Abstract
Introduction
Androgenetic alopecia (AGA) is a prevalent, multifactorial form of hair loss involving complex aetiological factors, such as altered androgen regulation and energy metabolism. Existing treatments offer limited success, thus highlighting the need for advanced, personalised therapeutic strategies. This study focuses on correlating the genetic mechanisms of AGA with molecular targets involved in the response to current treatment modalities.
Methods
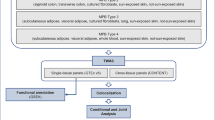
An anonymised database including 26,607 patients was subjected to analysis. The dataset included information on patients’ genotypes in 26 single nucleotide polymorphisms (SNPs), specifically, and diagnosed AGA grades, representing a broad range of ethnic backgrounds.
Results
In our sample, 64.6% of males and 35.4% of females were diagnosed with female pattern hair loss. This distribution aligns well with prior studies, thus validating the representativeness of our dataset. AGA grading was classified using the Hamilton–Norwood and Ludwig scales, although no association was found to the grade of the disease. SNP association analysis revealed eight SNPs, namely rs13283456 (PTGES2), rs523349 (SRD5A2), rs1800012 (COL1A1), rs4343 (ACE), rs10782665 (PTGFR), rs533116 (PTGDR2), rs12724719 (CRABP2) and rs545659 (PTGDR2), to be statistically significant with a p-value below 0.05.
Conclusions
The study establishes a preliminary association between eight specific SNPs and AGA. These genetic markers offer insights into the variability of therapeutic responses, thus underlining the importance of personalised treatment approaches. Our findings show the potential for more targeted research to understand these SNPs’ and further roles in AGA pathophysiology and in modulating treatment response.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Androgenetic alopecia (AGA) affects 64.6% of males and 35.4% of females in a dataset of 26,607 patients, validating its representativeness. |
Analysis of 26 single nucleotide polymorphisms (SNPs) identifies eight markers significantly associated with androgenetic alopecia, highlighting the genetic basis of hair loss. |
The study establishes a robust association between eight specific single nucleotide polymorphisms and androgenetic alopecia, providing valuable insights for personalised treatment approaches. |
Introduction
Androgenetic alopecia (AGA) represents the most common form of hair loss, affecting a significant proportion of both male and female populations. In men, it typically commences between adolescence and 30 years of age, with early onset indicating a more severe progression. For women, AGA, female pattern hair loss (FPHL), usually manifests in two age peaks, namely around 20 years (premenopausal) and 55 years (postmenopausal) [1]. The disorder is mediated by dihydrotestosterone (DHT), a testosterone metabolite that negatively influences the hair follicle, resulting in follicular atrophy, a decrease in hair diameter and shortening of the anagen phase leading to more frequent hair cycles. Other key contributing factors include alterations in prostaglandin pathways and Wnt/β-catenin pathways, highlighting the multifactorial aetiology of this condition [1, 2].
Various treatment modalities have been developed to combat AGA, with the ultimate goal of enhancing hair density, thickness and growth and, importantly, patient self-perception of hair health. Pharmacological treatments, such as minoxidil, finasteride and dutasteride, work via different mechanisms—arteriolar vasodilation and 5-α-reductase inhibition, respectively—to alleviate the symptoms of AGA. Other interventions, such as low-level laser light therapy, and surgical methods, such as hair transplants, also play significant roles in the treatment landscape[1,2,3,4].
However, despite the variety of available treatments, the success of pharmacological therapy remains limited. Meta-analysis of randomised controlled trials indicate that all treatments, including 5% minoxidil in men and 2% minoxidil in women, 1 mg finasteride in men, and low-level laser light therapy, prove superior to placebo, but they still present significant limitations. A 12-month observational study demonstrated that a 5% minoxidil topical solution decreased the affected area in 62.0% of subjects, with hair regrowth rated as effective or very effective in 63.7% of patients[5]. Nevertheless, physicians and patients perceive the efficacy of the treatments differently, with the former often rating the effectiveness higher than the latter. Therefore, it becomes evident that approaches to enhance treatment efficacy or predict treatment response could provide a significant advantage. Such advancements could pave the way towards personalised, targeted therapies, ultimately improving outcomes for patients suffering from AGA [3,4,5,6,7,8].
