Abstract
Molecular characterization was carried out on samples of historical grapevine populations that were gathered from within and around the medieval walls of Siena. Forty-nine grapevines were selected based on their age, historical site of growth, grapevines’ ampelography, and for being relict accessions, obsolete to cultivation. SSR profiling data were compared to 44 known grapevines, revealing six functional genetic groups with significant similarity to grapevine types generally grown in Tuscany. The Sienese germplasm is enriched with rare grapevines at risk of extinction, such as Zuccaccio, Gorgottesco, Tenerone, Prugnolo gentile, Occhio di Pernice, Procanico, Rossone, Mammolo, and Canina. Population genetics analysis revealed the existence of five subpopulations structure (-k5) in analogy with cluster analysis.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The importance of characterizing the traditional grapevine germplasm was recently reaffirmed by the EU directives (prm_leg_future_prm-study_swd-2021-90(3).pdf). The study aimed to enhance the indigenous viticultural heritage and the historical forms of cultivation of the grapevines in the walled city of Siena (Tuscany, Italy) and its suburban spaces. Siena is a privileged setting since it has preserved small areas of vineyard landscapes of historical origin. The "land units” are generally limited green spaces, such as cloistered convents or private gardens, characterized by the presence of historical grapevines of considerable age.
The implementation of genetic breeding programs requires the maintenance of a high degree of genetic diversity in the grapevine germplasm that should, hopefully, include resilient individuals that had survived over the years with no intensive care by man. Local, resilient grapevines represent a potential added value for the economy and tourism as they present a low environmental impact in favor of environmental and agricultural sustainability.
Molecular markers have been widely used for studies of the population structure, demographic history, dynamics, and the evolution of plants’ genetic populations [1,2,3,4]. The recovery of the ancient varieties is a recent trend, promoted especially by local communities and governmental entities to contrast the erosion of grapevine genetic diversity. In this view, many researchers nowadays focus on recovering abandoned, regional, relict varieties [5,6,7]. The local sites in Siena appear to be of extraordinary historic importance, even if they are only a small proportion of the numerous locations in Tuscany hosting local, uncharacterized ancient grapevine strains which deserve to be studied and protected.
Within a multidisciplinary study involving the prospection of local, historical vineyards in Siena and its peri-urban spaces, here, we demonstrate genetic evidence of the existence of rare grapevines at risk of extinction, branching from functional groups characterized by predominant similarities with known main grapevine strains mainly found in Tuscany.
Material and Methods
Plant Material
Ninety-three accessions from the Vitis spp. were analyzed; 49 from different sites within the medieval walls of Siena or in the immediate surroundings of the Town (Fig. 1), namely: Strada di Istieto (IS), Strada Certosa di Maggiano (CM), Strada Cassia Sud (CS), Istituto San Girolamo (ISG), Orto de’ Pecci (OdP), Porta San Marco (PSM), Convento San Domenico (CSD), Strada di Busseto—Podere Ponticini (PP), Strada del Linaiolo (SLI), Via Aretina (VAR), Via Peruzzi (BP), Belriguardo (BLR), Palazzo Sergardi Biringucci (PSE), Strada di Ventena (VE).
Grapevines used as varietal references for genetic comparison derive from the Italian National Varietal Register (Table 1).
DNA Extraction and Genotyping
DNA extraction and genotyping at SSR loci: VVMD21 [8], VVMD31, VVMD36, VrZAG47, VVMD25, VVMD27 [9], VVS2 [10], VVMD7 [11] were carried out according to Vignani et al. [12].
Cluster analysis of the grapevines was performed by the UPGMA (unweighted pair group arithmetic means average) method using the SAHN subprogram of NTSYS 2.0 software (Exeter Software, East Setauket, NY).
Genetic Diversity Analyses
GenAlex v. 6.5 was used for descriptive statistics: alleles per locus (Na), the effective number of alleles per locus (Ne), allele frequency, any Wahlund effect, observed heterozygosity (HO), expected heterozygosity (HE), Fixation index (F), Principal Coordinates Analysis (PCoA), private alleles (Pa), [13, 14], and the hierarchical analysis of molecular variance (AMOVA).
