Abstract
Freshwater ecosystems are under multiple stressors and it is crucial to find methods to better describe, manage, and sustain aquatic ecosystems. Ecosystem modelling has become an important tool in integrating trophic relationships into food webs, assessing important nodes using network analysis, and making predictions via simulations. Fortunately, several modelling techniques exist, but the question is which approach is relevant and applicable when? In this study, we compare three modelling frameworks (Ecopath, Loop Analysis in R, STELLA software) using a case study of a small aquatic network (8 nodes). The choice of framework depends on the research question and data availability. We approach this topic from a methodological aspect by describing the data requirements and by comparing the applicability and limitations of each modelling approach. Each modelling framework has its specific focus, but some functionalities and outcomes can be compared. The predictions of Loop Analysis as compared to Ecopath’s Mixed Trophic Impact plot are in good agreement at the top and bottom trophic levels, but the middle trophic levels are less similar. This suggests that further comparisons are needed of networks of varying resolution and size. Generally, when data are limiting, Loop Analysis can provide qualitative predictions, while the other two methods provide quantitative results, yet rely on more data.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ecosystem modelling has greatly evolved over the past decades with several software frameworks available to scientists and resource managers (Geary et al., 2020). Ecosystem models have been used as a tool to integrate ecosystem components with processes (e.g., trophic interactions) into a representative model. Here, we focus on network-based trophic models in which the components of a system are described in terms of nodes (e.g., species or functional groups) and their trophic interactions (e.g., diet matrix) (Jordán & Scheuring, 2004; Belgrano et al., 2005). These are simplified versions of an ecosystem, focusing on connecting consumer(s) with the resource(s) (e.g., predator–prey interactions). These interactions can be based on presence-absence (i.e., connectance web) or weighted (quantitative) connections (Woodward et al., 2005). Each model has limitations and assumptions, under which the question of interest can be examined, which necessarily means that there is no perfect model. However, depending on data availability (e.g., quantitative vs qualitative), there are considerations on which modelling approach is most suitable for the research question in a context-dependent way.
Our objective is to compare three commonly used but rarely compared modelling frameworks (Ecopath, STELLA, Loop Analysis) using a case study of a general lake model. We discuss the data requirements and the potential of incorporating social systems (e.g., socio-ecological models). It is imperative to apply interdisciplinary, system-level thinking to the complex problems facing society today (Richmond, 1993; Saviano et al., 2019). As aquatic ecosystems are under increasing anthropogenic pressure and their biodiversity is threatened globally (Dudgeon et al. 2019; Sala et al., 2000), the need to better describe and understand these systems is urgently needed.
First, we provide a brief overview of the modelling frameworks. In Table 1, a summary table was compiled describing several practical aspects of the three frameworks and their main references. Then, we discuss the preliminary model outcomes using a case study of a freshwater lake by comparing the STELLA and Loop Analysis models to the Ecopath model.
Ecopath
Ecopath with Ecosim (EwE) is a comprehensive modelling framework initially developed in the early 1980s, based on the theoretical approach by Polovina (1984a, 1984b), and since then, continuously improved with various extensions and specific functionalities (Christensen et al., 2005; Steenbeek et al., 2016). It has three major parts: Ecopath (static, mass-balanced system); Ecosim (time-dynamic simulation); and Ecospace (spatial and temporal simulation) (Christensen et al., 2005). Its online data repository, EcoBase (Colléter et al., 2013) holds over 200 freely accessible models (to date). The objective of these trophic models is to address general ecological questions (e.g., food web network analysis, Adebola & de Mutsert, 2019; D’Alelio et al., 2016; Rahman et al., 2019) and to investigate more complex issues (e.g., fisheries management, policy options, Kao et al., 2014; Mackinson & Daskalov, 2007).
STELLA
STELLA (Systems Thinking, Experimental Learning Laboratory with Animation) is a visual programming software developed in 1985 (Richmond et al. 1985). The idea behind the creation of STELLA was to approach problems requiring "systems thinking" by utilizing four building blocks (stock, flow, connector, converter) (Richmond, 1994). The program applies numerical simulation by solving ordinary differential equations (ODE), although only to 2-digit-precision (Cellier, 2008). Complex socio-economic models have been integrated using STELLA, such as the World3 model on global sustainability (Meadows et al., 1974, 2004). As an icon-based dynamic modelling and simulation tool, it has been used in education with interdisciplinary applications in environmental systems (Deaton & Winebrake, 1999; Ford, 2010). Recently, an extension to R open-source framework was developed (Naimi & Voinov, 2012).
Loop Analysis
Loop Analysis is a qualitative model (Levins, 1974), that provides a method that is useful where species and their natural history are well-known, but not quantified (Dambacher et al., 2003). Using the qualitative modelling framework of Loop Analysis, one can analyze pathways and feedback in the system, making predictions about the response of variables to perturbations. For example, these can be the addition (increased biomass) or deletion (decreased biomass) of other nodes. Based on feedback and pathways, one can qualitatively specify the direction of change. For getting these predictions, Loop Analysis uses differential equations (Bodini, 2000; Bodini & Clerici, 2016; Fábián, 2021).
