Abstract
We examined how various soil characteristics are associated with Scirpus mariqueter growth and spatial heterogeneity in Shanghai Nanhui Dongtan wetlands and addressed a major knowledge gap regarding the effects of reclamation on microbial communities in the soil. Biolog was used to examine soil carbon resources, diversity, abundance, and community structure of S. mariqueter soil microbes after a 72-h culture. Tidal elevation influenced soil microbes, which used carbon resources at higher rates, exhibited more community diversity and had greater species richness in areas with dense S. mariqueter cover than in bare tidal flats (P < 0.05). Microbial functional diversity also differed significantly across the study region in a manner that reflected S. mariqueter spatial heterogeneity (P < 0.05). In terms of soil characteristics, microbial community diversity was positively correlated with soil salinity, organic carbon, and total phosphorus (P < 0.05), whereas negatively correlated with soil particle size (P < 0.05). The results of this study provide insight into plant–soil interactions of disturbed S. mariqueter wetland ecosystems through clarifying factors that influence soil microbes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Scirpus mariqueter is an important native wetland plant in the Yangtze River delta and is widely distributed along the southeastern coast of Shanghai [1, 2]. This species can remove pollutants from the environment through absorption and also provides habitats for native waterfowl [3]. A hardy plant that can survive in anaerobic and high salinity environments, S. mariqueter is a common pioneer species in Yangtze River estuaries [4]. However, heavy population declines due to high-intensity reclamation [5] and massive Spartina alterniflora invasion [6] has led to a major need for remediation.
Previous researchers have applied multiple techniques to investigate reclamation’s effects on plant species, soil, and soil bacteria. Several studies have demonstrated the effects of salt concentration on relative abundance and composition diversity [7] or the correlation between enzymatic activities and soil physiochemical factors [8]. Soil characteristics play a critical role in the growth and distribution of salt marsh plants [9, 10], and reclamation clearly alters soil physicochemical properties [11]. However, although soil microbes are involved in most soil biochemical reactions and played a key role in organic matter decomposition and nutrient cycling [12], we know little about how they respond to reclamation [7]. Because soil microbial biomass [13] and diversity [14] reflect how the environment influences vegetation-soil systems, studies aiming to understand the ecological impact of reclamation should examine both plants and their associated soil microbes.
Microorganisms metabolic diversity classification is an easy but important way to reveal microbial community diversity [15]. Biolog is a powerful method currently known for studying the metabolic diversity of microorganisms. Biolog reflects overall microbial community activity and characterizes microbial functional diversity without the need to analyze individual microorganisms. This simple method of high-resolution has been widely used in microbial ecology and environmental microbial detection over the past 20 years [16]. Biolog established characteristic metabolic fingerprints for each microorganism. The aim of microorganism’s identification can be realized by comparing the fingerprint of the carbon source of the identified microorganism with the database by software. Eco-plates were commonly used to determine functional diversity of microbial communities in recent studies [17]. Yan studied the seasonal changes of microbial community diversity of sea cucumber pond by Biolog microplate method [18]. Lin et al. studied the effects of planting Pennisetum sp. (Giant juncao) on soil microbial functional diversity by using Biolog microplate [19]. Graham and Haynes analyzed the catabolic diversity of soil microbial communities under sugarcane and other land uses by Biolog [20]. Braun et al. applied Biolog method to the characterization of soil microbial diversity in urban soil [21].
In this study, Biolog method was used to determine the difference in utilization of carbon sources by soil microorganisms under different S. mariqueter coverage. And through principal component analysis combined with soil physical and chemical properties, the difference in functional diversity of soil microbial communities was preliminarily revealed. The results should improve our understanding of the relationship between soil characteristics and microbial diversity. In turn, we would gain further insight on S. mariqueter performance in reclaimed lands, which contributes to future coastal remediation efforts in the region.
