Abstract
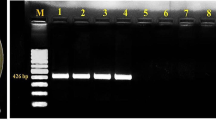
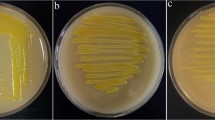
Bacterial panicle blight (BPB) of rice (Oryza sativa) caused by Burkholderia glumae (BG) is an emerging disease of rice that substantially limits rice productivity. It has become a major concern in basmati-growing regions of North-western India. In the present study, 10 BG strains were studied for genetic and pathogenic diversity. The colonies were non-spore forming, rod shaped with rounded ends; looked gray, yellow, and brisk yellow on PSA, King’s B, and SPG agar plate, respectively. Rep-PCR DNA fingerprint analysis exhibited different phyletic grouping, viz., five in BOX-PCR, two in ERIC-PCR, and one in rep-PCR. The phyletic group 1 of BOX PCR consisted of three isolates BG1, BG2, and BG3 and phyletic group 2 had three isolates, viz., BG4, BG8, and BG10, from the same agroclimatic condition along with high similarity matrix. The first phyletic group of ERIC PCR exhibited two isolates BG1 and BG10 from Uttar Pradesh (UP); however, grouping of isolates irrespective of agroclimatic condition was observed in phyletic group 2. The strains BG5 and BG6 (Delhi) that grouped together in phyletic group 3 in the BOX analysis also grouped together in the ERIC analysis. The BOX-PCR formed greater number of phyletics as well as amplicons and the difference in Jaccard’s similarity matrix was more than ERIC- and rep-PCR. The phylogenetic analysis of gyrB gene showed simultaneous grouping of eight isolates from UP in one clade and two isolates from Delhi in another clade. Based on pathogenicity assays, strains varied in their ability to produce variable disease scores when artificially inoculated onto the susceptible rice plants. The strains BG1 and BG3 were highly virulent while strains BG7 and BG10 were of least virulent types. Altogether, the present study brings an overview on the variability of BG strains from India and this detailed insight would help the breeders for disease-resistance screening.







Similar content being viewed by others
Data availability
The manuscript has data included as electronic supplementary material.
References
Baghaee-Ravari S, Rahimian H, Shams-Bakhsh M, Lopez Solanilla E, Antúnez-Lamas M, Rodríguez-Palenzuela P (2011) Characterization of Pectobacterium spp from Iran using biochemical and molecular methods. European Journal of Plant Pathology 129:413–425
Bassler BL (2001) Quorum sensing in bacteria. Annual Review of Microbiology 55(1):165–199
Brenner DJ, Krieg NR, Staley JT (2005) Bergey’s manual of systematic bacteriology, 2nd edn. Springer, New York, USA
Cha KH, Lee YH, Ko SJ, Park SK, Park IJ (2001) Influence of weather condition at heading period on the development of rice bacterial grain rot caused by Burkholderia glumae. Research in Plant Disease 7:150–154
Chalupowicz L, Zellermann EM, Fluegel M, Dror O, Eichenlaub R, Gartemann KH, Savidor A, Sessa G, Iraki N, Barash I, Manulis-Sasson S (2012) Colonization and movement of GFP-labeled Clavibacter michiganensis subsp. michiganensis during tomato infection. Phytopathology 102(1):23–31
Chen R, Barphagha IK, Karki HS, Ham JH (2012) Dissection of quorum-sensing genes in Burkholderiaglumae reveals non-canonical regulation and the new regulatory gene tofM for toxoflavin production. PLOS ONE 7(12):e52150
Cottyn B, Cerez MT, Van Outryve MF, Barroga J, Swings J, Mew TW (1996) Bacterial diseases of rice. I. Pathogenic bacteria associated with sheath rot complex and grain discoloration of rice in the Philippines. Plant Disease 80(4):429–437
Cui ZB, Xie G, Li B, Huang S (2016) Research status and prospect of Burkholderia glumae, the pathogen causing bacterial panicle blight. Rice Science 23:111–118
Dickey RS (1979) Erwinia chrysanthemi: A comparative study of phenotypic properties of strains from several hosts and other Erwinia species. Phytopathology 69:324–329
Fory PA, Triplett L, Ballen C, Abello JF, Duitama J, Aricapa MG, Prado GA, Correa F, Hamilton J, Leach JE, Tohme J, Mosquera GM (2014) Comparative analysis of two emerging rice bacterial pathogens. Phytopathology 104:436–444
Fukagawa NK, Ziska LH (2019) Rice: importance for global nutrition. Journal of Nutritional Science and Vitaminology 65:S2–S3
Goto M (1992) Fundamentals of bacterial plant pathology. Academic Press, San Diego, CA
Goto K, Ohata K (1956) New bacterial diseases of rice (brown stripe and grain rot). Annals of the Phytopathological Society of Japan 21:46–47
Goto M, Takikawa Y (1984) Methods for identification of plant pathogenic bacteria. Plant Protection 38:339–344
Gowda HRA, Tripathi R, Tewari R, Vishunavat K (2022) Morphological and molecular characterization of Burkholderiaglumae causing panicle blight of paddy. Physiological and Molecular Plant Pathology 117:101755
Hikichi Y, Okuno K, Furusawa I (1993) Immunofluorescent antibody technique for detecting Pseudomonas glumae on rice plants. Annals of the Phytopathological Society of Japan 59:477–480