Population Structure Analysis
STRUCTURE v. 2.3.4 was used for population analysis [15]. All simulations were performed with 100,000 replicates for burn-in and 1,000,000 replicates for Markov chain Monte Carlo (MCMC) processes in 10 independent runs. The number of K clusters was determined firstly by simulating a range of K-values from 1 to 10, without putative population origin for each individual. Since after K = 2, there were no other appreciable peaks, STRUCTURE was run again with 10 runs for K-values ranging from 1 to 11, with putative population origin for each individual following the 6 functional genetic groups identified by the dendrogram of similarity. The STRUCTURE outputs were analyzed by Structure Harvester [16] and CLUMPAK [17]. Principal coordinate analysis (PCoA) was run via a Covariance matrix with data standardization to infer the distribution of grapevine accessions among structure groups.
Results and Discussion
Cluster Analysis
Grapevines were selected based on their unusual phenotypic traits that seemed to deviate from the standard morphology of main Tuscan grapevine types according to OIV ampelographic determination (data not shown).
The genetic variability of the Sienese grapevines is graphically represented in Fig. 2. A threshold of significance seems to be deductible in agreement with ampelography observations done on the same plants. Only five grapevines were highly related genetically to known references (> 90%), while the majority ranged from 40 to 70% similarity with known grapevines. The high degree of grapevine diversity may be explained by considering the “land units” have acted as "time capsules" for grapevine diversity conservation.
Six main similarity groups were detected. Namely:
-
(i)
The homologs of white-berry Tuscan grapes Procanico, Ansonica d’Arcille, and Ansonica were used as controls for identity. There was only one vine from the Northern area—42 BP—while the remaining—40 SLI, 22 ISG, 35 CSD, and 29 PSM—were from the town of Siena or its Southern peri-urban area. These are white-berry grapevines genetically homolog to Trebbiano, considered an endemic variety in Tuscany;
-
(ii)
The homologs of Sangiovese, including Prugnolo Gentile and Sangiovese Piccolo Precoce, were used as controls, ranging from a minimum of 80% identifying with Sangiovese (Sangiovese numbers 9, 73, and 75 in Table 1). 30 PSM is related to Prugnolo Gentile and Sangiovese Piccolo Precoce, 5 IS is a Sangiovese, and 23 ISG, 28 OdP are about 85% similar to Sangiovese. 26 OdP is very close to Rossone, while 2 IS, 3 IS, 15 CS were found in this group close to Mammolo (Mammolo 2ISV, number 82 in Table 1). Only three grapevines from the Northern area of Siena were found in this group: 44 BLR, 46 PSE, and 49 VE. 46 PSE is an ancient grapevine over 100 years old; it was taken from an inner garden located at Sergardi Biringucci Palace, a noble residence built in the eighteenth century (1744) along the path of the Via Francigena. 23 ISG was isolated from a convent (Istituto di San Girolamo), and 5 IS, 21 IS, 31 IS, 15 CS and 26 OdP were from the Southern area of Siena;
-
(iii) and (iv)
Heterogeneous, traditional Tuscan grapevines that include some red-berry aromatic grapevines as controls, such as Occhio di Pernice, Moscatello Nero, Malvasia nera, and several aromatic white-berry grapevines such as Moscato Bianco di Lucca, and Malvasia Bianca. Only two accessions from the Northern suburban area, 45 BLR and 47 VE, cluster with low genetic similarity with the standards in this group. Two accessions from the Northern suburban area, 45 BLR and 47 VE, cluster with low genetic similarity to the standards in this group. 24 ISG found at the Convent of San Girolamo (Siena) is highly related to Occhio di Pernice which is still used for Vin Santo, a sweet wine used for religious celebrations. 16 CS found along the Francigena road is genetically identical to Tenerone. 25 ODP and 34 CSD are > 90% related to 14 CM, where CM refers to Certosa di Maggiano, an ancient charterhouse founded in 1314.
-
(v) and (vi)
The most important details are that 33 CSD, 13 CM, and 9 CM are 80% genetically related to the red-berry grapevine Gorgottesco, 36 PP branches with 60% homology with the Gorgottesco, Zuccaccio, and Canaiolo Nero, and 12 CM is 77% genetically similar to the "Canina". 32 PSM and 38 PP are close to Giacchè, while 43 BP appears unrelated to any control. The Giacchè shows some rare alleles (141 bp at the VVS2 locus, 270 bp at VVMD36, and 238 bp at VVMD21), and its biological origin and correlation with non-vinifera and color-releasing grapes remain uncertain.
In the (v) and (vi) groups, we found accessions that were genetically and morphologically similar to American non-vinifera grapevines. These might be natural hybrids from V. vinifera x non-vinifera spp. widely used as rootstocks.