2 Materials and methods
2.1 Study site and soil sampling
The study area is located on the coasts of Nanhui, Pudong New District Area, Shanghai, (30.860°–30.870°N, 121.921°–121.944°E; Fig. 1). Because the biomass of S. mariqueter reached the maximum in October [22], soil sampling was performed during September to October 2015 from 101 plots to test soil and vegetation characteristics. Site elevation was measured using an optical level (DS3, Shanghai Wahgon, China). S. mariqueter characteristics were obtained from a 1 m × 1 m plot with three replicates. Vegetation density was manually counted. Plant height was computed from the average of the five tallest plants at each sampling point. Coverage was estimated through visual assessment. Aboveground biomass was measured via harvesting material from 25 cm × 25 cm quadrat and then dried them at 105 °C to a constant weight [23]. Soil samples from depths of 0–20 cm were collected in 20 cm × 20 cm plots [24]. These samples were used for soil physiochemical characteristics analyses.
Nanhui Wetland is located at the junction of the Yangtze River estuary and Hangzhou Bay. It is a north subtropical monsoon climate, with abundant sunshine, abundant rainfall, warm, and humid climate, annual average temperature of 15.3 °C, annual precipitation of 1022 mm. The largest coastal wetland of Yangtze River estuary with a total area of about 33,100 h m2. The study area is located in the south of the East China Sea Bridge, in the range of 30.860°–30.870°N and 121.921°–121.944°E
Then, S. mariqueter community was divided into three categories based on calculated coverage: bare tidal flats (0–10%, unvegetated area), sparse areas (10–30%, vegetated area), and dense areas (30–60%, vegetated area) (Fig. 1). Soil samples from these three coverage categories were randomly collected in October 2016 to analyze soil microbes. Soil sampling procedures and methods are the same as the samplings in 2015. Soil samples were 9 in number, each 3 replicates and 27 in total [8]. One subsample of each plot was stored at 4 °C about 3 days for microbial analyses and then stored at − 80 °C. A second subsample was air-dried and sieved by 0.15 mm screen (Shanghai Precision Instrument, China).
2.2 Soil physiochemical characteristics analysis
Soil organic carbon was tested by potassium dichromate-external heating method. Total nitrogen was determined by Kjeldahl method, and total phosphorus was determined by alkali fusion molybdenum antimony spectro-photometric method. Nitrate nitrogen, nitrite nitrogen, and ammonia nitrogen were measured by potassium chloride liquor extract-Spectrophotometry. Data were obtained using an UV-1800 spectrophotometer (SHIMADZU, Japan). The soil pH was measured by soil extract of water soil ratio 2.5:1. Soil salinity was expressed by conductivity, which was measured using a portable multi-parameter water quality analyzer by the extracted liquid of water soil ratio 5:1 [25]. Soil particle size was measured using a Malvern laser particle size analyzer. The specific measures of soil indexes mentioned above refer to the article of Bao [26].
2.3 Biolog experiments
The functional diversity of soil microbes was evaluated through their use of 31 carbon resource types on Biolog ECO-plates. Microbe-specific redox reactions during carbon metabolism will alter the plate color, from colorless for oxidation to purple or red for reduction [27]. Microbial classification was realized from Gen III Microstation automated microbial identification system (American Biolog Corporation).
First, the 96 micro-pores (8 × 12) of an ECO plate were separated into three groups, 32 each group, with the control micro-pore containing water and the remaining 31 containing a different organic carbon source and tetrazole violet dye. Carbon resources were divided into seven categories: monosaccharides/glycosides (4), amino acids (6), glycosides (4), esters (4), alcohols (3), amines (3), and acids (7) [27, 28]. Next, soil samples (1 g) were added to 99 ml sterile water, shaken (250 rpm) for 20 min, and left to stand for 30 min (4 °C)with chemical flocculants (1 g CaCO3 and CaCl2) (Dang et al. 2015). Aliquots (150 μL) were then added to each micro-pore. The ECO plate was cultured at 25 °C for 120 h and monitored every 24 h. The OD (optical density) of each micro-pore was measured using a Biolog reader at 590 nm to 750 nm.
