host cell from energetic limitation to expand DNA content. Replication in eukaryotes is limited to the S phase of the cell cycle, where sufficient time and (stored) energy are available for the duplication of the genome. From this innovation (the emergence of compartmentalized energy-converting metabolism), genome size was no longer restricted by size constraints. This allowed the development of vastly expanded genomes, providing the genetic basis for further changes in biological complexity (Fig. 4).
Thus, every novel gene arising primarily in an individual genome could spread within a given species by sex (i.e., biased gene conversion), rather than by natural selection. For example, if a genetic lineage (population) creates a novel Hox paralog, while another one generates a novel Igf receptor-encoding gene, individuals can accumulate both novel genetic innovations in the population even without a selection pressure. Thus, there is no need for the organism to decide which of the innovations (the Hox or Igf paralog) is better (i.e., which of them provides a more significant advantage to the population over the other). Both novel genes can spread within the species from the individuals in which they emerged by biased gene conversion.
Conclusions
Here, we presented the so-called genome loading model for the origin and maintenance of sexual reproduction. Accordingly, sex primarily evolved to restore formerly lost DNA sequences (repair of deletions) and to copy newly arisen sequences into allelic positions. Both processes are driven by biased gene conversion. The former function of sex reduces mutation rate in populations through reducing the proportion of deletions in the gametes generated during meiosis. The latter increases genetic variability through elevating the proportion of novel sequences, each of which primarily arose in an individual genome. This model simultaneously involves Bengtsson’s hypothesis (deletions can be recognized as DNA gaps during chromosome pairing and filled by biased gene conversion; Bengtsson 1985), the environmental stochastic hypotheses (more rapid accumulation of beneficial and neutral insertions; and the mutational deterministic hypotheses (an increased efficiency of selection against synergistically interacting deleterious insertions). Both elimination of (heterozygous) deletions and accumulation of beneficial insertions can provide immediate (individual) advantage to the host, explaining the emergence and short-term maintenance of sex. Note that beneficial point mutations can be accumulated, while deleterious ones can be eliminated, more rapidly in populations by crossing over (according to the environmental stochastic and mutational deterministic hypotheses), but these effects generally rely on long-acting (group) selection pressures. Together, the genome reloading model for the origin of sexual reproduction postulates that sex provides both the advantage of generating novel, adaptive gene combinations (neutral and beneficial insertions) and preventing the accumulation of deleterious mutations (deletions).
Availability of supporting data
Not applicable.
Abbreviations
- CDK-S:
-
Cyclin-dependent kinase Cdc28
- COMPASS:
-
Complex Proteins Associated with Set1
- D-loop:
-
Displacement loop
- DDK:
-
The Dbf4-dependent Cdc7 kinase
- dHJ:
-
Double Holliday junction formation
- DNA:
-
Deoxyribonucleic acid
- DSB:
-
DNA double-strand break
- DSBR:
-
DNA double-strand break repair
- Hox:
-
Homeobox-containing homeotic genes
- H2AK5ac:
-
Histone H2A lysine 5 acetylation
- H3K4me3:
-
Trimethylation of lysine 4 residue on the histone H3 protein subunit
- LLR:
-
Large loop repair
- MMR:
-
Mismatch repair
- PCNA:
-
