Abstract
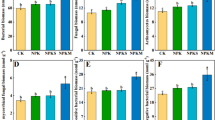
An experiment was executed at field condition to evaluate ecological impact of Bt cotton (Mech 162 + Bt) on soil microbes and Bt protein behavior in soil. Root exudates of Bt cotton plant containing Bt protein enter into soil which persisted in soil for atleast 235 days after sowing (DAS). During vegetative and flowering stage, Bt protein was not detectable in both qualitative and quantitative assay but, during onset of boll formation at 150 DAS, a level of 0.0013 μg/g of Cry 1Ac protein was quantified in rhizospheric soil which further increased exponentially up to 0.0033 μg/g at 195 DAS and then maintained at the same level till 235 DAS with Cry1Ac expression level of 0.0029 μg/g soil. Rhizospheric soil of Bt cotton had significantly higher (p < 0.05) abundance of Fatty Acid Methyl Esters (FAME) of gram positive bacteria. Abundance of polyunsaturated FAMEs that are indicators of fungi, were slightly higher in Bt as compared to non-Bt cotton soils. The metabolic profiling, also called community level physiological profiling (CLPP) showed more microbial functional diversity (Shannon–Wiener Diversity) in soil samples of different growth stages of Bt cotton than its corresponding non-Bt variety. Our results showed that Cry 1AC Bt protein persists in clay loamy soil even after harvesting and significantly altered microbial community composition and functional diversity in rhizospheric soil of Bt cotton than non-Bt. Any possible and unintentional changes in composition of root exudes of Bt cotton could regulate the selection of different microbial communities and thus can alter functional responses.
Graphic abstract








Similar content being viewed by others
Data availability
Will be shared as per request.
Code availability
(software application or custom code): NA.
References
Aulakh MS, Wassmann R, Bueno C, Kreuzwieser J, Rennenberg H (2008) Characterization of root exudates at different growth stages of ten rice (Oryza sativa L.) cultivars. Pl Biol. https://doi.org/10.1055/s-2001-12905
Bai Q, Gattinger A, Zelles L (1999) Characteristic of microbial consortia in paddy soil by analysis of PLFA. FEMS Microbiol Ecol 39(4):273–281
Birch ANE, Geoghegan IE, Majerus MEN et al (1999) Tritrophic interactions involving pest aphids, predatory 2-spot ladybirds and transgenic potatoes expressing snowdrop lectin for aphid resistance. Mol Breed 5:75–83
Brusetti L, Francia P, Bertolini C et al (2005) Bacterial communities associated with the rhizosphere of transgenic Bt 176 maize (Zea mays) and its non transgenic counterpart. Pl Soil. https://doi.org/10.1007/s11104-005-5399-x
Cetinic V (2019) Cotton pesticide statistics you need to hear. Cotton pesticides. Wriggly Toes publishing. https://wrigglytoes.com.au/blog/cotton-pesticides/cotton-pesticide-statistics-you-need-to-hear. Accessed 5 June 2021
Das NR, Chaudhary A, Aggarwal PK et al (2009) Detection and persistence of Bt protein in decomposition study of Bt leaves of transgenic cotton. J Environ Res Dev 3:859–866
Donegan KK, Schaller DL, Stone JK et al (1996) Microbial populations, fungal species diversity and plant pathogen levels in field plots of potato plants expressing the Bacillus thuringiensis var. tenebrionis endoprotein. Trans Res 5:25–35
Frostegard A, Tunlid A, Baath E (1993) Shifts in the structure of soil microbial communities in limed forests as revealed by phospholipid fatty acid analysis. Soil Biol Biochem 25:723–730
Garland JL (1996) Analytical approaches to the characterization of samples of microbial communities using patterns of potential C source utilization. Soil Biol Biochem 28(2):213–221
Grayston SJ, Wang S, Campbell CD, Edwards AC (1998) Selective influence of plant species on microbial diversity in the rhizosphere. Soil Biol Biochem 30:369–378
Gupta VSR, Watson S (2004) Ecological impact of GM cotton on soil biodiversity. CSIRO project report. https://www.environment.gov.au/system/files/pages/ca807cf6-8ad9-478f-9d7d-f3894704642d/files/bt-cotton.pdf. Accessed 5 June 2021
Head G, Surber JB, Waston JA, Martin JW, Duan JJ (2002) No detection of Cry 1Ac protein insoil after multiple years transgenic Bt cotton (Bollgard) use. Environ Ent 31(1):30–36. https://doi.org/10.1603/0046-225X-31.1.30
Hopkins DW, Gregorich EG (2003) Detection and decay of Bt endoprotein in soil from a field trial with genetically modified maize. Eur J Soil Sci 54:793–800
James C (2017) Global status of commercialized biotech/GM crops in 2017: Biotech crop adoption surges as economic benefits accumulate in 22 years. ISAAA Briefs No. 53
Jensen LS, Sorensen J (1994) Microscale fumigation-extraction and substrate-induced respiration methods for measuring microbial biomass in barley rhizosphere. Plant Soil 162:151–161
Karuri H, Amata R, Amugune N, Waturu C (2013) Effect of Bt cotton expressing Cry1Ac and Cry2Ab2 protein on soil nematode community assemblages in Mwea. J Anim Plant Sci 19:2864–2879
Koskella J, Stotzky G (1997) Microbial utilization of free and clay-bound insecticidal proteins from Bacillus thuringiensis and their retention of insecticidal activity after incubation with microbes. Appl Environ Microbiol 63(9):3561–3568