Interestingly, the identified accessions relate to grapevines that are listed in the Tuscan germplasm collection as rare grapevine types, such as Mammolo, Zuccaccio, Canina, Rossone, Salamanna, Gorgottesco, Occhio di Pernice and Tenerone, some of them being classified as varieties risking extinction. These traditional varieties have been neglected over the years due to several features, mainly related to scarce productivity that conversely was a priority trait during clonal selection starting from the eighties. The increment of grapevine biodiversity can be a strategic resource to face climate change, proving how the selection criterion has been modified over the years.
Population Structure
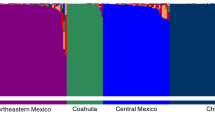
STRUCTURE analysis suggested K = 2 (Fig. 3) and K = 5 (Fig. 4) as the uppermost hierarchical levels of structure [20]. A membership coefficient (q-value) threshold of 0.7 for genetic sub-population assignment SSR-group 1 and SSR-group 2 was considered for K = 2. 48 (51.6%) and 40 (43%) genotypes were assigned to SSR-group 1 and SSR-group 2, respectively. The percentage of admixed genotypes was 5.4% corresponding to five genotypes: 45 BLR, 47 VE, 33 CSD, “Vaiano,” “Uva Vecchia Bianca” (Fig. 3). SSR-group 1 is composed of Italian varieties including mostly red-berry grapevines such as ‘Sangiovese’ and local grapevine strains that are widely distributed in Tuscany as major cultivars for PDO wine production, such as Prugnolo, Malvasia nera, Mammolo, Poverina, Rossone, Tenerone, and Pisciancio. Furthermore, among the white berry grapevines Ansonica, Procanico, Biancone, and Zuccaccio are found in SSR-group 1. SSR-group 2 includes “Color” grapevine types, such as members of the Colorino family, and control vines closely related to the non-Vinifera species like the “uva Americana” (V. Labrusca). Among the local strains are Gorgottesco, Abrusco/Abrostine, Occhio di Pernice, Canina, Pisciancione, and Giacchè.
a Best K as determined by calculating ln(K) and ΔK using STRUCTURE HARVESTER. Best K = 2. b Bayesian clustering results were obtained using STRUCTURE and c CLUMPAK clustering visualization of the Structure results at K = 2. The number in Bayesian clustering results (b) refers to the grapevine samples (Table 1)
According to K = 5 stratification, genotypes are divided into five major genetic groups and almost exactly overlap the functional groups in the dendrogram, except for groups v) and vi) that merged (Figs. 2, 4). The population stratification observed may be due to local agronomical practices such as clonal asexual propagation that allowed the conservation of traditional grapevines by fixing genetic traits in the respective strains.
The two-dimensional projections of PCoA analysis (Fig. 5) were plotted in a 2-D dimension scattered plot. PCoA1 and PCoA2 explained 11.76 and 9.42% of the total variation, respectively. SSR-groups 1 and SSR-group 2 members were partially discriminated by this analysis; along the PCoA1, the majority of Sienese grapevines were found in the SSR-group 1 with the varietal control that include the Sangiovese family. The highest molecular variation associated with the Sangiovese can be explained based on the wide diffusion of this variety in the region of Tuscany. Sangiovese, which is considered a “difficult” variety on an international basis, is well adapted to the climate of Tuscany and central Italian regions which explained the wide diffusion of this variety for the red wine-making industry of Chianti and other renowned DOC and DOCG wines from central Italy. Several local grapevine strains (Pisciancio, Tenerone, Malvasia, Canina, Canaiolo, Occhio di Pernice, Vaiano, Biancone, and Procanico) belonged to the SSR-group 2, together with the Colorino family, and Gorgottesco.
Along the PCoA2, the population showed a lower degree of differentiation, even if the Sangiovese and Colorino families are clustering together.
Genetic Diversity
The allelic frequencies (Fig. 6) distinguished the local vines from the varietal references. Some alleles, namely 135 at the VVS2 locus, 241 at the locus VVMD25, 246 at the VVMD7, 160 at the locus Zag47 and 183 at the locus VVMD27 approximately showed a doubled frequency in the Siena grapevines with respect to the references, a phenomenon that expresses a certain tendency to genetic fixation. In other words, among the 176 alleles observed overall in the two groups, some appeared more frequently in Siena vines compared to the total control group.