2.4 Calculation of carbon use and diversity indices
The average well color development (AWCD) of each micro-pore after obtaining OD was calculated as follows:
where Ci is the absorbance of the ith non-control micro-pore, R is the absorbance of the control micro-pore, and n is the number of the carbon resource species in the culture medium [29, 30].
After 72-h cultivation, microbial diversity indices were computed, including the Shannon diversity index (H′; measurement of species abundance), McIntosh evenness index (U; measurement of community uniformity), and Simpson index (D; measurement of community diversity). For all three, higher index values represent higher levels of the relevant variable.
The formulas were as follows:
Pi is the ratio of absorbance from the ith non-control micro-pore to the sum of absorbance from all non-control micro-pores, \(n_{i}\) is the number of species [31, 32].
2.5 Statistical analysis
GraphPad Prism 5 was used for mapping S. mariqueter distribution. Correlations between physicochemical factors and microbial diversity and principal component analysis (PCA) of carbon resource use were performed in SPSS 19.0 [33]. Canoco for Windows 4.5 was used for the canonical correspondence analysis (CCA). Microbial AWCD values were relatively stable throughout cultivation (0–120 h), so the72-h cultivation data were used for analysis [32] of carbon resource use. Data are presented as mean ± standard deviation. Significance was set at P < 0.05.
3 Results and discussion
3.1 Soil microbial community functional diversity
3.1.1 Spatial differentiation of microbial community diversity
Scirpus mariqueter soil microbial communities differed significantly (P < 0.05) in their H′, U, and D values across coverage levels, in the following order: dense > sparse > bare tidal flat (Table 1). Thus, their functional diversity was far higher in areas with dense S. mariqueter cover than in bare tidal flats.
3.1.2 Functional diversity of soil microbial communities based on their use of single and multiple carbon resources
The ability of soil microbial community to use single carbon sources is an important indicator of their metabolic activity [16]. AWCD changes over time showed a normal growth curve with time, including the color change, color change index, and early stage final stable period (or maximum level). The change of AWCD value of each sample with time can be used to indicate the average activity of microorganisms, which is used to reflect the reaction speed and the final extent of the microbial community [15]. We found that AWCD values of samples from all three coverage categories gradually increased with culture time, before stabilizing (Fig. 2). This outcome can be explained by the fact that microbial growth logarithmically increases when encountering a new culture environment, but when carbon resources are exhausted (in this experiment, by 72 h), microbial growth will eventually stop [34].
The three regions of the AWCD values are gradually increased with the culture time enter the logarithmic growth phase to 72 h and then changes gently and gradually stabilized. The results showed that the utilization rate of soil microbial biomass was significantly higher than that of sparse area than bare mudflat, and the difference of the three regions remained stable. The results showed that the microbial activity of soil was higher, and the microbial abundance was higher in dense area. One-way analysis of variance was used in SPSS(19.0) to show significant differences among the three area of samples (P < 0.05)
AWCD in dense vegetation area was bigger than other areas; AWCD in bare tidal flats was increased after a 72 h culture and was lower than the other two coverage categories. Overall, the data showed that soil microbe carbon use rates (and thus, biomass) were higher in dense areas than in sparse areas or bare tidal flats, and this difference remained stable over time.
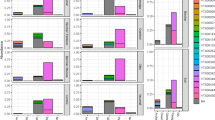
Microbial use rates of six carbon resource types differed across the three coverage categories (Fig. 3), in the following order: dense > sparse > bare tidal flat. Generally, the carbon hydrate had the highest utilization in all the soils. Sugars, alcohols, and acids were used more in dense areas, while sugars, esters, and acids were higher in bare tidal flats. Together, these data suggest that the same species of soil microorganisms were present across the three coverage types. Moreover, sugars and acids appear to be the main carbon resources for their growth.
