Abstract
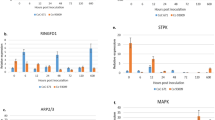
Sugarcane is one of the most important commercial crops in India, supporting the second largest agro-based sugar industry. However for more than a century, the production is imperiled by the most devastating fungus Colletotrichum falcatum Went, which causes red rot in sugarcane. Complex polyploidy and limited understanding on the inheritance to red rot resistance makes breeding efforts more complicated in sugarcane. Transcription factors (TFs) play a key role in regulating defense response, which offers much promise for the manipulation of metabolic pathways, potential in regulating multiple signaling networks leading to plant defense against pathogens. In order to better manage red rot disease in sugarcane, we sought to screen the TFs possibly involving in pathogen defense by analyzing 5 major defense-related TF classes (WRKY, bZIP, MYB, NAC and TLP). In this study two parallel sets of experiments were carried out to compare the differential regulation of these classes of TFs upon pathogen challenge and SAR priming. Among the 41 TFs studied, differential regulation of 24 TFs upon pathogen challenge and 15 TFs upon SAR inducer priming were observed. Comparison of incompatible interaction and SAR inducer priming revealed that 8 TFs were highly induced in both the cases. Collectively, the results showed that TFs which are significantly induced early may involve actively in triggering or co-ordinating defense against pathogen invasion.



Similar content being viewed by others
Abbreviations
- TFs:
-
Transcription factors
- SAR:
-
Systemic acquired resistance
- BTH:
-
Benzothiadiazole
- SA:
-
Salicylic acid
- Cf :
-
Colletotrichum falcatum
- hpi:
-
hours post inoculation
References
Bell JN, Ryder TB, Vincent PM, Lamb CJ et al (1986) Differential accumulation of plant defense gene transcripts in a compatible and incompatible plant-pathogen interaction. Mol Cell Biol 1615–1623
Chen R, Zhongfu N, Nie X, Qin Y, Dong G, Sun Q (2005) Isolation and characterization of genes encoding MYB transcription factor in wheat (Triticum aestivem L.). Plant Sci 169(6):1146–1154
Chen WJ, Zhu T (2004) Networks of transcription factors with roles in environmental stress response. Trends Plant Sci 9(12):591–596
Desveaux D, Subramaniam R, Despres C, Mess JN, Lévesque C, Fobert PR, Dangl JL, Brisson N (2004) A “whirly” plant transcription factor is required for salicylic acid-dependent disease resistance in Arabidopsis. Dev Cell 6:229–240
Durrant WE, Dong X (2004) Systemic acquired resistance. Annu Rev Phytopathol 42:185–209
Gray J, Bevan M, Brutnell T, Buell R, Cone K, Hake S, Jackson D, Kellogg E, Lawrence C, McCouch S, Mockler T, Moose S, Paterson A, Peterson T, Rokshar D, Souza GM, Springer N, Stein N, Timmermans M, Wang GL, Grotewold E (2009) Naming transcription factors. Plant Physiol 149:4–6
Guo AY, Chen X, Gao G, Zhang H, Zhu QH, Liu XC, Zhong YF, Gu XC, He K, Luo JC (2008) PlantTFDB: a comprehensive plant transcription factor database. Nucleic Acids Res 36:D966–D969
Kim CY, Lee SH, Park HC, Bae CG, Cheong YH, Choi YJ, Han CD, Lee SY, Lim CO, Cho MJ (2000) Identification of rice blast fungal elicitor-responsive genes by differential display analysis. Mol Plant Microbe Interact 13:470–474
Latchman DS (1997) Transcription factors: an overview. Int J Biochem Cell Biol 29(12):1305–1312
Liu J, Que Y, Guo J, Xu L, Wu J, Chen R (2012) Molecular cloning and expression analysis of a WRKY transcription factor in sugarcane. Afr J Biotechnol 11(24):6434–6444
Lopes MA, Hora Junior BT, Dias CV, Santos GC, Gramacho KP, Cascardo JCM, Gesteira AS, Micheli F (2010) Expression analysis of transcription factors from the interaction between cacao and Moniliophthora perniciosa (Tricholomataceae). Genet Mol Res 9(3):1279–1297
Naoumkina M, He X, Dixon R (2008) Elicitor-induced transcription factors for metabolic reprogramming of secondary metabolism in Medicago truncatula. BMC Plant Biol 8:132
Nogueira FTS, De Rosa VE, Menossi M, Ulian EC, Arruda P (2003) RNA expression profiles and data mining of sugarcane response to low temperature. Plant Physiol 132(4):1811–1824
Ooka H, Satoh K, Doi K, Nagata T, Otomo Y, Murakami K, Matsubara K, Osato N, Kawai J, Carninci P, Hayashizaki Y, Suzuki K, Kojima K, Takahara Y, Yamamoto K, Kikuchi S (2003) Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res 10(6):239–247
Ramesh Sundar A, Viswanathan R, Malathi P, Padmanaban P (2006) Mechanism of resistance induced by plant activators against Colletotrichum falcatum in sugarcane. Arch Phytopathol PFL 39(4):259–272
Ramesh Sundar A, Velazhahan R, Nagarathinam S, Vidhyasekaran P (2008) Induction of pathogenesis-related proteins in sugarcane leaves and cell-cultures by a glycoprotein elicitor isolated from Colletotrichum falcatum. Biol Plant 52(2):321–328