Data and methods
Data
Lake Balaton is a large (596 km2), shallow (3.25 m average depth), freshwater lake located in Hungary (Istvánovics et al., 2008). A freshwater lake model was created with 8 general functional groups and fishing pressure added, in three modelling frameworks: Ecopath, STELLA, and Loop Analysis. For Lake Balaton, our model ecosystem, an Ecopath model described in Bíró (2002) was taken as baseline. Functional groups were aggregated (Producers = Phytoplankton, Periphyton, Benthic algae; 7 fish species grouped into ‘Other Fishes’) to be more general. These functional groups are purposefully very general for simplicity and to provide the basis for the methodological comparison. In the next section, we describe the creation of each of the models.
Ecopath model
Diet matrix (Appendix A), biomass data (Appendix B-1, Appendix D), and fishing yield (Weiperth et al., 2014) were updated to recent values using literature data from 2000–2020 (manuscript in preparation). 8 nodes describe this lake system, of which six are aggregate functional groups and two nodes are for the main fish species in the lake (Fig. 1). In both Ecopath models, total biomass is comparable across trophic levels (measured in t/km2/year). Total primary production decreased significantly over the two time periods (changing from eutrophic to oligotrophic state, Bernát et al., 2020), being the main ecological difference between the earlier (Bíró, 2002) and this newer Ecopath model. Pre-balance (PREBAL) diagnostics (Link, 2010) were run to check the model's compliance with basic ecological principles (Appendix B-1) and the Mixed Trophic Index (MTI) was obtained (Appendix C, Fig. 3). The MTI table is used to compare with the Loop Analysis predictions.
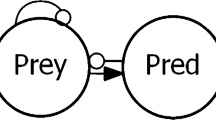
Trophic structure of the Ecopath model (based on Bíró (2002) aggregated and modified with recent data). The model contains eight nodes and fishing pressure. Six nodes are aggregate functional groups (Detritus, Producers, Zooplankton, OBI = Other Benthic Invertebrates, Mollusca, OFish = Other Fish Species), and two nodes represent the main fish species in the lake (Bream = Abramis brama, Pike = Sander lucioperca)
STELLA model
The STELLA model was created in isee systems software (STELLA, 2021; Fig. 2). We used the same input data (as in Ecopath) for the stocks (biomass of functional groups), flows (diet matrix, annual fishing yield), and converters (mortality) (Appendix D). The model was parametrized starting from the bottom up (i.e., producer stock and output flows, then invertebrate stocks and flows, fishes, and finally detritus), ensuring that the relative contribution of diet groups is accurately represented. Some additional assumptions had to be made in order to be able to run the model: 1) production input is constant (oligotrophic state), 2) natural mortality at 18 °C (average annual water temperature) was retrieved for Sander lucioperca and Abramis brama from FishBase (and their average taken for OFish), 3) for invertebrates, natural losses at the stock outflows were estimated to be highest for Zooplankton (75%), and lower for Mollusca (40%) and OBI (25%) (to balance inflows and outflows), 4) living to non-living flows are unknown (not parametrized) and are not connected to detritus (otherwise the detritus stock would accumulate these), except for producers. Further settings are the following: time step (DT = 1 year), Runge–Kutta 2 integration method.
STELLA model with 8 stocks (biomass of functional groups), flows (trophic interactions, fishing yield, and other losses), and converters (natural mortality). Model parameters and equations are found in Appendix D. STELLA symbology: stock (rectangle), flow (double lines with the arrow indicating direction), converter (circle), connector (red arrow)
Loop analysis model
The Loop Analysis model was created in R software (R Development Core Team, 2020). We used MASS 7.3–51.5 and nlme 3.1–148 R packages for the analyzes (Pinheiro et al., 2013; Venables & Ripley, 2002). For simulating sign predictions, we used the R code in Bodini and Clerici (2016). Optionally, a signed digraph figure can be created using GVEdit Graph File Editor For Graphviz version: 1.02 and 2.38 (Ellson et al., 2004), but was not applied here. For making the community matrix, Ecopath model’s diet matrix was used. The community matrix got a 1 (− 1) value where in the diet matrix was a prey-predator (predator–prey) relationship. 0 means there is no trophic connection between two groups. The diagonal terms of the community matrix are self-effects of system variables, represented in signed digraphs as links connecting variables with themselves. These links are self-dampening with self-limiting growth rate, except detritus, because self-limitation was considered only for living groups (Table 2).
We followed the routine described in Bodini and Clerici (2016) to get the predictions for our network (Table 3). The loop formula is used for calculating the equilibrium value of the variables following a perturbation, so it can be deduced how the abundance of a certain variable change (Bodini, 2000):
On the left side, xj is the variable with the equilibrium value being calculated and c is the changing parameter (e.g., mortality, fecundity, abundance). On the right side, f is the growth rate, ∂fi /∂c designates whether the growth rate of the ith variable is increasing or decreasing (positive or negative input, respectively), pji(k) is the pathway connecting the variable to the changed biomass variable (where the perturbation enters the system), Fn-k(comp) is the complementary feedback, which buffers or reverses the effects of the pathway and Fn designates the overall feedback of the system, which is a measure of the inertia of the whole system to change (Bodini, 2000; Bodini & Clerici, 2016). See also Puccia and Levins (1985) for the discussion of the correspondence between matrix algebra and Loop Analysis.