There are obvious differences in the utilization rates of six kinds of carbon sources in three different areas of S. mariqueter. The dense area > sparse area > bare mudflat, the utilization rate of the same area microorganism are also different. The utilization rate of sugars, alcohols, and acids was higher in the dense area, and the utilization rate of sugars, esters, and acids was higher in the bare mudflat, indicating that the three regions soil microorganisms are basically the same species, and the sugars and acids are the main carbon sources for the growth of soil microorganisms. Tukey’s test was adopted to obtain the above results. Different letters on the figure represent significant differences (P < 0.05)
The utilization of 31 carbon sources is analyzed by principal component analysis [35, 36]. The eigenvalue, variance contribution rate, and cumulative variance contribution rate of principal components were shown in Table 2, and the load values of the two principal components of the 31 carbon sources are shown in Table 3. The first principal component (PC1) explained 76.307% of the variation in carbon use, which is mainly influenced by 25 substrates (five sugars, six amino acids, five acids);while the second principal component (PC2) explained 23.693%, which is mainly influenced by 5 substrates (two monosaccharides, one amino acid, one acid) (Table 3).
Substrates with high load values in the two axes reflect differences and similarities in microbial carbon source use across rhizosphere soils. Thus, most carbons used by microbes were sugar, amino acids, and acids. Furthermore, it is clear that soil microbial communities in areas with dense S. mariqueter exhibited greater biomass, diversity, uniformity, and dominance than microbes in bare tidal flats.
Coordinate mapping of PC1 and PC2 showed significant differences across coverage categories. On the PC1 axis, dense and sparse areas are distributed in the positive direction, while bare tidal flat is distributed in the negative direction (Fig. 4), indicating that dense and sparse areas had an absolute advantage in the utilization of 25 substrates (five sugars, six amino acids, five acids). On the PC2 axis, only dense areas are distributed in the positive direction, while bare tidal flat areas remain close to 0, indicated that dense area had an absolute advantage in the utilization of five substrates (two monosaccharides, one amino acid, one acid). Thus, we conclude that the soil microbial community of S. mariqueter has strong carbon source utilization in densely covered areas [37].
The first two principal components were selected for analysis. PC1 was used as the horizontal axis and PC2 as the vertical axis, and with different regions in the main components of the score for the coordinate mapping. There are significant differences in the PC axis in different areas of the beach. On the PC1 axis, the dense area and the sparse area are distributed in the positive direction. The bare mudflat is distributed in the negative direction. On the PC2 axis, dense area is distributed in the positive direction, and rare area is distributed in the negative direction. The soil microbial community of S. mariqueter has strong carbon source utilization in dense area
3.1.3 Physiochemical characteristics influencing soil microorganisms
Soil characteristics differed across coverage categories, with conductivity, particle size, and organic carbon exhibiting greater larger variance (Table 4).
Soil salinity was significantly correlated the Shannon diversity index (P < 0.01) (Table 5), indicating that salinity exerts a strong effect on microbial activity and community diversity. This relationship may be due to the impact of salt on microbial cells. Soil salinity determines osmotic pressure during microbial cell growth; if salinity is too low, the influx of water into microbial cells results in cell death, whereas excessive salinity also leads to cell death through water loss. The maintenance of appropriate osmotic pressure is a major reason that cultivation of coastal microorganisms uses potassium chloride solution.
Soil particle size was negatively correlated with all three diversity indices (P < 0.01). Therefore, soil particle size was also a major factor affecting soil microbial diversity in S. mariqueter soil. As S. mariqueter coverage decreased, soil microbial diversity decreased while particle size gradually increased. This pattern is likely associated with the scouring action of the tides. In areas with dense S. mariqueter cover, root systems are well developed; weakening the tide’s scouring force and elevating sediment deposition. These factors then improved soil structure, increasing water, and nutrient retention of the soil, thus providing a suitable environment for microbe growth. In contrast, bare tidal flats discourage sediment deposition, meaning only larger sediment particles are left behind, leading to larger soil pores [9] and more evaporation. Such conditions are more inhospitable to soil microorganisms. Overall, our results demonstrate the clear importance of soil organic carbon and total phosphorus to soil microbial diversity (P < 0.05), in line with previous studies [38].