Kumar R, Tian JC, Naranjo SE, Shelton AM (2014) Effects of Bt cotton on Thrips tabaci (Thysanoptera: Thripidae) and its predator, Orius insidiosus (Hemiptera: Anthocoridae). J Econ Ent 107:927–932
Kranthi KR (2020) Stone GD (2020) Long-term impacts of Bt cotton in India. Nat Plants 6:188–196. https://doi.org/10.1038/s41477-020-0615-5
Losey JE, Rayor LS, Carter ME (1999) Transgenic pollen harms monarc larvae. Nature 399:214
Mulder C, Wouterse M, Raubuch M, Roelofs W, Rutgers M (2006) Can transgenic maize affect soil microbial communities. PLOS Comput Biol 2(9):128. https://doi.org/10.1371/journal.pcbi.0020128
Mansouri H, Petit A, Oger P, Dessaux Y (2002) Engineered rhizosphere: the trophic bias is independent of the opine type, the soil origin, and the plant species. Appl Environ Microbiol 68:2562–2566
Normander B, Prosser JJ (2000) Bacterial origin and community composition in the barley photosphere as a function of habitat and presowing condition. Appl Environ Microbiol 66:4372–4377
Palm CJ, Donegan KK, Harris D, Siedler RJ (1994) Quantification in soil of Bacillus thuringiensis var. kurstaki Δendoprotein from transgenic plants. Mol Ecol 3:145–151
Perlak FJ, Deaton RW, Armstrong TA et al (1993) Insect resistant cotton plant. Bio Technology 8:939–943
Quim M, Zilberman D (2003) Yield effects of genetically modified Crops in developing countries. Science 299:900–901
Shera PS, Prasun K, Sudhendu S, Sangha KS (2018) Impact of Bt cotton expressing single (Cry1Ac) and dual toxins (Cry1Ac and Cry2Ab) on the fitness of the predator Chrysoperla zastrowi sillemi (Esben-Petersen): prey-mediated tri-trophic analysis. Egypt J Biol Pest Co. https://doi.org/10.1186/s41938-018-0102-8
Sanaullah Y, Hafiz AN, Ahmad F et al (2016) Impact of Bt-cotton on soil microbiological and biochemical attributes. Plant Prod Sci 19(4):458–467. https://doi.org/10.1080/1343943X.2016.1185637
Saxena and Stotzky G, (2000) Insecticidal protein from Bt is released from root of transgenic Bt corn in vitro and in situ. FEMS Microbiol Ecol 33:35–39
Saxena D, Flores S, Stotzky G (2002) Bt protein is released in root exudates from 12 transgenic corn hybrids representing three transformation events. Soil Biol Biochem 34:133–137
Sims SR, Ream JE (1997) Soil inactivation of the Bacillus thuringiensisvar.kurstaki Cry IIA insecticidal protein within transgenic cotton tissue: laboratory microcosm and field studies. J Agric Food Chem 45:1502–1505
Singh RJ, Ahlawat IP, Singh S (2013) Effects of transgenic Bt cotton on soil fertility and biology under field conditions in subtropical inceptisol. Environ Monit Assess 185:485–495
Stotzky G (2000) Persistence and biological activity in soil of insecticidal proteins from Bacillus thuringiensisand of bacterial DNA bound on clays and humic acids. J Environ Qual 29:691–770
Tapp H, Stotzky G (1998) Persistence of the insecticidal protein from Bacillus thuringiensis subsp. Kurstaki in Soil. Soil Biol Biochem 30(4):471–476
Venkateswerlu G, Stotzky G (1992) Binding of proprotein and protein proteins from Survival of Bacillus thuringiensis subsp. kurstaki on clay minerals. Curr Microbiol 25:1–9
Velmourougane K, Sahu A (2013) Impact of transgenic cottons expressing Cry 1Ac on soil biological attributes. Pl Soil Environ 59(3):108–114
Yan WD et al (2007) Overexpression of a foreign Bt gene in cotton affects the low-molecular-weight components in root exudates. Pedosphere 17:324–330
Zwahlen C, Hilbeck A, Gugerli P, Nentwig W (2003) Degradation of the Cry1Ab protein within transgenic Bacillus thuringiensis corn tissue in the field. Mol Ecol 12:765–775
Acknowledgements
Authors duly acknowledge Post Graduate School, Indian Agricultural Research Institute (IARI), Pusa, New Delhi, India for providing all facilities and Indian Council of Agricultural Research (ICAR), India for funding to execute this experiment.
Funding
Funding in the form of Junior Research Fellowship from Post Graduate School, Indian Agricultural Research Institute (IARI), Pusa, New Delhi, India, was received by Dr. Namita Das Saha for executing this experiment.
Author information
Authors and Affiliations
Contributions
Dr. NDS and Mr. AKS had executed the experiment, Dr. AC conceptualized the idea and monitored the experimental execution, Dr. AB helped in statistical analysis of data.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest for any part of this research article.
Consent to participate
(include appropriate statements): NA.
Consent for publication
(include appropriate statements): All the authors have given consent for this publication.
Additional information
Editorial responsibility: Jing Chen.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Saha, N.D., Chaudhary, A., Singh, A.K. et al. Evaluation of Bt cotton effects on belowground microbial community structure and function in tropical western India. Int. J. Environ. Sci. Technol. 19, 7411–7424 (2022). https://doi.org/10.1007/s13762-021-03543-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13762-021-03543-4