A perturbation on variable j (in this case the perturbation is the increase in the biomass of j) has a net effect (the sum of the direct and indirect effects) on variable i given by the j – ith element of the inverse community matrix [A]−1 (see Levins, 1974; Puccia & Levins, 1985; Raymond et al., 2011). The sign of the coefficients of [A]−1 gives the direction of the expected changes for the variables (Bodini & Clerici, 2016). To make predictions, we used a routine that randomly assigns numerical values from a uniform distribution to the coefficients of the community matrix (these coefficients belong to the links of the signed digraph). This was performed 100 * N2 times, where N is the number of variables in the system. Matrices satisfying the asymptotic Lyapunov criteria were accepted and inverted. The routine of Bodini and Clerici (2016) calculated predictions for the probabilities based on the percentage of positive and negative signs and zeroes in the inverted matrices. They defined a set of rules to make a final table of predictions only from signs (Appendix A in Bodini & Clerici, 2016) which is what we applied in this study (Table 3). Using this routine, from stable matrices we obtained the simulated tables of percentages of ± and 0 and the table of predictions generated from the tables (Table 3).
Results and discussion
The STELLA and Loop Analysis models were compared to the Ecopath model (which is the most comprehensive framework).
Comparison: ecopath and loop analysis
The Mixed Trophic Impact plot from Ecopath (Fig. 3A) could be compared with the predictions of Loop Analysis (Fig. 3B). The MTI quantifies how an infinitesimal increase of any of the impacting groups is predicted to have on the impacted groups (Christensen et al., 2005), while Loop Analysis gives qualitative predictions. The advantage of an MTI plot is also to highlight interactions (and system components) whose importance otherwise might not have been realized (Ulanowicz & Puccia, 1990), including direct and indirect interactions (e.g. competition) in the system (Christensen et al., 2005). For the MTI, a net impact matrix is constructed of all system components in which the element qij indicates the net impact of i upon j. Specifically, the net impact (qij) will equal the difference between gij (the amount that i serves as a prey item for j) and fji (the detrimental impact the consumer i has on the resource j). Also, the indirect effects are taken into account (if i has an effect on j, and j influences k, then there is an indirect pathway between i and k), see Ulanowicz & Puccia, 1990. Fishing pressure is regarded as “predator”, and for detritus gi,j is set to zero. MTI values fall within the range of -1 to + 1, inclusive. The MTI routine based on the net impact matrix by Ulanowicz and Puccia (1990) is automatically implemented in Ecopath and the MTI plot (and numerical values) can be accessed under the Ecopath output tab.
Mixed Trophic Impact plot A and Loop Analysis predictions B. The plots showing positive (blue), negative (red), and neutral (white) combined direct and indirect trophic impacts between the 8 functional groups and fishing pressure. Darker color indicates a stronger effect of the impacting group on the impacted group
For the correlation, we excluded the self-effects due to methodological differences between MTI and Loop Analysis. MTI predicts self-limiting growth (negative effect), so if a group’s abundance or biomass increases, it will not increase infinitely. In contrast, in the prediction matrix of Loop Analysis, the affecting groups are increasing as the representation of the perturbation.
We found a positive correlation between the percentage of positive predictions in Loop Analysis (LA + (%)) and the quantitative values of MTI (r = 0.62, Fig. 4). According to both methods, the groups with the most positive impact on others are Producers and Detritus (Fig. 3). The strongest negative impacts in Loop Analysis matched those of the MTI plot (i.e., Pike-OFish, Producers-Detritus; Fig. 3). The predictions of Loop Analysis as compared to the outcome of the MTI are in good agreement at the top and bottom trophic levels, but the middle trophic levels (e.g., OBI) are less similar. Most of the directions of predictions are similar, but their strength in the MTI plot is stronger (Fig. 3). The quantitative method results in stronger strength of impacts (Fig. 3A) as compared to the qualitative method (Fig. 3B). The difference between the qualitative and quantitative methods is apparent if we observe the Fishing-Pike effect. Fishing has a single-step direct negative impact on the group Pike (Figs. 1 and 4B), while in the quantified network the interaction strength makes the effect more subtle (Fig. 3A). Some of the visibly outlier points of the network (e.g., Detritus-Fishing, Bream-Producers, Producers-Bream, Figs. 3–4) are nodes that represent the most distant groups within the network (having the most trophic steps between them), which means that their predictability is most difficult due to pathway redundancy (Ulanowicz, 1980; Ulanowicz & Puccia, 1990).