Hu FP, Young JM, Triggs CM, Park DC, Saul DJ (2001) Relationships within the Proteobacteria of plant pathogenic Acidovorax species and subspecies, Burkholderia species, and Herbaspirillum rubrisubalbicans by sequence analysis of 16S rDNA, numerical analysis and determinative tests. Antonie Van Leeuwenhoek 80:201–213
Iiyama K, Furuya N, Hara K, Nakashima N, Takanami Y, Matsuyama N (1994) Phytotoxic produced by Pseudomonas glumae Kurita et Tabei, a causal bacterium of the grain and seedling rot of rice [Oryza sativa]. Journal of the Faculty of Agriculture Kyushu University 38:175–181
Jeong Y, Kim J, Kim S, Kang Y, Nagamatsu T, Hwang I (2003) Toxoflavin produced by Burkholderia glumae causing rice grain rot is responsible for inducing bacterial wilt in many field crops. Plant Disease 87(8):890–895
Jurtshuk P Jr, McQuitty DN (1976) Use of a quantitative oxidase test for characterizing oxidative metabolism in bacteria. Applied and Environmental Microbiology 31(5):668–679
Karki HS, Shrestha BK, Han JW, Groth DE, Barphagha IK, Rush MC, Melanson RA, Kim BS, Ham JH (2012) Diversities in virulence, antifungal activity, pigmentation and DNA fingerprinting among strains of B. glumae. PLoS One 7:45376
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution 35:1547–1549
Louws FJ, Fulbright D, Stephens C, Bruijn FD (1994) Specific genomic fingerprints of phytopathogenic Xanthomonas and Pseudomonas pathovars and strains generated with repetitive sequences and PCR. Applied and Environmental Microbiology 60:2286–2295
Louws FJ, Rademaker JLW, de Bruijn FJ (1999) The three Ds of PCR-based genomic analysis of phytobacteria: diversity, detection, and disease diagnosis. Annual Review of Phytopathology 37:81–125
Maeda Y, Shinohara H, Kiba A, Ohnishi K, Furuya N, Kawamura Y, Ezaki T, Vandamme P, Tsushima S, Hikichi Y (2006) Phylogenetic study and multiplex PCR-based detection of Burkholderia plantarii, Burkholderia glumae and Burkholderia gladioli using gyrB and rpoD sequences. International Journal of Systematic and Evolutionary Microbiology 56:1031–1038
Maeda Y, Horita M, Shinohara H (2007) Analysis of sources of oxolinic acid-resistant field strains of Burkholderia glumae based on rep-PCR analysis and nucleotide sequences of gyrB and rpoD. Journal of General Plant Pathology 73:46–52
Mondal KK, Mani C, Verma G (2015) Emergence of bacterial panicle blight caused by Burkholderia glumae in North India. Plant Disease 99(9):1268–1268
Mulaw T, Wamishe Y, Jia Y (2018) Characterization and in plant detection of bacteria that cause bacterial panicle blight of rice. American Journal of Plant Sciences 9:667–684
Murray M, Thompson W (1980) Rapid isolation of higher weight DNA. Nucleic Acids Research 8:4321–4325
Nandakumar R, Shahjahan AKM, Yuan XL, Dickstein ER, Groth DE, Clark CA, Cartwright RD, Rush MC (2009) Burkholderia glumae and B. gladioli cause bacterial panicle blight in rice in the southern United States. Plant Disease 93:896–905
Pet-amphai W, Watcharachaiyakup J, Patarapuwadol S, Kositratana W (2017) Identification of bacterial pathogens causing panicle blight and dirty panicle of rice by multilocus sequence analysis. Journal of Agricultural Science 48:297–311
Prakash O, Verma M, Sharma P, Kumar M, Kumari K, Singh A, Kumari H, Jit S, Khanna SK, Gupta M (2007) Polyphasic approach of bacterial classification—an overview of recent advances. Indian Journal of Microbiology 47:98–108
Ramachandran K, Vijaya SI, Ahmad FN (2021) Characterization and identification of Burkholderia glumae as the causal pathogen of bacterial panicle blight of rice (Oryza sativa L.) in Malaysian rice granaries. Journal of General Plant Pathology 87:164–169
Saichuk J (ed) (2009) Louisiana rice production handbook. Baton Rouge, LA, USA: LSU Agricultural Center Pub. 2321 (3M) 6/09 Rev
Salles JF, De Souza FA, van Elsas JD (2002) Molecular method to assess the diversity of Burkholderia species in environmental samples. Applied and Environmental Microbiology 68:1595–1603
Sayler RJ, Cartwright RD, Yang Y (2006) Genetic characterization and real-time PCR detection of Burkholderia glumae, a newly emerging bacterial pathogen of rice in the United States. Plant Disease 90(5):603–610
Schaad NW, Jones JB, Lacy GH (2001) Laboratory guide for identification of plant-pathogenic bacteria. APS Press, St Paul, MN, USA
Shew AM, Durand-Morat A, Nalley LL, Zhou XG, Rojas C, Thoma G (2019) Warming increases bacterial panicle blight (Burkholderia glumae) occurrences and impacts on USA rice production. PLoS ONE 14:e0219199
Singh D, Vishunavat K (2015) Identification of a seed-borne rice bacterium, Burkholderia glumae using cultural, morphological and biochemical methods. Journal of Applied and Natural Science 7:562–566
Streeter JG (2007) Factors affecting the survival of Bradyrhizobium applied in liquid cultures to soya bean [Glycine max (L.) Merr.] seeds. Journal of Applied Microbiology 103:1282–1290.