AGA’s molecular mechanisms involve primarily the increased sensitivity of the hair follicles’ androgen hormones, especially dihydrotestosterone (DHT). This hormone is synthesised from testosterone via the enzymatic action of 5-alpha-reductases. A pertinent feature of AGA is the increased levels of unbound, or active, testosterone, underlining the pivotal role of androgens in the pathophysiology of AGA. Additionally, the androgen receptor gene on the X chromosome has been identified as a genetic susceptibility loci for AGA, further underscoring the link between androgen regulation and the development of AGA [6, 9].
Recently, specific molecular pathways associated with energy metabolism have been implicated in the pathogenesis of AGA. For instance, the peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PGC1a) is found to be upregulated in miniaturised hair samples from patients with AGA. PGC1a is known to be a key regulator of mitochondrial biogenesis and energy metabolism, suggesting its potential role in hair growth and follicular miniaturisation [10]. Additionally, a differential expression of microRNAs (miRNAs) has been noted. Specific miRNAs that bind to PGC1a and androgen receptor transcripts display altered expression profiles in hair miniaturisation, hinting at a possible miRNA-mediated epigenetic regulation in AGA [11]. Furthermore, upregulation of PPARγ and retinoic acid receptor alpha (RAR-a), along with PGC1a, suggests potential interactions between the PGC1a/PPARγ pathway and retinoid pathways in AGA progression. This emerging understanding of the molecular mechanisms of AGA provides novel insights and potential therapeutic targets in the management of this common condition [6, 9,10,11,12,13,14].
AGA’s hereditary characteristics have been illustrated through multiple genetic studies that have identified single-nucleotide polymorphisms (SNPs) in eight different genomic loci associated with AGA development. However, these identified loci do not account for the entire heritable risk, indicating that additional, yet unidentified, risk loci exist. It is also worth noting that an in-depth understanding of AGA’s pathophysiology remains to be fully achieved, and the discovery of each newly associated locus provides novel insights into the contributing biological pathways. Among the genetic basis of AGA, four genetic risk loci have been identified with genome-wide significance on chromosomes 2q35, 3q25,1, 5q33,3 and 12p12,115,16. Notably, the strongest association has been detected for the rs7349332 SNP on chromosome 2q35, which is located intronically in the WNT10A gene. Research has indicated that the WNT10A gene, a member of the WNT gene family involved in various developmental processes and adult tissue homeostasis, has a substantial role in hair development and hair cycle regulation [15, 16].Furthermore, evidence points to changes in WNT10A expression being potentially crucial in AGA aetiology. These changes might delay the telogen to anagen transition and shorten anagen duration, characteristics closely linked with AGA’s clinical presentation. Consequently, this evidence underscores the potential importance of WNT signalling pathways in the aetiology of AGA and adds to the complexity of AGA’s genetic landscape. Despite these advancements, more in-depth genetic and functional studies are needed to elucidate further the role of these identified risk loci and WNT10A in AGA [15, 16].
Expanding on this comprehensive understanding of the genetic and molecular mechanisms that underlie AGA, it becomes clear that numerous intricate biological pathways and genetic loci contribute to the development of this disorder. This evidence underscores the necessity for ongoing exploration to elucidate the underlying pathophysiology thoroughly [2, 6]. Accordingly, the forthcoming study intends to delve further into the genetic landscape of AGA, specifically focusing on evaluating 26 SNPs of interest. These SNPs, chosen for their potential association with AGA, are hypothesised to correlate with the molecular mechanisms that play a role in response to principal drugs used in alopecia treatment. By comprehending the genetic basis of response variability to treatment modalities, such as minoxidil, finasteride and dutasteride, this investigation may provide valuable insights, facilitating the development of personalised therapeutic strategies to counter this common condition [4,5,6,7,8]. This study aims to shed further light on the pathway towards a more effective, individualised therapeutic approach for patients suffering from AGA.