Comparison: ecopath and STELLA
Simple dynamic changes can be visualized (graphical and numerical outputs) in the STELLA model, for example, one could easily double the predation rate of pike on bream, which would change system dynamics. However, such predictions from the STELLA framework are not as comprehensive as Ecopath’s MTI, therefore we did not compare them with each other. When focusing on trophic connections between the living groups, we found it possible to recreate Ecopath’s trophic structure with our simple assumptions, meaning that the predicted biomass magnitudes are meaningful and comparable to Ecopath’s biomass values shown in Fig. 1. For example, for fishes, biomass (t/km2/year) in the steady-state are comparable between the two frameworks (Bream: 16.88, OFish: 17.34 in STELLA at the end of the simulation; and Bream: 14.03, OFish: 16.64 in Ecopath). The STELLA model initially overestimated the biomass of pike, therefore an additional fixed loss of 2.4 t/km2/year was subtracted. The primary production is constant, which represents the overall production in the lake during an oligotrophic state (minor interannual deviations are negligible as long as the lake remains in an oligotrophic state). Ultimately, trophic efficiency (called “ecotrophic efficiency” in Ecopath, which is the proportion of the production utilized in the system (e.g. accumulating or moving onto higher trophic levels)) in our model was estimated to be high for producers (~ 90%) and invertebrates (~ 70%), and lower for fish and zooplankton (~ 20–30%) (meaning that primary production and invertebrates are readily consumed, while zooplankton and fish species are mainly lost via natural mortality and diseases), following best practices detailed in Heymans et al. (2016). For the invertebrate groups, both models showed deviation from the expected value (Ecopath gave high values for the Mollusca group (Appendix B-1), and STELLA predicted high values for the Zooplankton group 17.77 vs the input of 9.7 t/km2/year in Ecopath. This difference probably comes from the simplicity of processes (flows) of the STELLA model at the second trophic level.
Fath et al. (2007) discuss the applicability of STELLA numerical simulation as an input to an Ecopath model. To test this, we plugged in the biomass results from the end of numerical simulation (when steady state was reached) into Ecopath (all other settings remained the same as in the original Ecopath model), resulting in comparable PREBAL diagnostics to the original Ecopath model (Appendix B-2). While the Ecopath framework allows for some uncertainty if data is limited (e.g., the software can estimate one input parameter per group), the STELLA model needs to be fully parameterized in order to run, which in some cases leads to inevitable oversimplification of the system (e.g., omission of living to detritus flows due to lack of quantitative data).
Conclusion
In our freshwater lake case study, we found that the output of Ecopath’s MTI plot and the Loop Analysis model are in good agreement. Since Loop Analysis relies only on network topology (assuming no knowledge of interaction strengths in the input; Novak et al., 2011), and Ecopath needs more details of the network (quantitative interaction strength; Ulanowicz & Puccia, 1990), MTI can provide more robust results. Bondavalli et al. (2009) present a hybrid approach, based on a quantitative model of a low-resolution lagoon (7 nodes), in which the net impact matrix was amended with an additional variable (capturing exchanges between the ecosystem components and the surrounding environment), after which the loop analysis procedure was performed. The resolution of the food web can influence many aspects of network analysis (e.g., Martinez et al. 1991; Giacomuzzo & Jordán, 2021). The connectance of the network influences the proportion of correct predictions, thus the reliability of predictions diminishes as network size and connectance increase (Novak et al., 2011). As the number of groups increases within a network, the predictability of Loop Analysis decreases due to pathway redundancy (Ulanowicz, 1980; Ulanowicz & Puccia, 1990), thus it is best recommended for small networks (< 24 nodes, Novak et al., 2011).
Our findings agree with Fath et al. (2007), suggesting transferability between the two frameworks, STELLA and Ecopath. Our recommendation is, that when a system is well documented with quantitative data and the processes are clear, STELLA models can be a great way to better understand the system as a whole (Gertseva et al., 2004; Power et al., 1995; Xuan & Chang, 2014), to highlight important feedback loops (Hayes, 2012; Richmond, 1994), and even to raise awareness about an environmental problem (Jørgensen & De Bernardi, 1997, 1998), and educate the public with its interactive interface (isee Exchange, 2021). Cellier (2008) details further advantages and disadvantages of using the STELLA software for dynamic modelling. Due to the details of parametrization, STELLA is best applicable for small networks (< 10 nodes).
Both frameworks, Loop Analysis and STELLA can be used to complement an existing Ecopath model, for example, if time dynamics predictions are needed, but Ecosim models are not yet available. A limitation of our study is that we only used one small network. It would be important to compare these methods using several networks (of different size, resolution, etc.). Discrepancies between quantitative and qualitative approaches are not surprising and expected to some extent (e.g., less detail and/or information loss is inevitable in qualitative methods compared to quantitative methods). In general, data availability (e.g., lack of information on nodes or on interaction strength) influences food web properties and sampling effects need to be taken into consideration (Berlow et al., 2004; Goldwasser & Roughgarden, 1997). However, not every research question requires numerically parametrized models. For example, it can be qualitatively shown that the effect of an external factor (e.g., aquaculture) negatively impacts certain groups (primary producers, zooplankton, and deposit-feeders), and positively impacts predators and scavengers (Forget et al., 2020). Modelling is thus important to bring out complex, network-level interactions, which might not be evident simply from single parts. The challenge now is to extend these modelling frameworks to social-ecological systems (Martone et al., 2017; Niquil et al., 2021; Rodriguez et al., 2021).
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Adebola, T., & de Mutsert, K. (2019). Comparative network analyses for Nigerian coastal waters using two ecopath models developed for the years 1985 and 2000. Fisheries Research, 213, 33–41. https://doi.org/10.1016/j.fishres.2018.12.028
Belgrano, A., Scharler, U. M., Dunne, J., & Ulanowicz, R. E. (Eds.). (2005). Aquatic food webs: an ecosystem approach. London: Oxford University Press.
