Abstract
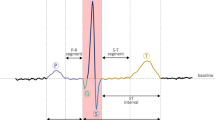
The electrocardiogram (ECG) is a critical, non-invasive tool for diagnosing cardiovascular diseases, offering insights into heart function. However, analyzing extended ECG data can be complex, requiring advanced computerized systems for effective diagnosis and classification. These systems must detect arrhythmias, manage data noise, and adapt to individual waveform variations, while ensuring model robustness across different populations and settings. The main goal of this study is to develop an ECG signal processing system that can accurately detect and classify various cardiac conditions. We propose a novel hybrid approach, classifying ECG signals into categories such as normal, left bundle branch block (LBBB), paced beat, right bundle branch block (RBBB), and supraventricular contraction (SVC) using a PhysioNet database. By applying discrete wavelet transform (DWT) and principal component analysis (PCA), we extracted six relevant features from each ECG category. These features were analyzed using an adaptive neuro-fuzzy inference system (ANFIS) classifier, achieving an overall classification accuracy of 99.44%, with average sensitivity and specificity of 99.36% and 99.84%, respectively. This system shows significant promise in enhancing the accuracy and efficiency of diagnosing cardiovascular diseases through ECG analysis.

















Similar content being viewed by others
Data Availability
In this study, we used the publicly available MIT-BIH Arrhythmia Database from PhysioNet. Link to the original dataset: https://www.physionet.org/content/mitdb/1.0.0/.
References
Jevon P (2010) An introduction to electrocardiogram interpretation: part 1. Emerg Nurse 18:28–36. https://doi.org/10.7748/en2010.04.18.1.28.c7689
Elhaj FA, Salim N, Harris AR et al (2016) Arrhythmia recognition and classification using combined linear and nonlinear features of ECG signals. Comput Methods Programs Biomed 127:52–63. https://doi.org/10.1016/j.cmpb.2015.12.024
Shah AJ, Hocini M, Pascale P et al (2013) Body surface Electrocardiographic Mapping for non-invasive identification of arrhythmic sources. Arrhythmia Electrophysiol Rev 2:16. https://doi.org/10.15420/aer.2013.2.1.16
Kishore B, Gopal Reddy AN, Kumar Chillara A et al (2022) An Innovative Machine Learning Approach for Classifying ECG Signals in Healthcare Devices. J Healthc Eng 2022:. https://doi.org/10.1155/2022/7194419
Joukar S (2021) A comparative review on heart ion channels, action potentials and electrocardiogram in rodents and human: extrapolation of experimental insights to clinic. Lab Anim Res 37:1–15. https://doi.org/10.1186/s42826-021-00102-3
Martinek R, Ladrova M, Sidikova M et al (2021) Advanced bioelectrical signal processing methods: past, present and future approach—part II: brain signals. Sensors 21:1–32. https://doi.org/10.3390/s21196343
Rafie N, Kashou AH, Noseworthy PA (2021) ECG interpretation: clinical relevance, challenges, and advances. Hearts 2:505–513. https://doi.org/10.3390/hearts2040039
Mahapatra S, Mohanta D, Mohanty P et al (2016) A neuro-fuzzy based model for analysis of an ECG Signal using Wavelet Packet Tree. Procedia Comput Sci 92:175–180. https://doi.org/10.1016/j.procs.2016.07.343
Niyigena Ingabire H, Wu K, Toluwani Amos J et al (2022) Analysis of ECG signals by Dynamic Mode Decomposition. IEEE J Biomed Heal Inf 26:2124–2135. https://doi.org/10.1109/JBHI.2021.3130275
Al ZA, Thapa K (2021) Mode selected Energy and adaptive window sizing. Algorithm with Decision Tree Algorithm
Barki H, Chung WY (2023) Mental stress detection using a wearable In-Ear plethysmography. Biosensors 13. https://doi.org/10.3390/bios13030397
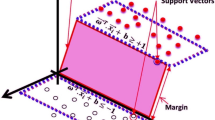
Kohli N, Verma NK, Roy A (2010) SVM based methods for arrhythmia classification in ECG. 2010 Int Conf Comput Commun Technol ICCCT-2010 486–490. https://doi.org/10.1109/ICCCT.2010.5640480
Saini I, Singh D, Khosla A (2014) Electrocardiogram beat classification using empirical mode decomposition and multiclass directed acyclic graph support vector machine. Comput Electr Eng 40:1774–1787. https://doi.org/10.1016/j.compeleceng.2014.04.004
Li H, Yuan D, Wang Y et al (2016) Arrhythmia classification based on multi-domain feature extraction for an ECG recognition system. Sens (Switzerland) 16. https://doi.org/10.3390/s16101744
Turnip A, Ilham Rizqywan M, Kusumandari DE et al (2018) Classification of ECG signal with support Vector Machine Method for Arrhythmia Detection. J Phys Conf Ser 970. https://doi.org/10.1088/1742-6596/970/1/012012
Ilbeigipour S, Albadvi A, Akhondzadeh Noughabi E (2021) Real-Time Heart Arrhythmia Detection using Apache Spark Structured Streaming. J Healthc Eng 2021. https://doi.org/10.1155/2021/6624829