Methods
Study Design and Data Acquisition
We analysed a comprehensive database containing matched genotypes of patients for the 26 SNPs examined in relation to their previously diagnosed AGA at different grades. The complete data consisted of information from 26,607 patients whose genotypes were correlated to the diagnostic of AGA and, tentatively, AGA grading, at the time of DNA collection.
This dataset was fully anonymised prior to any data processing, ensuring that only the AGA grade and genotype for each patient were compiled. These patients had their genotypes and clinical data securely stored as medical records within a repository incorporating individuals from diverse ethnic backgrounds. Ethics committee approval was not required to perform the current study as all the data derive from medical records generated as an industry by-product and the whole dataset was anonymised as required by the Helsinki Declaration.
Genotyping and SNPs
Briefly, all the genotyping assays were performed by qPCR Taqman Assays (Thermo Fisher Scientific) from genomic DNA previously collected from the patients. The SNPs analysed are summarised on Table 1.
Association Analyses and Statistical Framework
The association between genotype and diagnostic was analysed for patients in the database described previously. The patients’ genotypes were correlated with their clinically diagnosed AGA. Subsequently, the genotypes were analysed in relation to the clinical presentation of AGA. Additional data were also evaluated for significance; we grouped SNPs on genes associated with the molecular mechanisms of drugs used to treat alopecia and performed statistical analyses to evaluate the potential epistatic effect.
Statistical analyses were executed utilising R (version 4.2.2) as the computational environment, with the SNPassoc package selected for the necessary computations [17]. We assessed the genotype–phenotype association without restrictions to the dominance model and implemented Bonferroni correction. In line with the specific study of each SNP, we endeavoured to assess the clinical and molecular significance. Statistical significance was defined as p < 0.01 or p < 0.005. Subsequent to statistical analysis, a comprehensive literature review was carried out to gain further insights into the potential functional relevance of the analysed SNPs.
Results and Discussion
Population Description
The data gathered from the patient database allowed an evaluation of both the proportion of patients diagnosed with alopecia and the corresponding grades, nevertheless, in the further SNP-correlation analyses we did not find any correlation to alopecia grading. The population encompassed patients of both sexes, with ages spanning from 17 to 93 years. Table 2 summarises the composition of the population of patients.
Within the patient data utilised for the present study, a diagnosis of androgenetic alopecia is demonstrated in 64.6% of male and 35.4% of female patients [18, 19]. This proportional distribution aligns consistently with the prevalence patterns found in previously published findings, thus underscoring the representativeness and adequacy of our population sample.
SNP Association Analysis
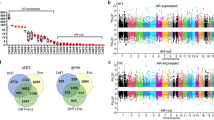
Statistical analysis of the correlation between SNPs and the diagnosis of androgenetic alopecia identified eight SNPs exhibiting statistical significance at a threshold of p < 0.05. A summarised report of this statistical analysis, including the respective p-values, is provided in Table 3.
Figure 1 shows a summarised view of the statistical findings associated with all analysed SNPs evaluated under diverse statistical models.
Age Interaction with the Statistically Significant SNPs
This study conducted a statistical investigation of the interaction between age and the SNPs, yielding a p-value of less than 0.05, considering its critical role in alopecia diagnosis across the entire examined population. Interestingly, the study found no interaction with age for any of the analysed SNPs.
SNPs Associated with the Vasodilation-Related Enzymes
Among the SNPs notably correlated with androgenetic alopecia, the SNPs rs13283456 (PTGES2, p = 0.00005) and rs4343 (ACE, p = 0.0129) have been previously linked to various mechanisms involved in the regulation of the vasodilatory system [20,21,22]. A statistical examination of the correlation between these two SNPs revealed a p-value of 0.0068426, representing a significant trend when rs13283456 occurs within rs4343. This emphasises the influence of functional alterations in genes related to vasodilation control in the pathogenesis of alopecia, a novel insight from our data.