Berlow, E. L., Neutel, A.-M., Cohen, J. E., de Ruiter, P. C., Ebenman, Bo., Emmerson, M., Fox, J. W., Vincent, A. A., Jansen, J. I., Jones, G. D., Kokkoris, D. O., Logofet, A. J., McKane, J. M., & Montoya, O. P. (2004). Interaction strengths in food webs: issues and opportunities. Journal of Animal Ecology, 73(3), 585–598. https://doi.org/10.1111/j.0021-8790.2004.00833.x
Bernát, G., Boross, N., Somogyi, B., Vörös, L., László, G., & Boros, G. (2020). Oligotrophication of Lake Balaton over a 20-year period and its implications for the relationship between phytoplankton and zooplankton biomass. Hydrobiologia, 847(19), 3999–4013.
Bíró, P. (2002). A Balaton halállományának hosszúidejű változásai. Állattani Közlemények., 87, 63–77.
Bodini, A. (2000). Reconstructing trophic interactions as a tool for understanding and managing ecosystems: Application to a shallow eutrophic lake. Canadian Journal of Fisheries and Aquatic Sciences, 57, 1999–2009.
Bodini, A., & Clerici, N. (2016). Vegetation, herbivores and fires in savanna ecosystems: A network perspective. Ecological Complexity, 28, 36–46.
Bondavalli, C., Favilla, S., & Bodini, A. (2009). Quantitative versus qualitative modeling: A complementary approach in ecosystem study. Computational Biology and Chemistry, 33(1), 22–28.
Cellier, F. E. (2008). World3 in Modelica: Creating System Dynamics Models in the Modelica Framework. In Proc. 6th International Modelica Conference. Bielefeld, Germany, 2, 393–400.
Christensen, V., Walters, C. J., & Pauly, D. (2005). Ecopath with Ecosim: A user’s guide (p. 154). University of British Columbia, Vancouver.
Colléter M., Valls A. E., Guitton J., Morissette L., Arreguín-Sánchez F. F., Christensen V, Gascuel D, Pauly D (2013) EcoBase: a repository solution to gather and communicate information from EwE models.
D’Alelio, D., Libralato, S., Wyatt, T., & Ribera d’Alcalà, M. (2016). Ecological-network models link diversity, structure and function in the plankton food-web. Science and Reports, 6, 21806. https://doi.org/10.1038/srep21806
Dambacher, J. M., Li, H. W., & Rossignol, P. A. (2003). Qualitative predictions in model ecosystems. Ecological Modelling, 161, 79–93.
Deaton, M. L., & Winebrake, J. J. (2000). Dynamic modeling of environmental systems. New York, NY: Springer New York. https://doi.org/10.1007/978-1-4612-1300-0
Dinno A (2018) LoopAnalyst: A Collection of Tools to Conduct Levins' Loop Analysis. https://CRAN.R-project.org/package=LoopAnalyst Accessed on March 23, 2022
Dudgeon, D. (2019). Multiple threats imperil freshwater biodiversity in the Anthropocene. Current Biology, 29, R960–R967.
Ellson, J., Gansner, E. R., Koutsofios, E., North, S. C., & Woodhull, G. (2004). Graphviz and Dynagraph — Static and Dynamic Graph Drawing Tools. In M. Jünger & P. Mutzel (Eds.), Graph Drawing Software (pp. 127–148). Berlin, Heidelberg: Springer Berlin Heidelberg. https://doi.org/10.1007/978-3-642-18638-7_6
Fábián, V. (2021). Predicting the sign of trophic effects: Individual-based simulation versus loop analysis. Community Ecology, 22(3), 441–451.
Fath, B. D., Scharler, U. M., Ulanowicz, R. E., & Hannon, B. (2007). Ecological network analysis: Network construction. Ecological Modelling, 208(1), 49–558.
Ford A (2010) Modeling the environment. Second edition. Island Press. 1718 Connecticut Avenue, NW, Suite 300, Washington DC, USA.
Forget, N. L., Duplisea, D. E., Sardenne, F., & McKindsey, C. W. (2020). Using qualitative network models to assess the influence of mussel culture on ecosystem dynamics. Ecological Modelling, 430, 109070.
Geary, W. L., Bode, M., Doherty, T. S., Fulton, E. A., Nimmo, D. G., Tulloch, A. I., Tulloch, V. J., & Ritchie, E. G. (2020). A guide to ecosystem models and their environmental applications. Nature Ecology & Evolution, 4(11), 1459–1471.
Gertseva, V. V., Schindler, J. E., Gertsev, V. I., Ponomarev, N. Y., & English, W. R. (2004). A simulation model of the dynamics of aquatic macroinvertebrate communities. Ecological Modelling, 176(1–2), 173–186.
Giacomuzzo, E., & Jordán, F. (2021). Food web aggregation: Effects on key positions. Oikos, 130, 2170–2181. https://doi.org/10.1111/oik.08541
Goldwasser, L., & Roughgarden, J. (1997). Sampling effects and the estimation of food-web properties. Ecology, 78, 41–54.
Hayes, B. (2012). Computation and the human predicament. American Scientist, 100, 186–191.