At present, few studies are available on how soil microbial community structure could be used to evaluate S. mariqueter growth distribution. Soil nutrients and microbial diversity of tea trees at different ages were studied by single sampling [39]. Carbon source metabolic activity of Karst plateau wetland in different stages of vegetation succession in the rhizosphere soil microorganism was studied and found that environmental factors have important impacts on microbial metabolic activity with one time sampling [40];the single carbon source utilization of soil microbial community under different land use types in Poyang Lake wetland was determined by single sampling and found that the soil microbial community structure was changed by reclamation [41]. Here, we provided one sampling data that clarified the soil-related factors associated with changes to S. mariqueter soil microbial communities.
By the results of factors analysis influencing S. mariqueter soil microbial communities, technology of micro-topography manipulation should be the heart work of S. mariqueter wetlands remediation. The change of soil physical and chemical properties (salinity, soil water content, and total phosphorus) through geomorphic regulation could be developed to change soil microbial community diversity and then promoting the frequent occurrence of geochemical cycle and the accumulation of soil nutrients, and lastly promoting the survival of S. mariqueter population.
Biolog microplate technology is efficient and easy to operate. However, this technique also has some limitations. Biolog microplate technique can reflect the information mainly on the microbial carbon metabolism and functional diversity, limited in indicating microbial community structure and diversity, so further research would be performed to identify the gene structure and species using molecular biology technology such as high-throughput sequencing technology, etc. This study is a pioneer research which proved that microbe community diversity has a significant relation with Scirpus spatial heterogeneity. How using microbe to promote coastal Scirpus wetlands remediation is the heart issue we want to do in the near future.
4 Conclusions
Carbon source utilization rate and functional diversity were significantly higher in areas of dense S. mariqueter coverage than in sparsely covered areas and bare tidal flats (no coverage). In the utilization of carbon sources, carbohydrates, amino acids, and acids are the main carbon sources of S. mariqueter soil microorganisms, while the utilization rate of amine carbon source is the lowest.
The main factors affecting soil microbial diversity were salinity, particle size, organic carbon, and soil moisture. The fact that S. mariqueter growth was primarily influenced by elevation, salinity, particle size, and total phosphorus implies that these factors are critical for S. mariqueter remediation.
References
Sun C, Liu Y, Zhao S, Li H, Sun J (2016) Salt marshes response to human activities on a prograding coast revealed by a dual-scale time-series strategy. Estuar Coasts 40(2):1–18
Sun S, Cai Y, Tian X (2003) Salt marsh vegetation change after a short-term tidal restriction in the Changjiang Estuary. Wetlands 23(2):257–266
Sun S, Cai Y, An S (2002) Differences in morphology and biomass allocation of Scirpus mariqueter between creekside and inland communities in the Changjiang Estuary, China. Wetlands 22(4):786–793
Wang CH, Lu M, Yang B, Yang Q, Zhang XD, Toshihiko H, Li B (2010) Effects of environmental gradients on the performances of four dominant plants in a Chinese saltmarsh: implications for plant zonation. Ecol Res 25(2):347–358