Proliferating cell nuclear antigen
- PHD:
-
Plant Homeo Domain
- SDSA:
-
Synthesis-dependent strand annealing
- ssDNA:
-
Single-strand deoxyribonucleic acid
- RPA:
-
Replication protein A
References
Acquaviva L, Székvölgyi L, Dichtl B, Dichtl BS, de La Roche Saint André C, Nicolas A, Géli V (2013) The COMPASS subunit Spp1 links histone methylation to initiation of meiotic recombination. Science 339:215–218
Archetti M (2003) A selfish origin for recombination. J Theor Biol 223:335–346
Atcheson CL, DiDomenico B, Frackman S, Esposito RE, Elder RT (1987) Isolation, DNA sequence, and regulation of a meiosis-specific eukaryotic recombination gene. Proc Natl Acad Sci USA 84:8035–8039
Barton NH, Charlesworth B (1998) Why sex and recombination? Science 281:1986–1990
Baudat F, Buard J, Grey C, Fledel-Alon A, Ober C, Przeworski M, Coop G, de Massy B (2010) PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science 327:836–840
Bengtsson BO (1986) Biased conversion as the primary function of recombination. Genet Res 47:77–80
Benjamin KR, Zhang C, Shokat KM, Herskowitz I (2003) Control of landmark events in meiosis by the CDK Cdc28 and the meiosis-specific kinase Ime2. Genes Dev 17:1524–1539
Bergerat A, de Massy B, Gadelle D, Varoutas PC, Nicolas A, Forterre P (1997) An atypical topoisomerase II from Archaea with implications for meiotic recombination. Nature 386:414–417
Bernstein C, Bernstein H (1991) Aging, sex and DNA repair. Academic Press
Borde V, Robine N, Lin W, Bonfils S, Géli V, Nicolas A (2009) Histone H3 lysine 4 trimethylation marks meiotic recombination initiation sites. EMBO J 28:99–111
Brachet E, Sommermeyer V, Borde V (2012) Interplay between modifications of chromatin and meiotic recombination hotspots. Biol Cell 104:51–69
Butlin R (2002) Evolution of sex: the costs and benefits of sex: new insights from old asexual lineages. Nat Rev Genet 3:311–317
Charlesworth B (1976) Recombination modification in a fluctuating environment. Genetics 83:181–195
Chen JF, Lu F, Chen SS, Tao SH (2006) Significant positive correlation between the recombination rate and GC content in the human pseudoautosomal region. Genome 49:413–419
Chen J-M, Cooper DN, Chuzhanova N, Férec C, Patrinos GP (2007) Gene conversion: mechanisms, evolution and human disease. Nat Rev Genet 8:762–775
Chen X, Cui D, Papusha A, Zhang X, Chu CD, Tang J, Chen K, Pan X, Ira G (2012) The Fun30 nucleosome remodeller promotes resection of DNA double-strand break ends. Nature 489:576–580
Cloud V, Chan Y-L, Grubb J, Budke B, Bishop DK (2012) Rad51 is an accessory factor for Dmc1-mediated joint molecule formation during meiosis. Science 337:1222–1225
Corrette-Bennett SE, Mohlman NL, Rosado Z, Miret JJ, Hess PM, Parker BO, Lahue R (2001) Efficient repair of large DNA loops in Saccharomyces cerevisiae. Nucleic Acids Res 29:4134–4143
Crow JF, Simmons MJ (1983) The mutation load in Drosophila. In: Ashburner M, Carson HL, Thompson JN (eds) The genetics and biology of Drosophila, vol 3. Academic Press, pp 1–35
Cummings WJ, Yabuki M, Ordinario EC, Bednarski DW (2007) Chromatin structure regulates gene conversion. PLoS Biol 5:e246
Danilowicz C, Lee CH, Kim K, Hatch K, Coljee VW, Kleckner N, Prentiss M (2009) Single molecule detection of direct, homologous, DNA/DNA pairing. Proc Natl Acad Sci USA 106:19824–19829
Dawkins R (1976) The selfish gene. Oxford University Press, Oxford
de Massy B (2003) Distribution of meiotic recombination sites. Trends Genet 19:514–522
Doolittle WF (2013) Is junk DNA bunk? A critique of ENCODE. Proc Natl Acad Sci USA 110:5294–5300
Doolittle WF, Sapienza C (1980) Selfish genes, the phenotype paradigm and genome evolution. Nature 284:601–603