Heymans, J. J., Coll, M., Libralato, S., Morissette, L., & Christensen, V. (2014). Global patterns in ecological indicators of marine food webs: A modelling approach. PLoS ONE, 9(4), e95845.
Heymans, J. J., Coll, M., Link, J. S., Mackinson, S., Steenbeek, J., Walters, C., & Christensen, V. (2016). Best practice in Ecopath with Ecosim food-web models for ecosystem-based management. Ecological Modelling, 331, 173–184. https://doi.org/10.1016/j.ecolmodel.2015.12.007
isee Exchange (2021) isee Exchange. https://exchange.iseesystems.com/ Accessed on January 3, 2022
Istvánovics, V., Honti, M., Kovács, Á., & Osztoics, A. (2008). Distribution of submerged macrophytes along environmental gradients in large, shallow Lake Balaton (Hungary). Aquatic Botany, 88(4), 317–330.
Jordán, F., & Scheuring, I. (2004). Network ecology: Topological constraints on ecosystem dynamics. Physics of Life Reviews, 1(3), 139–172.
Jørgensen, S. E., & De Bernardi, R. (1997). The application of a model with dynamic structure to simulate the effect of mass fish mortality on zooplankton structure in Lago di Annone. Hydrobiologia, 356(1), 87–96.
Jørgensen, S. E., & De Bernardi, R. (1998). The use of structural dynamic models to explain successes and failures of biomanipulation. Hydrobiologia, 379(1), 147–158.
Kao, Y.-C., Adlerstein, S., & Rutherford, E. (2014). The relative impacts of nutrient loads and invasive species on a Great Lakes food web: An Ecopath with Ecosim analysis. Journal of Great Lakes Research, 40, 35–52. https://doi.org/10.1016/j.jglr.2014.01.010
Levins, R. (1974). Qualitative analysis of partially specified systems. Annals New York Academy of Sciences, 231, 123–138.
Link, J. S. (2010). Adding rigor to ecological network models by evaluating a set of pre-balance diagnostics: a plea for PREBAL. Ecological Modelling, 221(12), 1580–1591. https://doi.org/10.1016/j.ecolmodel.2010.03.012
Mackinson, S., & Daskalov, G. (2007). An ecosystem model of the North Sea to support an ecosystem approach to fisheries management: Description and parameterisation. Cefas Science Series Technical Report, 142, 196.
Martinez, N. D. (1991). Artifacts or attributes? Effects of resolution on the Little Rock Lake food web. Ecological Monographs, 61(4), 367–392.
Martone, R. G., Bodini, A., & Micheli, F. (2017). Identifying potential consequences of natural perturbations and management decisions on a coastal fishery social-ecological system using qualitative loop analysis. Ecology and Society, 22(1), 34. https://doi.org/10.5751/ES-08825-220134
Meadows DH, Randers J, Meadows DL (2004) Limits to Growth: The 30-Year Update, Chelsea Green, White River Junction, Vermont.
Meadows, D. L., Behrens, W. W., Meadows, D. H., Naill, R. F., Randers, J., & Zahn, E. (1974). Dynamics of growth in a finite world. Wright-Allen Press.
Naimi, B., & Voinov, A. (2012). StellaR: A software to translate Stella models into R open-source environment. Environmental Modelling & Software, 38, 117–118.
Niquil, N., Scotti, M., Fofack-Garcia, R., Haraldsson, M., Thermes, M., Raoux, A., Loc’h, L., & Mazé, C. (2021). The Merits of Loop Analysis for the Qualitative Modeling of Social-Ecological Systems in Presence of Offshore Wind Farms. Frontiers in Ecology and Evolution, 9, 35.
Novak, M., Wootton, J. T., Doak, D. F., Emmerson, M., Estes, J. A., & Tinker, M. T. (2011). Predicting community responses to perturbations in the face of imperfect knowledge and network complexity. Ecology, 92(4), 836–846.
Ortiz, M., & Wolff, M. (2008). Mass-balanced trophic and loop models of complex benthic systems in northern Chile (SE Pacific) to improve sustainable interventions: A comparative analysis. Hydrobiologia, 605(1), 1–10.
Pinheiro J, Bates D, DebRoy S, Sarkar D, R Development Core Team (2013) nlme: Linear and Nonlinear Mixed Effects Models. R package version 3.1–108
Polovina JJ (1984a) Model of a coral reef ecosystems I. The ECOPATH model and its application to French Frigate Shoals. Coral Reefs, 3(1):1–11
Polovina, J. J. (1984b). An overview of the ECOPATH model. Fishbyte, 2(2), 5–7.
Power, M. E., Sun, A., Parker, G., Dietrich, W. E., & Wootton, J. T. (1995). Hydraulic food-chain models. BioScience, 45(3), 159–167.
Puccia, C. J., & Levins, R. (1985). Qualitative modeling of complex systems: An Introduction to Loop Analysis and Time Averaging. Harvard University Press. https://doi.org/10.4159/harvard.9780674435070
R Development Core Team. (2020). R: A Language and Environment for Statistical Computing. Austria.