The number of different alleles (Na) for Siena grapevines was lower than the varietal references, 77 versus 99, while the values of alleles with a frequency of more than 5% were comparable between the two groups (37 and 40, respectively) (Table 2).
Several Private Alleles (No. Private Alleles) were associated selectively with each group, wherein the varietal references the No. was approximately doubled that in the Siena grapevines: mean values 5.5 and 2.75, respectively. The list of Private Alleles per locus in each accession is reported in the Supplementary material (S1). Analysis of molecular variance (AMOVA) for Siena grapevines and varietal accessions is reported in the supplementary material (S2). The average Expected Heterozygosity (He) shown in the auxiliary vertical axis varies between 0.66 (Siena vines) and 0.82 (varietal references). The diversity expressed by the I value, the Shannon index (a measure of the proportion of individuals belonging to the same genotype) is comparable for the Siena vines and the control (Fig. 7).
Allelic patterns across the two analyzed groups. Na = No. of Different Alleles; Na (Freq ≥ 5%) = No. of Different Alleles with a Frequency ≥ 5%; Ne = No. of Effective Alleles = 1/(Sum pi^2); I = Shannon's Information Index = − 1* Sum (pi * Ln (pi)); No. Private Alleles = No. of Alleles Unique to a Single Population; There aren’t common alleles with a frequency over 5% linked to at least 25% of grapevines taken from the meta-group (Siena grapevines, control grapevines)
The diversity indices calculated for distinct genotypes of historical Siena vines and control grapes are shown in Table 3. Most of the markers were highly polymorphic in the two groups. In the Siena grapevines, the observed number of effective alleles (Ne) differed from 1.942 (VVS2) to 7.309 (VVMD7), and an average number of 4.725 effective alleles was obtained for SSR markers over the two groups, and loci.
The observed heterozygosity (Ho) in the historical local grapevines varied between 0.479 (VVS2) and 0.918 (Zag49); the lowest expected heterozygosity (He) was detected at the VVS2 locus with 0.485, and the highest one at VVMD7 locus with 0.863. The varietal references showed values of Ho ranging from 0.585 (VVMD31) to 0.909 (Zag47) and He values varying between 0.689 (VVMD31) and 0.883 (VVMD7). In both groups, most loci showed Ho values lower than He values under the HWE, suggesting a prevalent deficiency of heterozygous, also confirmed by a comparison between the two parameters based on the inbreeding coefficient (F, (He-Ho)/He = 1 − (Ho/He). The F value describes the statistically expected degree of a reduction in heterozygosity when compared to HWE (Hardy–Weinberg Equilibrium) expectation. The F value is used to gauge the strength of inbreeding, and specifically F is the probability that two alleles in an individual are identical by descent (IBD). In Sienese vines, three loci showed elevated levels of heterozygosity (VVMD25, F = −0.13; VVMD21, F = − 0.09; Zag47, F = − 0.13); five loci (VVMD27, VVS2, VVMD31, VVMD36, VVMD7) showed signs of heterozygote deficiency (F > 0.011).
Given the peculiar genetic connotations of the two grapevine groups (Siena and varietal references), the population genetics proved that the Sienese vines seemed to still be able to crossbreed with the most common vines. This agreed with evolutionary biological timing that is quite long if compared to secondary local domestication processes.
Conclusion
Tuscany has great historical and geographical value as regards viticulture. Studies from all over the world are being used to create databanks and gathering genotypes to preserve and prevent old cultivars from genetic erosion [21, 22].
The multidisciplinary approach applied to the urban area of Siena and its surroundings revealed the existence of a high degree of unexpected genetic variability in the local grapevine germplasm. Non-intensive local vineyards in Mediterranean countries are a cradle of unexplored biodiversity that deserve scientific efforts and attention in order to safeguard grapevine biodiversity [23,24,25,26,27,28,29,30].
The recovery and characterization of the old local varieties of Vitis vinifera in Siena and its surrounding areas are still going on in the study. Over the years, new cultivars have been identified and preserved in the germplasm bank, while some others remain still unknown to established genetic and ampelographic databases.
This study conducted on Siena's antique grapevines, in addition to having scientific importance can also be seen as a positive premise for future tourism development plans. It could be a path to organizing wine-based trekking trails, alternating exploring and tasting, allowing consumers discover unknown varieties.
