Abstract
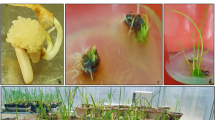
Transformation of rice was done through Agrobacterium mediated, utilizing a binary vector pBS, harboring a fungal gene endo α-1, 3-glucanase from Trichoderma harzianum under rice constitutive promoter actin2 and Agrobacterium nopaline synthase (nos) transcriptional terminator in a T-DNA. Likewise, a selectable marker gene hygromycin phosphotransferese (hpt) resistant to Hygromycin B was cloned in the middle of actin2 and nos terminator in a similar T-DNA. The expression of endo α-1, 3-glucanase was affirmed by cloning gfp gene after the fungal gene in the transformation DNA cassette. In the first generation of transgenic rice lines, out of 912 just 209 plants were false positive affirmed through PCR based screening for the transgene. The positive transgenic lines were tried with fungal infection by leaf cut test in vitro and foliar leaf shower technique in vivo. They demonstrated an exceptional protection against sheath blight disease. Further, seeds of all positive transgenic plants of the first generation were developed and screened in next generation. Just 62 plants were false positive out of 873 transgenic lines in this generation. In the comparative way, they were tried against the fungal disease and they demonstrated the exceptional protection once more. In this way, in this investigation, a fungal gene endo α-1, 3-glucanase was transformed into rice (IR 64) effectively which demonstrated protection against fungus Rhizoctonia solani.





Similar content being viewed by others
Abbreviations
- PAMPs:
-
Pathogen associated molecular patterns
- PRRs:
-
Pattern recognition receptors
- hpt :
-
Hygromycin phosphotramferase
- YEM:
-
Yeast extract manitol
- AAM:
-
A protocol for Agrobacterium mediated transformation method medium
References
Akagi A, Jiang CJ, Takatsuji H (2015) Magnaporthe oryzae inoculation of rice seedlings by spraying with a spore suspension. Bio-protocol 5:e1486. https://doi.org/10.21769/BioProtoc.1486
Chen L, Ai P, Zhang J, Deng Q, Wang S, Li S, Zhu J, Li P, Zheng A (2016a) RSIADB, a collective resource for genome and transcriptome analyses in Rhizoctonia solani AG1 IA. Database. https://doi.org/10.1093/database/baw031
Chen L, Wang Q, Chen H, Sun G, Liu H, Wang H (2016b) Agrobacterium tumefaciens-mediated transformation of Botryosphaeria dothidea. World J Microbiol Biotechnol 32:1–5. https://doi.org/10.1007/s11274-016-2045-0
Cota-Sánchez JH, Remarchuk K, Ubayasena K (2006) Ready-to-use DNA extracted with a CTAB method adapted for herbarium specimens and mucilaginous plant tissue. Plant Mol Biol Rep 24:161
de Vetten N, Wolters AM, Raemakers K, van der Meer I, ter Stege R, Heeres E, Visser R (2003) A transformation method for obtaining marker-free plants of a cross-pollinating and vegetatively propagated crop. Nat Biotechnol 21:439–442. https://doi.org/10.1038/nbt801
Eziashi EI, Omamor IB, Dimaro-Oruade EA, Ogunkanmi LA (2007) Control of phytotoxin from Ceratocystis paradoxa using Trichoderma species phytotoxins on oil palm (Elaeis quineensis Jacq.) sprouted seeds. Plant Pathol J 6:324–329. http://www.ansinet.org/ppj
Fujikawa T, Sakaguchi A, Nishizawa Y, Kouzai Y, Minami E, Yano S, Koga H, Meshi T, Nishimura M (2012) Surface α-1, 3-glucan facilitates fungal stealth infection by interfering with innate immunity in plants. PLoS Pathog 8:e1002882. https://doi.org/10.1371/journal.ppat.1002882
Mayo S, Gutierrez S, Malmierca MG, Lorenzana A, Campelo MP, Hermosa R, Casquero PA (2015) Influence of Rhizoctonia solani and Trichoderma spp. in growth of bean (Phaseolus vulgaris L.) and in the induction of plant defense-related genes. Front. Plant Sci 6:685. https://doi.org/10.3389/fpls.2015.00685
Metzker ML, Caskey CT (2009) Polymerase chain reaction (PCR). eLS. https://doi.org/10.1002/9780470015902.a0000998.pub2
Moradpour M, Abdullah SNA (2017) Cisgenesis and Intragenesis as new strategies for crop improvement. In Crop Improvement, pp 191–216. Springer, Cham. https://doi.org/10.3389/fpls.2016.01608
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Pettitt TR, Wainwright MF, Wakeham AJ, White JG (2011) A simple detached leaf assay provides rapid and inexpensive determination of pathogenicity of Pythium isolates to ‘all year round’ (AYR) chrysanthemum roots. Plant Pathol 60:946–956
Scaramelli L, Balestrazzi A, Bonadei M, Piano E, Carbonera D, Confalonieri M (2009) Production of transgenic barrel medic (Medicago truncatula Gaernt.) using the ipt-type MAT vector system and impairment of Recombinase-mediated excision events. Plant Cell Rep 28:197–211. https://doi.org/10.1007/s00299-008-0634-6
Sengupta S, Chakraborti D, Mondal HA, Das S (2010) Selectable antibiotic resistance marker gene-free transgenic rice harbouring the garlic leaf lectin gene exhibits resistance to sap-sucking planthoppers. Plant Cell Rep 29:261–271. https://doi.org/10.1007/s00442-009-1541-4
Svitashev S, Young J, Schwartz C, Gao H, Falco SC, Cigan AM (2015) Targeted mutagenesis, precise gene editing and site-specific gene insertion in maize using Cas9 and guide RNA. Plant Physiol. https://doi.org/10.1104/pp.15.00793
Tang W, Chen H, Xu C, Li X, Lin Y, Zhang Q (2006) Development of insect-resistant transgenic indica rice with a synthetic cry1C gene. Mol Breeding 18:1–10. https://doi.org/10.1007/s11032-006-9002-9
Toriyama K, Hinata K (1985) Cell suspension and protoplast culture in rice. Plant Sci 41:179–183. https://doi.org/10.1016/0168-9452(85)90086-X
Xue M, Yang J, Li Z, Hu S, Yao N, Dean RA, Liu L (2012) Comparative analysis of the genomes of two field isolates of the rice blast fungus Magnaporthe oryzae. PLoS Genet 8:e1002869. https://doi.org/10.1371/journal.pgen.1002869
Acknowledgements
The study was supported by Visargha Agrisciences Pvt. Ltd., TBI, Campus-11, Kalinga Institute of Industrial Technology (KIIT), Deemed to be University, Bhubaneswar-751024, Odisha, India in association with School of Biotechnology, Campus-11, Kalinga Institute of Industrial Technology (KIIT), Deemed to be University, Bhubaneswar-751024, Odisha, India.
Author information
Authors and Affiliations
Contributions
RK designed and did the most of the experiments and wrote the MS. KK, KCH and LK did some part of experiments. BKB edited the MS.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumar, R., Kumari, K., Hembram, K.C. et al. Expression of an endo α-1, 3-Glucanase gene from Trichoderma harzianum in rice induces resistance against sheath blight. J. Plant Biochem. Biotechnol. 28, 84–90 (2019). https://doi.org/10.1007/s13562-018-0465-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-018-0465-7