Dougherty EC (1955) Comparative evolution and the origin of sexuality. Syst Zool 4:145–169
Eshel I, Feldman MW (1970) On the evolutionary effect of recombination. Theor Popul Biol 1:88–100
Ettinger L (1986) Meiosis: a selection stage preserving the genome’s pattern of organization. Evol Theor 8:17–26
Eyre-Walker A, Hurst LD (2001) The evolution of isochores. Nat Rev Genet 2:549–555
Fisher RA (1930) The genetical theory of natural selection: a complete, variorum edn. Oxford University Press
Garcia V, Gray S, Allison RM, Cooper TJ, Neale MJ (2015) Tel1(ATM)-mediated interference suppresses clustered meiotic double-strand-break formation. Nature 520:114–118
Garcia-Fernàndez J (2005) The genesis and evolution of homeobox gene clusters. Nat Rev Genet 6:881–892
Geodakyan VA (1965) Role of the sexes in the transmission and transformation of genetic information. Probl Peredachi Inf 1:105–112
Gladyshev E, Kleckner N (2014) Direct recognition of homology between double helices of DNA in Neurospora crassa. Nat Commun 5:3509
Grabarz A, Barascu A, Guirouilh-Barbat J, Lopez BS (2012) Initiation of DNA double strand break repair: signaling and single-stranded resection dictate the choice between homologous recombination, non-homologous end-joining and alternative end-joining. Am J Cancer Res 2:249–268
Haber JE, Ira G, Malkova A, Sugawara N (2004) Repairing a double-strand chromosome break by homologous recombination: revisiting Robin Holliday’s model. Philos Trans R Soc B 359:79–86
Holliday R (1988) A possible role for meiotic recombination in germline reprogramming and maintenance in the evolution of sex. Sinauer Associates, Sunderland
Holzenberger M, Dupont J, Ducos B, Leneuve P, Geloen A, Even PC, Cervera P, Le Bouc Y (2003) IGF-1 receptor regulates lifespan and resistance to oxidative stress in mice. Nature 421:182–187
Huertas P (2010) DNA resection in eukaryotes: deciding how to fix the break. Nat Struct Mol Biol 17:11–16
Jensen LE, Jauert PA, Kirkpatrick DT (2005) The large loop repair and mismatch repair pathways of Saccharomyces cerevisiae act on distinct substrates during meiosis. Genetics 170:1033–1043
Jiricny J (2006) The multifaceted mismatch-repair system. Nat Rev Mol Cell Biol 7:335–346
Johnson-Schlitz DM, Engels WR (1993) P-element-induced interallelic gene conversion of insertions and deletions in Drosophila melanogaster. Mol Cell Biol 13:7006–7018
Kearney HM, Kirkpatrick DT, Gerton JL, Petes TD (2001) Meiotic recombination involving heterozygous large insertions in Saccharomyces cerevisiae: formation and repair of large, unpaired DNA loops. Genetics 158:1457–1476
Keeney S, Kleckner N (1995) Covalent protein-DNA complexes at the 5’ strand termini of meiosis-specific double-strand breaks in yeast. Proc Natl Acad Sci USA 92:11274–11278
Keeney S, Lange J, Mohibullah N (2014) Self-organization of meiotic recombination initiation: general principles and molecular pathways. Annu Rev Genet 48:187–214
Kimura M (1968) Evolutionary rate at the molecular level. Nature 217:624–626
Kirkpatrick DT (1999) Roles of the DNA mismatch repair and nucleotide excision repair proteins during meiosis. Cell Mol Life Sci 55:437–449
Kondrashov AS (1982) Selection against harmful mutations in large sexual and asexual populations. Genet Res 40:325–332
Kondrashov AS (1993) Classification of hypotheses on the advantage of amphimixis. J Heredity 84:372–387
Kornyshev AA, Leikin S (2001) Sequence recognition in the pairing of DNA duplexes. Phys Rev Lett 86:3666–3669
Kumar R, Bourbon H-M, de Massy B (2010) Functional conservation of Mei4 for meiotic DNA double-strand break formation from yeasts to mice. Genes Dev 24:1266–1280
Kusano K, Sonuhara Y, Takahasi N, Yoshikura H, Kobayashi I (1994) DNA double-strand break repair: genetic determinants of flanking crossing over. Proc Natl Acad Sci USA 91:1173–1177