One of the long-term objectives of this study is, in fact, to improve the economy of local communities linked to traditional landscapes and crops, including the perspective to promote local wines that use recovered, local grapevines for cultivation.
The genetic importance and consequent recovery of rare, traditional grapevine strains can be seen as an important key factor favoring the biological future of these varieties.
References
Phumichai C, Phumichai T, Wongkaew A (2015) Novel chloroplast Microsatellite (cpSSR) markers for genetic diversity assessment of cultivated and wild Hevea rubber. Plant Mol Biol Rep 33:5. https://doi.org/10.1007/s11105-014-0850-x
Zoghlami N, Riahi L, Laucou V, Mliki A, Ghorbel A, This P (2013) Genetic structure of endangered wild grapevine Vitis vinifera ssp. sylvestris populations from Tunisia: implications for conservation and management. For Ecol Manage 310:896–902
Crespan M (2020) Genes, special issue: genetics and diversity of grapevine. ISSN 2073–4425:2020
Crespan M, Migliaro D, Larger S, Pindo M, Palmisano M, Manni A, Manni E, Polidori E, Sbaffi F, Silvestri Q, Silvestroni O, Velasco R, Virgili S, Camilli G (2021) Grapevine (Vitis vinifera L.) varietal assortment and evolution in the Marche region (central Italy). OENO One 55(3):25. https://doi.org/10.20870/oeno-one.2021.55.3.4628
Fanelli V, Roseti V, Savoia MA, Miazzi MM, Venerito P, Savino VN, Pirolo C, La Notte P, Falbo M, Petrillo F et al (2021) New insight into the identity of Italian grapevine varieties: the case study of Calabrian Germplasm. Agronomy 11:1538. https://doi.org/10.3390/agronomy11081538
De Lorenzis G, Mercati F, Bergamini C et al (2019) SNP genotyping elucidates the genetic diversity of Magna Graecia grapevine germplasm and its historical origin and dissemination. BMC Plant Biol 19:7. https://doi.org/10.1186/s12870-018-1576-y
D’Onofrio C, Tumino G, Gardiman M, Crespan M, Bignami C, de Palma L, Barbagallo MG, Muganu M, Morcia C, Novello V, Schneider A, Terzi V (2021) Parentage Atlas of Italian Grapevine varieties as inferred from SNP genotyping. Front Plant Sci 11:605934. https://doi.org/10.3389/fpls.2020.605934
Bowers J, Boursiquot JM, This P, Chu K, Johansson H, Meredith C (1999) Historical genetics: the parentage of Chardonnay, Gamay, and other wine grapes of Northeastern France. Science 285(5433):1562–1565. https://doi.org/10.1126/science.285.5433.1562
Sefc KM, Regner F, Turetschek E, Glossl J, Steinkellner H (1999) Identification of microsatellite sequences in Vitis riparia and their applicability for genotyping of different Vitis species. Genome 42:367–373
Thomas MR, Scott NS (1993) Microsatellite repeats in grapevine reveal DNA polymorphism when analyzed as sequence-tagged sites (STSs). Theor Appl Genet 86(8):985–990. https://doi.org/10.1007/BF00211051
Bowers J, Dangl GS, Vignani R, Meredith C (1996) Isolation and characterization of new polymorphic simple sequence repeat loci in grape (Vitis vinifera L). Genome 39(4):628–633. https://doi.org/10.1139/g96-080
Vignani R, Liò P, Scali M (2019) How to integrate wet lab and bioinformatics procedures for wine DNA admixture analysis and compositional profiling: case studies and perspectives. PLoS ONE 14(2):e0211962. https://doi.org/10.1371/journal.pone.0211962
Peakall R, Smouse PE (2012) GenAlex 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38(6):1358–1370. https://doi.org/10.1111/j.1558-5646.1984.tb05657.x
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959. https://doi.org/10.1093/genetics/155.2.945
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Cons Genet Res 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I (2015) Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15(5):1179–1191. https://doi.org/10.1111/1755-0998.12387
Scali M, Zifferero A, Vignani R (2018) Distribution and characterization of the Vitis vinifera L. subsp sylvestris in Southern Tuscany. Recent Patents on Biotechnol 12(3):208–220. https://doi.org/10.2174/1872208312666180125102138