Li J, Wei GX, Ji HY, Wu Y, Yu K (2014) Analysis on promotion methods of large-scale deposition promotion projects in shallow tidal-lands in Yangtze River Estuary. Yangtze River 45(7):61–65
Chen ZY (2004) Local competitive effects of introduced Spartina alterniflora on Scirpusmariqueter at Dongtan of Chongming Island, the Yangtze River estuary and their potential ecological consequences. Hydrobiologia 528(1–3):99–106
Zhang Y, Cui B, Tian X, Wang Q, Yan J (2016) Gradient distribution patterns of rhizosphere bacteria associated with the coastal reclamation. Wetlands 36(1):69–80
Fu QL, Liu C, Ding NF, Lin YC, Guo B, Luo JF, Wang HL (2012) Soil microbial communities and enzyme activities in a reclaimed coastal soil chronosequence under rice–barley cropping. J Soils Sediments 12(7):1134–1144
He YL, Li XZ, Craft C, Ma ZG, Sun YG (2011) Relationships between vegetation zonation and environmental factors in newly formed tidal marshes of the Yangtze River Estuary. Wetl Ecol Manag 19(4):341–349
Levine JM, Brewer JS, Bertness MD (1998) Nutrients, competition and plant zonation in a New England salt marsh. J Ecol 86(2):285–292
Sun YG, Li XZ, Mander Ü, He YL, Jia Y, Ma ZG, Guo WY, Xin ZJ (2011) Effect of reclamation time and land use on soil properties in Changjiang River Estuary, China. Chin Geogr Sci 21(4):403
Zeller V, Bardgett RD, Tappeiner U (2001) Site and management effects on soil microbial properties of subalpine meadows: a study of land abandonment along a north–south gradient in the European Alps. Soil Biol Biochem 33(4):639–649
Böhme L, Langer U, Böhme F (2005) Microbial biomass, enzyme activities and microbial community structure in two European long-term field experiments. Agric Ecosyst Environ 109(1):141–152
Garland JL (1997) Analysis and interpretation of community-level physiological profiles in microbial ecology. FEMS Microbiol Ecol 24(4):289–300
Tian YN, Wang HQ (2011) Application of Biolog to study of environmental microbial function diversity. Environ Sci Technol 34(3):50–57
Zhang H, Huang T, Chen S, Lin G, Liu T, Xiao Y (2014) Microbial community functional diversity and enzymatic activity in the sediments of drinking water reservoirs, Northwest China. Desalin Water Treat 52(7–9):1608–1614
Zheng H, Yun OZ, Fang ZG (2004) Application of BIOLOG to study on soil microbial community functional diversity. Acta PedologicaSinica 41(3):456–461
Yan FJ, Tian XL, Dong SL, Yang G, Liu RJ, Zhang K (2014) Seasonal variation of functional diversity of microbial communities in sediment and shelter of sea cucumber (Apostichopus japonicus) cultural ponds. Acta EcologicaSinica 34(11):2996–3006
Lin X, Lin Z, Lin D, Lin H, Luo H, Hu Y, Lin C, Zhu C (2014) Effects of planting Pennisetum sp. (Giant Juncao) on soil microbial functional diversity and fertility in the barren hillside. Acta EcologicaSinica 34(15):4304–4312
Graham MH, Haynes RJ (2005) Catabolic diversity of soil microbial communities under sugarcane and other land uses estimated by Biolog and substrate-induced respiration methods. Appl Soil Ecol 29(2):155–164
Braun B, Böckelmann U, Grohmann E, Szewzyk U (2006) Polyphasic characterization of the bacterial community in an urban soil profile with in situ and culture-dependent methods. Appl Soil Ecol 31(3):267–279
Mei XY, Zhang XF (2007) Carbon storage and fixation function of Scirpus mariqueter in Changjiang River Estuary: a case study of Chongming Dongtan Wetland. J Agro Environ Sci 1:360–363
Xu ZG, Yan BX, Yan HE, Song CC (2007) Analysis of soil environmental factors of wetland plants of Sanjiang Plain. China Environ Sci 32(35):8381–8382
Tian B, Zhang LQ, Wang XR, Zhou YX, Zhang W (2010) Forecasting the effects of sea-level rise at Chongming Dongtan nature reserve in the Yangtze Delta, Shanghai, China. Ecol Eng 36(10):1383–1388