Lange J, Pan J, Cole F, Thelen MP, Jasin M, Keeney S (2011) ATM controls meiotic double-strand-break formation. Nature 479:237–240
Lloyd DG (1980) Sexual strategies in plants I. A hypothesis of serial adjustment of maternal investment during one reproductive session. New Phytol 86:69–79
Lynch M (2007) The frailty of adaptive hypotheses for the origins of organismal complexity. Proc Natl Acad Sci USA 104(Suppl 1):8597–8604
Lynch M, Conery JS (2003) The origins of genome complexity. Science 302:1401–1404
Manning JT (1982) Sex and the fixation of single favourable mutations. J Theor Biol 94:905–908
Manning JT, Jenkins J (1980) The “balance” argument and the evolution of sex. J Theor Biol 86:593–601
Mather K (1943) Polygenic inheritance and natural selection. Biol Rev 18:32–64
Maynard-Smith J (1978) The evolution of sex. Cambridge University Press, Cambridge
McMahill MS, Sham CW, Bishop DK (2007) Synthesis-dependent strand annealing in meiosis. PLoS Biol 5:e299
McNight GL, Cardillo TS, Sherman F (1981) An extensive deletion causing overproduction of yeast iso-2-cytochrome c. Cell 25:409–419
Morgan TH (1913) Heredity and sex. Columbia University Press
Morgan TH (1919) The physical basis of heredity. JB. Lippincott Company
Muller HJ (1932) Some genetic aspects of sex. Am Nat 66:118–138
Muller HJ (1964) The relation of recombination to mutational advance. Mutat Res 1:2–9
Murakami H, Keeney S (2014) Temporospatial coordination of meiotic DNA replication and recombination via DDK recruitment to replisomes. Cell 158:861–873
Nagylaki T, Petes TD (1982) Intrachromosomal gene conversion and the maintenance of sequence homogeneity among repeated genes. Genetics 100:315–337
Neale MJ, Keeney S (2006) Clarifying the mechanics of DNA strand exchange in meiotic recombination. Nature 442:153–158
Nobrega MA, Zhu Y, Plajzer-Frick I, Afzal V, Rubin EM (2004) Megabase deletions of gene deserts result in viable mice. Nature 431:988–993
Ohta T (2002) Near-neutrality in evolution of genes and gene regulation. Proc Natl Acad Sci USA 99:16134–16137
Ortutay C, Gáspári Z, Tóth G, Jáger E, Vida G, Orosz L, Vellai T (2003) Speciation in Chlamydia: genomewide phylogenetic analyses identified a reliable set of acquired genes. J Mol Evol 57:672–680
Otto SP, Lenormand T (2002) Resolving the paradox of sex and recombination. Nat Rev Genet 3:252–261
Pan J, Sasaki M, Kniewel R, Murakami H, Blitzblau HG, Tischfield SE, Zhu X, Neale MJ, Jasin M, Socci ND, Hochwagen A, Keeney S (2011) A hierarchical combination of factors shapes the genome-wide topography of yeast meiotic recombination initiation. Cell 144:719–731
Panizza S, Mendoza MA, Berlinger M, Huang L, Nicolas A, Shirahige K, Klein F (2011) Spo11-accessory proteins link double-strand break sites to the chromosome axis in early meiotic recombination. Cell 146:372–383
Paques F, Haber J (1999) Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 49:33–58
Partridge L, Hurst LD (1998) Sex and conflict. Science 281:2003–2008
Petes TD (2001) Meiotic recombination hot spots and cold spots. Nat Rev Genet 2:360–368
Ponting CP, Hardison RC (2011) What fraction of the human genome is functional? Genome Res 21:1769–1776
Prahlad V, Pilgrim D, Goodwin EB (2003) Roles for mating and environment in C. elegans sex determination. Science 302:1046–1049
Robert T, Nore A, Brun C, Maffre C, Crimi B, Bourbon HM, de Massy B (2016) The TopoVIB-Like protein family is required for meiotic DNA double-strand break formation. Science 351:943–949
Serrentino M-E, Borde V (2012) The spatial regulation of meiotic recombination hotspots: are all DSB hotspots crossover hotspots? Exp Cell Res 318:1347–1352