Ciacci A, Giannace M (2012) “Il Progetto “Senarum Vinea” e il paesaggio storico della vite nella città di Siena: metodo, risultati, prospettive di ricerca. In: Archeologia della Vite e del Vino in Toscana e nel Lazio, Dalle tecniche dell’indagine archeologica alle prospettive della biologia molecolare. Ed. All’insegna del Giglio s.a.s., Italy, pp 731–779
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Mena A, Martìnez J, Fernàndez-Gonzàlez M (2014) Recovery, identification and relationships by microsatellite analysis of ancient grapevine cultivars from Castilla-La Mancha: the largest wine growing region in the world. Genet Resour Crop Evol 61:625–637. https://doi.org/10.1007/s10722-013-0064-3
Margaryan K, Melyan G, Röckel F, Töpfer R, Maul E (2021) Genetic diversity of Armenian Grapevine (Vitis vinifera L.) Germplasm: molecular characterization and parentage analysis. Biology 10:(12)-1279. https://doi.org/10.3390/biology10121279
Frioni T, Bertoloni G, Squeri C, Garavani A, Ronney L, Poni S (2020) Biodiversity of local Vitis vinifera L. germplasm: a powerful tool toward adaptation to global warming and desired grape composition. Front Plant Sci 11:608. https://doi.org/10.3389/fpls.2020.00608
Sargolzaei M, Rustioni L, Cola G, Ricciardi V, Bianco Piero A, Maghradze D, Failla O, Quaglino F, Toffolatti SL, De Lorenzis G (2021) Georgian grapevine cultivars: ancient biodiversity for future viticulture. Front Plant Sci 12:630122. https://doi.org/10.3389/fpls.2021.630122
Vesna M, Vladan B, Giannetto S, Crespan M (2014) SSR molecular marker analysis of the grapevine germplasm of Montenegro. J Int Sci Vign Vin 48(2):87–97. https://doi.org/10.20870/oeno-one.2014.48.2.1562
Villano C, Aiese Cigliano R, Esposito S, D’Amelia V, Iovene M, Caputo D, Aversano R (2022) DNA-based technologies for grapevine biodiversity exploitation: state of the art and future perspectives. Agronomy 12(2):491. https://doi.org/10.3390/agronomy12020491
Biniari K, Stravakaki M (2019) Genetic study of native grapevine varieties of Northern, Western and Central Greece with the use of ampelographic and molecular methods. Not Bot Horti Agrobo 47(1):46–53. 47.15835/nbha47111213
Augusto D, Ibáñez J, Pinto-Sintra AL, Falco V, Leal F, Martínez-Zapater JM, Oliveira AA, Castro I (2021) Grapevine diversity and genetic relationships in Northeast Portugal Old Vineyards. Plants 10:2755. https://doi.org/10.3390/plants10122755
Buhner-Zaharieva T, Moussaoui S, Lorente M, Andreu J, Núñez R, Ortiz JM, Gogorcena Y (2010) Preservation and molecular characterization of ancient varieties in Spanish Grapevine germplasm collections. Am J Enol Vitic 61:557–562. https://doi.org/10.5344/ajev.2010.09129
Aversano R, Basile B, Buonincontri MP, Carucci F, Caputo D, Frusciante L et al (2017) Dating the beginning of the Roman viticultural model in the Western Mediterranean: the case study of Chianti (Central Italy). PLoS ONE 12(11):e0186298. https://doi.org/10.1371/journal.pone.0186298
Acknowledgements
The ampelography was done by Studio Consulenza Agronomica Gambassi e Zorzi. We thank Associazione Nazionale Città del Vino for the constant support of the Senarum Vinea Project. Thanks to Massimilano Canuti for the information about Gacchè.
Funding
Open access funding provided by Università degli Studi di Siena within the CRUI-CARE Agreement. The study was partially funded by Fondazione Monte dei Paschi di Siena. The authors declare no conflicts of interest. Rita Vignani is on the board of directors of Serge-genomics Srl and receives no compensation from the company.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Significance statement: A multidisciplinary study, including genetic characterization of the grapevine (Vitis vinifera L.) germplasm in the urban area of Siena (Italy), revealed the existence of rare strains and a high degree of biodiversity. Population genetics showed the existence of population sub-structures.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vignani, R., Scali, M. & Ciacci, A. Grapevine (Vitis vinifera L.) Germplasm from Siena (Italy) Includes Rare Strains and Reveals Population Structuring. Proc. Natl. Acad. Sci., India, Sect. B Biol. Sci. 94, 659–669 (2024). https://doi.org/10.1007/s40011-024-01584-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40011-024-01584-6