Zhang YB, Deng AY, Zhuang TC (2003) Relation between soil salinity in intertidal zone and electric conductivity. Ecol Environ Sci 12(2):164–165
Bao SD (2000) Soil and agricultural chemistry analysis. China Agriculture Press, Beijing, pp 263–270
Stefanowicz A (2006) The Biolog plates technique as a tool in ecological studies of microbial communities. Pol J Environ Stud 15(5):669–676
Juliet P, Boddy L, Peter FR (2002) Analysis of microbial community functional diversity using sole-carbon-source utilisation profiles—a critique. FEMS Microbiol Ecol 42(1):1–14
Choi KH, Dobbs FC (1999) Comparison of two kinds of Biolog microplates (GN and ECO) in their ability to distinguish among aquatic microbial communities. J Microbiol Methods 36(3):203–213
Smalla K, Wachtendorf U, Heuer H, Liu WT, Forney L (1998) Analysis of BIOLOG GN substrate utilization patterns by microbial communities. Appl Environ Microbiol 64(4):1220–1225
Liu B, Li YX, Zhang XL, Wang J, Gao M (2015) Effects of chlortetracycline on soil microbial communities: comparisons of enzyme activities to the functional diversity via BiologEcoPlates™. Eur J Soil Biol 68:69–76
Tam L, Derry AM, Kevan PG, Trevors JT (2001) Functional diversity and community structure of microorganisms in rhizosphere and non-rhizosphere Canadian arctic soils. Biodivers Conserv 10(11):1933–1947
Valeri L, Vanderweele TJ (2013) Mediation analysis allowing for exposure-mediator interactions and causal interpretation: theoretical assumptions and implementation with SAS and SPSS macros. Psychol Methods 18(2):137–150
Yao H, He Z, Wilson MJ, Campbell CD (2000) Microbial biomass and community structure in a sequence of soils with increasing fertility and changing land use. Microb Ecol 40(3):223–237
Hong Q, Zhuang P, Yang G, Zhang T, Hou J, Zhang L, Zhi Y, Zhang H, Yang Y (2012) Feeding habits of Lateolabrax maculatus in Yangtze River Estuary. Acta EcologicaSinica 32(13):4181–4190
Yang G, Hong QQ, Zhang T, Hou JL, Zhi YL, Zhuang P, Zhang LZ (2012) Fish community structure in intertidal area of Acipensersinensis natural reserve in Yangtze River Estuary. Chin J Ecol 31(5):1194–1201
Jones DG, Beston BR, Murphy KM (2007) Novel application of principal component analysis to understanding visual cortical development. BMC Neurosci 8(Suppl 2):P188
Nannipieri P, Ascher J, Ceccherini MT, Landi L (2010) Microbial diversity and soil functions. Eur J Soil Sci 54(4):655–670
Lin S, Zhang JQ, Chen T, Zhang AJ, Zhou MM, Lin WX (2012) Analysis of nutrient and microbial Biolog function diversity in tea soils with different planting years in Fujian Anxi. Chin J Eco Agric 20(11):1471–1477
Xia PH, Koi YZ, Yu LF (2015) Carbon metabolic soil microbial community in Caohai Karst Plateau degraded wetland: a case study in southwest China. Acta Sci Circumst 35(8):2549–2555
Zhang J, Hu W, Liu YZ, Ge G, Wu L (2015) Response of soil microbial functional diversity to different land-use types in wetland of Poyang Lake, China. Acta EcologicaSinica 35(4):965–971
Acknowledgements
This work was supported by the National Basic Research Program of China (Grant Number 2013CB430400), Shanghai Oceanography Bureau HHK 2015-02, and National Key R&D Program of China (Grant Number 2017YFC0506002).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There are no conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhao, M., Yin, C., Tao, Y. et al. Diversity of soil microbial community identified by Biolog method and the associated soil characteristics on reclaimed Scirpus mariqueter wetlands. SN Appl. Sci. 1, 1408 (2019). https://doi.org/10.1007/s42452-019-1443-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-019-1443-y