Sostak JW, Orr-Weaver TL, Rothstein RJ, Stahl FW (1983) The double-strand break repair model for recombination. Cell 33:25–35
Stern C (1943) The Hardy–Weinberg law. Science 97:137–138
Sturtevant AH, Mather K (1938) The interrelations of inversions, heterosis and recombination. Am Nat 72:447–452
Szathmáry E (2015) Toward major evolutionary transitions theory 2.0. Proc Natl Acad Sci USA 112:10104–10111
Székvölgyi L, Ohta K, Nicolas A (2015) Initiation of meiotic homologous recombination: flexibility, impact of histone modifications, and chromatin remodeling. Cold Spring Harb Perspect Biol 7:a016527-a116516
Szöllősi G, Derényi I, Vellai T (2006) The maintenance of sex in bacteria is ensured by its potential to reload genes. Genetics 174:2173–2180
Treisman M (1976) The evolution of sexual reproduction: a model which assumes individual selection. J Theor Biol 60:421–431
Trivers RL, Hare H (1976) Haploidploidy and the evolution of the social insect. Science 191:249–263
Vellai T, Vida G (1999) The origin of eukaryotes: the difference between prokaryotic and eukaryotic cells. Proc R Soc Lond B 266:1571–1577
Vellai T, Takács K, Vida G (1998) A new aspect to the origin of eukaryotes. J Mol Evol 46:499–507
Vellai T, Kovacs AL, Kovacs G, Ortutay C, Vida G (1999) Genome economization and a new approach to the species concept in bacteria. Proc R Soc Lond B 266:1953–1958
Venter JC, Adams MD, Myers EW, Li PW, Mural RJ et al (2001) The sequence of the human genome. Science 291:1304–1351
Wagner CR, Kuervers L, Baillie DL, Yanowitz JL (2010) xnd-1 regulates the global recombination landscape in Caenorhabditis elegans. Nature 467:839–843
Webster MT, Hurst LD (2012) Direct and indirect consequences of meiotic recombination: implications for genome evolution. Trends Genet 28:101–109
Weismann A (1904) The evolution theory. Edward Arnold, London
Xu L, Kleckner N (1995) Sequence non-specific double-strand break and interhomolog interactions prior to double-strand break formation at a meiotic recombination hot spot in yeast. EMBO J 14:5115–5128
Xu X, Peng M, Fang Z, Xu X (2000) The direction of microsatellite mutations is dependent upon allele length. Nat Genet 24:396–399
Youds JL, Boulton SJ (2011) The choice in meiosis -defining the factors that influence crossover or non-crossover formation. J Cell Sci 124:501–513
Zickler D, Kleckner N (1999) Meiotic chromosomes: integrating structure and function. Annu Rev Genet 33:603–754
Zickler D, Kleckner N (2015) Recombination, pairing, and synapsis of homologs during meiosis. Cold Spring Harb Perspect Biol 7:a016626-16626
Acknowledgements
We are grateful to Gábor Vida and deceased István Molnár (Eötvös Loránd University, Department of Genetics, Budapest, Hungary) for their valuable comments on the genome (re)loading model of sex.
Funding
Open access funding provided by Eötvös Loránd University. Not available.
Author information
Authors and Affiliations
Contributions
AT, LS, and TV wrote the paper; AT and TV formulated the genome loading model for the origin of sex; AT and LS built up the mechanism of homologous recombination.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interest.
Ethical approval and consent to participate
Each author read the manuscript before submission and agreed with its content.
Consent for publication
Each author agreed with this submission to Biologica Futura.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tóth, A., Székvölgyi, L. & Vellai, T. The genome loading model for the origin and maintenance of sex in eukaryotes. BIOLOGIA FUTURA 73, 345–357 (2022). https://doi.org/10.1007/s42977-022-00148-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42977-022-00148-x