Ramesh Sundar A, Viswanathan R, Nagarathinam S (2009) Induction of systemic acquired resistance (SAR) using synthetic signal molecules against Colletotrichum falcatum in sugarcane. Sugar tech 11(3):274–281
Reitz MU, Bissue JK, Zocher K, Attard A, Huckelhoven R, Becker K, Imani J et al (2012) The subcellular localization of tubby-like proteins and participation in stress signaling and root colonization by the mutualist Piriformospora indica. Plant Physiol 160(1):349–364
Rushton PJ, Somssich IE (1998) Transcriptional control of plant genes responsive to pathogens. Curr Opin Plant Biol 1:311–315
Schenk PM, Kazan K, Manners JM, Anderson JP, Simpson RS, Wilson IW, Somerville SC et al (2003) Systemic gene expression in Arabidopsis during an incompatible interaction with Alternaria brassicicola. Plant Physiol 132:999–1010
Singh K, Foley RC, Onate-Sanchez L (2002) Transcription factors in plant defense and stress responses. Curr Opin Plant Biol (5):430–436
Singh K, Singh RP (1989) Red rot. In: Ricaud CG, Egan BT, Gillaspie AG, Hughes CG (eds) Sugarcane diseases of the world: major diseases. Elsevier, Amsterdam, pp 169–188
Srinivasan KV, Bhat NR (1961) Red rot of sugarcane criteria of grading resistance. J Indian Bot Soc 40:566–577
Thurow C, Schiermeyer A, Krawczyk S, Butterbrodt T et al (2005) Tobacco bZIP transcription factor TGA2.2 and related factor TGA2.1 have distinct roles in plant defense responses and plant development. Plant J 44:100–113
Torregrosa CA, Dumas B, Krajinski F, Esquerre-Tugaye MT, Jacquet C (2006) Transcriptomic approaches to unravel plant–pathogen interactions in legumes. Euphytica 147:25–36
Vailleau F, Daniel X et al (2002) A R2R3-MYB gene, AtMYB30, acts as a positive regulator of the hypersensitive cell death program in plants in response to pathogen attack. Proc Natl Acad Sci U S A 99(15):10179–10184
Vettore A, da Silva FR, Kemper EL, Arruda P (2001) The libraries that made SUCEST. Genet Mol Biol 24(1–4):1–7
Vincentz M, Schlogl PS, Correa LGG, Kuhne F, Leite A (2001) Phylogenetic relationship between Arabidopsis and bZIP transcriptional regulatory factors. Genet Mol Biol 24:55–60
Viswanathan R (2000) Possible involvement of anthocyanin compounds in resistance of sugarcane against red rot. Indian phytopath 53(3):311–313
Viswanathan R, Malathi P, Padmanaban P (2003) Variation in sugarcane red rot pathogen Colletotrichum falcatum Went. In: Rao GP, Manoharachari C, Bhat DJ, Rajak RC, Lakhanpal TN (eds) Frontiers of fungal diversity in India. International Book Distributing Co, Lucknow, pp 639–667
Viswanathan R, Ramesh sundar A, Malathi P, Rahul PR, Ganesh kumar V, Prathima PT, Raveendran M, Kumar KK, Balasubramanian P (2009) Interaction between sugarcane and Colletotrichum falcatum causing red rot: understanding disease resistance at transcription level. Sugar tech 11(1):44–50
Viswanathan R (2010) Plant disease: Red rot of sugarcane. Anmol pubilication pvt. ISBN: 978-81-261-4214-9
Wang D, Amornsiripanitch N, Dong X (2006) A genomic approach to identify regulatory nodes in the transcriptional network of systemic acquired resistance in plants. PLoS pathogens 2(11):e123. doi:10.1371/journal.ppat.0020123
Xia N, Zhang G, Liu XY, Deng L, Cai GL, Zhang Y, Wang XJ, Zhao J, Huang LL, Kang ZS (2010) Characterization of a novel wheat NAC transcription factor gene involved in defense response against stripe rust pathogen infection and abiotic stresses. Mol Biol Rep 37(8):3703–3712
Yanhui C, Xiaoyuan Y, Kun H, Meihua L, Jigang L, Zhaofeng G, Zhiqiang L, Yunfei Z, Xiaoxiao W, Xiaoming Q, Yunping S, Li Z, Xiaohui D, Jingchu L, Xing-Wang D, Zhangliang C, Hongya G, Li-Jia Q (2006) The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family. Plant Mol Biol 60(1):107–124
Zhang H, Jin JP, Tang L, Zhao Y, Gu XC, Gao G, Luo JC (2011) PlantTFDB 2.0: update and improvement of the comprehensive plant transcription factor database. Nucleic Acids Res 39:D1114–D1117
Acknowledgments
The authors are grateful to Dr.N. Vijayan Nair, Director of the institute for providing facilities and encouragement. The financial support received from Indian Council of Agricultural Research, Department of Biotechnology and the Department of Science of Technology, New Delhi is greatly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Muthiah, M., Ramadass, A., Amalraj, R.S. et al. Expression profiling of transcription factors (TFs) in sugarcane X Colletotrichum falcatum interaction. J. Plant Biochem. Biotechnol. 22, 286–294 (2013). https://doi.org/10.1007/s13562-012-0157-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-012-0157-7