Takeuchi T, Sawada H, Suzuki F, Matsuda I (1997) Specific detection for Burkholderia plantarii and B. glumae by PCR using primers selected from the 16S–23S rDNA spacer regions. Annals of the Phytopathological Society of Japan 63:455–462
Tamura K, Peterson D, Peterson N, Stecher G, Nei M (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution 28:2731–2739
Tsushima S (1996) Epidemiology of bacterial grain rot of rice caused by Pseudomonas glumae. Japan Agricultural Research Quarterly 30:85–89
Tsushima S, Wakimoto S, Mogi S (1986) Selective medium for detecting Pseudomonas glumae Kurita et Tabei, the causal bacterium of grain rot of rice. Japanese Journal of Phytopathology 52:253–259
Uematsu T, Yoshimura D, Nishiyama K, Ibaraki T, Fuji H (1976) Occurrence of bacterial seedling rot in nursery flat, caused by grain rot bacterium Pseudomonas glumae. Annals of the Phytopathological Society of Japan 42:310–312
Yabuuchi E, Kosako Y, Oyaizu H, Yano I, Hotta H, Hashimoto Y, Ezaki T, Arakawa M (1992) Proposal of Burkholderia genera and transfer of seven species of the genus Pseudomonas homology group II to the new genus, with the type species Burkholderia cepacia (Palleroni and Holmes 1981). Microbiology and Immunology 36(12):1251–1275
Yamamoto S, Harayama S (1998) Phylogenetic relationships of Pseudomonas putida strains deduced from the nucleotide sequences of gyrB, rpoD and 16S rRNA genes. International Journal of Systematic Bacteriology 48:813–819
Yoneyama K, Kono Y, Yamaguchi I, Horikoshi M, Hirooka T (1998) Toxoflavin is an essential factor for virulence of Burkholderia glumae causing rice seedling rot disease Japanese. Japanese Journal of Phytopathology 64:91–96
Yuan XL (2005) Identification of bacterial pathogens causing a blight. Master thesis, Louisiana State University. https://doi.org/10.31390/gradschool_theses.2883
Zhou XG (2019) Sustainable strategies for managing bacterial panicle blight in rice. In: Yulin J (ed), Protecting rice grains in the post-genomic era. Intech Open, London. https://doi.org/10.5772/intechopen.84882.
Acknowledgements
The authors are grateful to the Plant Bacteriology laboratory, Division of Plant Pathology, Indian Agricultural Research Institute, New Delhi for providing all the necessary infrastructure and materials so that the study could be conducted. No external financial assistance was provided for the study.
Author information
Authors and Affiliations
Contributions
KKM: conceptualized the research, supervised, analyzed the data, and edited the manuscript. SK, ALM: associated with data analysis, writing of draft manuscript. AK: primer designing. SK: performed research activities including BG isolation and virulence assay. AK, SB, ALM, MS, RER, KNS, SK: amplification of BOX, ERIC, and REP primers. CM, TG, AK: maintained the BG strains, helped in raising the rice crops and in inoculation.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kumar, S., Mondal, K.K., Ghoshal, T. et al. Genetic and pathogenic diversity analysis of Burkholderia glumae strains from Indian hot spot regions causing bacterial panicle blight of rice (Oryza sativa L.). Trop. plant pathol. 48, 139–153 (2023). https://doi.org/10.1007/s40858-023-00554-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40858-023-00554-z