Rahman, M. F., Qun, L., Xiujuan, S., et al. (2019). Temporal Changes of Structure and Functioning of the Bohai Sea Ecosystem: Insights from Ecopath Models. Thalassas, 35, 625–641. https://doi.org/10.1007/s41208-019-00139-1
Raymond, B., McInnes, J., Dambacher, J. M., Way, S., & Bergstrom, D. M. (2011). Qualitative modelling of invasive species eradication on subantarctic Macquarie Island. Journal of Applied Ecology, 48, 181–191.
Richmond B (1985) A User's Guide to STELLA. High Performance Systems. 45 Lyme Road, Hanover, NH 03755, USA
Richmond, B. (1993). Systems thinking: Critical thinking skills for the 1990s and beyond. System Dynamics Review, 9(2), 113–133.
Richmond, B. (1994). Systems thinking/system dynamics: Let’s just get on with it. System Dynamics Review, 10(2–3), 135–157.
Rodriguez, M., Bodini, A., Escobedo, F. J., & Clerici, N. (2021). Analyzing socio-ecological interactions through qualitative modeling: Forest conservation and implications for sustainability in the peri-urban Bogota (Colombia). Ecological Modelling, 439, 109344. https://doi.org/10.1016/j.ecolmodel.2020.109344
Sala, O. E., Chapin, F. S., Armesto, J. J., Berlow, E., Bloomfield, J., Dirzo, R., Huber-Sanwald, E., Huenneke, L. F., Jackson, R. B., Kinzig, A., & Leemans, R. (2000). Global biodiversity scenarios for the year 2100. Science, 287(5459), 1770–1774.
Saviano, M., Barile, S., Farioli, F., & Orecchini, F. (2019). Strengthening the science–policy–industry interface for progressing toward sustainability: A systems thinking view. Sustainability Science, 14(6), 1549–1564.
Steenbeek, J., Buszowski, J., Christensen, V., Akoglu, E., Aydin, K., Ellis, N., Felinto, D., Guitton, J., Lucey, S., Kearney, K., & Mackinson, S. (2016). Ecopath with Ecosim as a model-building toolbox: Source code capabilities, extensions, and variations. Ecological Modelling, 319, 178–189. https://doi.org/10.1016/j.ecolmodel.2015.06.031
STELLA (2021) https://www.iseesystems.com/. Accessed on January 1, 2021.
Ulanowicz RE, Puccia CJ (1990) Mixed trophic impacts in ecosystems. Coenoses, .7–16
Ulanowicz, R. E. (1980). An hypothesis on the development of natural communities. Journal of Theoretical Biology, 85, 223–245.
Venables, W. N., & Ripley, B. D. (2002). Modern Applied Statistics with S (4th ed.). Springer.
Weiperth, A., Ferincz, Á., Kováts, N., Hufnagel, L., Staszny, Á., Keresztessy, K., Szabó, I., Tátrai, I., & Paulovits, G. (2014). Effect of water level fluctuations on fishery and anglers’ catch data of economically utilised fish species of Lake Balaton between 1901 and 2011. Applied Ecology and Environmental Research, 12(1), 221–249.
Woodward, G., Speirs, D. C., Hildrew, A. G., & Hal, C. (2005). Quantification and resolution of a complex, size-structured food web. Advances in Ecological Research, 36, 85–135.
Xuan, Z., & Chang, N. B. (2014). Modeling the climate-induced changes of lake ecosystem structure under the cascade impacts of hurricanes and droughts. Ecological Modelling, 288, 79–93.
Acknowledgements
We would like to thank Central European University (CEU) for providing access to the STELLA software on campus as well as Ferenc Jordán for his helpful advice.
Funding
Open access funding provided by ELKH Centre for Ecological Research. This output reflects only the authors’ view and the European Union cannot be held responsible for any use of the information contained therein.
Author information
Authors and Affiliations
Contributions
VF. performed the Loop Analysis modelling and the statistical analysis, prepared manuscript text and figures. KP. performed the Ecopath and STELLA modelling, prepared manuscript text and figures.
Corresponding author
Ethics declarations
Conflict of interests
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Appendices
Appendix A
See Fig.
Appendix B-1,Bb-2
See Figs
PREBAL diagnostics obtained from Ecopath (Link, 2010). Groups are 1 = Pike, 2 = OFish, 3 = Bream, 4 = Mollusca, 5 = OBI, 6 = Zooplantkon, 7 = Producers. Blue bars indicate values estimated by the model (e.g., biomass data was not feasible to estimate from survey data). Biomass for Mollusca is estimated by Ecopath to be high and Zooplankton biomass (based on survey data) is low. Biomass data were obtained from the best available data from years 2000–2020 for Lake Balaton, Hungary (modified from Bíró, 2002)
.6 and
Appendix C
See Table
BREAM(t) = BREAM(t—dt) + (OBI__to_Bream + Zooplankton__to_Bream + Moll_to__Bream + Detritus__to_Bream—Bream_to_Pike—bream_yield—bream__mortality) * dt.
INIT BREAM = 14.
OBI__to_Bream = OBI*0.4
Zooplankton__to_Bream = ZOO*0.1
Moll_to__Bream = MOLL*0.4
Detritus__to_Bream = 2.
Bream_to_Pike = (BREAM/PIKE)*predation_rate.
bream_yield = 0.03.
bream__mortality = (BREAM*bream_natural__mortality_rate)*PIKE.