Houssein EH, Ibrahim IE, Neggaz N et al (2021) An efficient ECG arrhythmia classification method based on Manta ray foraging optimization. Expert Syst Appl 181:115131. https://doi.org/10.1016/j.eswa.2021.115131
Moody GB, Mark RG (2001) The impact of the MIT-BIH arrhythmia database. IEEE Eng Med Biol Mag 20:45–50. https://doi.org/10.1109/51.932724
Goldberger AL, Amaral LA, Glass L et al (2000) PhysioBank, PhysioToolkit, and PhysioNet: components of a new research resource for complex physiologic signals. Circulation 101. https://doi.org/10.1161/01.cir.101.23.e215
Rajini GK (2016) A comprehensive review on Wavelet transform and its applications. ARPN J Eng Appl Sci 11:11713–11723
Imah EM, Al Afif F, Ivan Fanany M et al (2011) A comparative study on Daubechies Wavelet Transformation, Kernel PCA and PCA as feature extractors for arrhythmia detection using SVM. IEEE Reg 10 Annu Int Conf Proceedings/TENCON 5–9. https://doi.org/10.1109/TENCON.2011.6129052
Dese K, Ayana G, Lamesgin Simegn G (2022) Low cost, non-invasive, and continuous vital signs monitoring device for pregnant women in low resource settings (lvital device). HardwareX 11:e00276. https://doi.org/10.1016/j.ohx.2022.e00276
Ayana G, Dese K, Raj H et al (2022) De-speckling breast Cancer ultrasound images using a rotationally invariant Block Matching Based Non-local Means (RIBM-NLM) Method. Diagnostics 12:862. https://doi.org/10.3390/diagnostics12040862
Heriana O, Al Misbah AM (2017) Comparison of Wavelet Family performances in ECG Signal Denoising. J Elektron Dan Telekomun 17:1. https://doi.org/10.14203/jet.v17.1-6
Junior EA, Valentim RADM, Brandão GB (2018) Real-time premature ventricular contractions detection based on redundant discrete wavelet transform. Res Biomed Eng 34:187–197. https://doi.org/10.1590/2446-4740.01618
Kitao A (2022) Principal component analysis and related methods for investigating the Dynamics of Biological Macromolecules. J 5:298–317. https://doi.org/10.3390/j5020021
Rajab S (2019) Handling interpretability issues in ANFIS using rule base simplification and constrained learning. Fuzzy Sets Syst 368:36–58. https://doi.org/10.1016/j.fss.2018.11.010
Senthilselvi A, Duela JS, Prabavathi R, Sara D (2021) Performance evaluation of adaptive neuro fuzzy system (ANFIS) over fuzzy inference system (FIS) with optimization algorithm in de-noising of images from salt and pepper noise. J Ambient Intell Humaniz Comput. https://doi.org/10.1007/s12652-021-03024-z
Viattchenin DA, Tati R, Damaratski A (2013) Designing Gaussian membership functions for fuzzy classifier generated by heuristic possibilistic clustering. J Inf Organ Sci 37:127–139
Arpitha Y, Madhumathi GL, Balaji N (2022) Spectrogram analysis of ECG signal and classification efficiency using MFCC feature extraction technique. J Ambient Intell Humaniz Comput 13:757–767. https://doi.org/10.1007/s12652-021-02926-2
Banerjee S, Mitra M (2013) ECG beat classification based on discrete wavelet transformation and nearest neighbour classifier. J Med Eng Technol 37:264–272. https://doi.org/10.3109/03091902.2013.794251
Dhyani S, Kumar A, Choudhury S (2023) Analysis of ECG-based arrhythmia detection system using machine learning. MethodsX 10:102195. https://doi.org/10.1016/j.mex.2023.102195
Sowmya S, Jose D (2022) Contemplate on ECG signals and classification of arrhythmia signals using CNN-LSTM deep learning model. Meas Sens 24:100558. https://doi.org/10.1016/j.measen.2022.100558
Funding
This work is supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (RS-2023-00240521) and project for Industry-University-Research Institute platform cooperation R&D funded by Korea Ministry of SMEs and Startups in 2022 (S3310765).
Author information
Authors and Affiliations
Contributions
All authors contributed to the research design of the study. A.M.A., and S.-w.C.: conceptualization, data creation, analysis, and interpretation of data, analyzed the results, and drafted the manuscript. G.A., A.A.D., data analysis, resource investigation, validation, proofreading and editing of the manuscript. H.B., formal analysis, critical revision of the manuscript. B.L.T., T.J and S.-w.C.: Conceptualization, supervision, visualization, writing review, and editing. All authors approved the final version of the manuscript and agreed to be accountable for all aspects of the work. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Conflict of interest
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Abagaro, A.M., Barki, H., Ayana, G. et al. Automated ECG Signals Analysis for Cardiac Abnormality Detection and Classification. J. Electr. Eng. Technol. (2024). https://doi.org/10.1007/s42835-024-01902-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42835-024-01902-y