DETRITUS(t) = DETRITUS(t—dt) + (Prod_to_Detritus—Detritus__to_Bream—Detritus__to_OFish—Detr_to_Moll—Detr_to_OBI—Detritus_to_Zoo) * dt.
INIT DETRITUS = 10.
Prod_to_Detritus = PRODUCERS*0.1
Detritus__to_Bream = 2.
Detritus__to_OFish = DETRITUS*0.1
Detr_to_Moll = DETRITUS*0.06.
Detr_to_OBI = DETRITUS*0.09.
Detritus_to_Zoo = DETRITUS*0.015.
MOLL(t) = MOLL(t—dt) + (Prod_to_Moll + Detr_to_Moll—Moll_to__Bream—Moll_to_OFish—Loss_at_Moll) * dt.
INIT MOLL = 56.
Prod_to_Moll = PRODUCERS*0.3
Detr_to_Moll = DETRITUS*0.06.
Moll_to__Bream = MOLL*0.4
Moll_to_OFish = MOLL*0.2
Loss_at_Moll = MOLL*0.40.
OBI(t) = OBI(t—dt) + (Prod_to_OBI + Detr_to_OBI—OBI__to_Bream—OBI_to_OFish—Loss_at_OBI) * dt.
INIT OBI = 44.
Prod_to_OBI = PRODUCERS*0.2
Detr_to_OBI = DETRITUS*0.09.
OBI__to_Bream = OBI*0.4
OBI_to_OFish = OBI*0.35.
Loss_at_OBI = OBI*0.25.
OFISH(t) = OFISH(t—dt) + (Prod_to_OFish + Moll_to_OFish + OBI_to_OFish + Detritus__to_OFish + Zooplankton_to_OFish—OFish_to_Pike—OFish__mortality—OFish_yield) * dt.
INIT OFISH = 16.6
Prod_to_OFish = PRODUCERS*0.07.
Moll_to_OFish = MOLL*0.2
OBI_to_OFish = OBI*0.35.
Detritus__to_OFish = DETRITUS*0.1
Zooplankton_to_OFish = ZOO*0.15.
OFish_to_Pike = (OFISH/PIKE)*0.32.
OFish__mortality = OFISH*PIKE*OFish_natural__mortality_rate.
OFish_yield = 0.018.
PIKE(t) = PIKE(t—dt) + (Bream_to_Pike + OFish_to_Pike—pike_yield—pike_mortality) * dt.
INIT PIKE = 1.63.
Bream_to_Pike = (BREAM/PIKE)*predation_rate.
OFish_to_Pike = (OFISH/PIKE)*0.32.
pike_yield = 0.011 + (PIKE*cannibalism).
pike_mortality = 2.4 + (PIKE*pike_natural__mortality_rate).
PRODUCERS(t) = PRODUCERS(t—dt) + (photosynthesis—Prod_to_Moll—Prod_to_OBI—Prod_to_Zooplankton—Prod_to_OFish—Prod_to_Detritus) * dt.
INIT PRODUCERS = 53.3
photosynthesis = production_input.
Prod_to_Moll = PRODUCERS*0.3
Prod_to_OBI = PRODUCERS*0.2
Prod_to_Zooplankton = PRODUCERS*0.33.
Prod_to_OFish = PRODUCERS*0.07.
Prod_to_Detritus = PRODUCERS*0.1
ZOO(t) = ZOO(t—dt) + (Prod_to_Zooplankton + Detritus_to_Zoo—Zooplankton__to_Bream—Zooplankton_to_OFish—Loss_at__Zooplankton) * dt.
INIT ZOO = 9.2
Prod_to_Zooplankton = PRODUCERS*0.33.
Detritus_to_Zoo = DETRITUS*0.015.
Zooplankton__to_Bream = ZOO*0.1
Zooplankton_to_OFish = ZOO*0.15.
Loss_at__Zooplankton = ZOO*0.75.
bream_natural__mortality_rate = 0.32.
cannibalism = PIKE*0.001.
OFish_natural__mortality_rate = 0.285.
pike_natural__mortality_rate = 0.25.
predation_rate = 0.15.
production_input = 53.3
Appendix D. STELLA model equation layer
INIT [group name] indicates the initial biomass (t/km2/yr), which are the same as in the Ecopath model (e.g., INIT ZOO = 9.2 t/km2/yr). Constant values are set for production input, natural mortality rates, and fishing yield. Others are expressed as rate of change: e.g., “Bream_to_Pike = (BREAM/PIKE)*predation_rate” means that the amount of biomass transferring from Bream to Pike is a function of the Bream to Pike ratio, multiplied by the predation rate. “PIKE(t) = PIKE(t—dt) + (Bream_to_Pike + OFish_to_Pike—pike_yield—pike_mortality) * dt” is the difference equation describing the dynamics of the pike stock, where inflow = (Bream_to_Pike + OFish_to_Pike); outflow = (pike_yield—pike_mortality), at each time step (dt). These equations are represented in the STELLA symbology layer (Fig. 2.).
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Patonai, K., Fábián, V.A. Comparison of three modelling frameworks for aquatic ecosystems: practical aspects and applicability. COMMUNITY ECOLOGY 23, 439–451 (2022). https://doi.org/10.1007/s42974-022-00117-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42974-022-00117-3