Notably, minoxidil, widely used as an initial treatment for alopecia, exerts its primary therapeutic effect through vasodilation. It acts as a potent arteriolar vasodilator, opening potassium channels on the peripheral artery’s smooth muscles, causing hyperpolarisation of the cell membrane. Minoxidil’s positive effect on hair growth is primarily due to its metabolite, minoxidil sulfate, converted by the enzyme sulfotransferase located in hair follicles [23, 24]. These factors make it likely that patients with alopecia, exhibiting genetic polymorphisms associated with reduced activity of PTGES2 and ACE, might respond effectively to minoxidil. This aligns with clinical observations and supports the hypothesis that genetic factors influencing vasodilation could contribute to variations in treatment response.
SNPs Associated with the Testosterone Metabolism
The three SNPs of interest in genes implicated in testosterone metabolism, namely rs2470152 (CYP19A1), rs39848 (SRD5A1) and rs523349 (SRD5A2), were analysed. However, in accordance with earlier discussions, the only variation associated with the diagnosis of androgenetic alopecia within our investigated population was rs523349, corresponding to the 5-alpha reductase enzyme type 2, with a p-value of 0.00023. The statistical interaction among these SNPs was also examined, but no significant correlation was revealed. It is relevant to highlight that the augmented activity of SRD5A2 is connected to enhanced DHT production. This hormone is renowned for its association with activating its receptors in hair follicle cells, resulting in diminished activity [25,26,27]. This observation also establishes a link to the clinical efficacy of finasteride and dutasteride, as this enzyme plays a vital role in their respective mechanisms of action [28]. Earlier findings had connected the activity of SRD5A2 and the SNP rs523349 to androgenetic alopecia and other alterations related to the metabolism of testosterone [29,30,31].
SNPs Associated with the Prostaglandin D2 Response
Emerging therapies for alopecia, e.g., cetirizine [34, 35] and phytotherapical preparations derived from Nigella sativa [36], are linked to levels and responses to prostaglandin D2. The hormonal responsiveness of PTGDS in other systems and the location of PGD2 receptors in the outer root sheath of the hair follicle further substantiates the integral role of prostaglandins in androgenetic alopecia. The elevated levels of PGD2 may also explain this condition’s notable increase in sebaceous gland size. This suggests that high levels of PGD2 may contribute to sebaceous gland hyperactivity, even though the precise role in alopecia’s pathogenesis is yet to be fully explored.
Two SNPs, specifically rs533116 (p = 0.0164) and rs545659 (p = 0.02996), have been discovered in the prostaglandin D2 receptor. Despite evaluating the interaction between these SNPs revealing no statistical significance with the set threshold (p = 0.054752), these SNPs are noteworthy. They are novel discoveries concerning the genetic predisposition to androgenetic alopecia, and they substantiate mechanisms that have been previously associated with the disease’s pathogenesis [32]. The significance of these SNPs is underlined by the known role of prostaglandins in modulating hair function, including the contrasting functions of PGE2 and PGF2a in promoting hair growth and PGD2 in inhibiting it [32, 33].
Early literature pertaining to the mechanism of minoxidil’s effect on blood pressure reduction indicated that it increases PGE2 levels, known to be diminished in androgenetic alopecia. Even though it is not yet definitively proven, this aligns with the irregular regulation of prostaglandins in AGA. Recent genetic analysis also identified associations with GPR44, approaching statistical significance, which implies the potential role of the PGD2 pathway in alopecia’s pathogenesis. Thus, these findings underscore the importance of the understudied field of lipid biology, specifically prostaglandins and their SNPs, in understanding AGA [32, 33].
SNP Associated with the Prostaglandin F Response
Furthermore, attention was drawn to the SNP rs10782665, located on the prostaglandin F receptor, which demonstrated an association with the diagnosis of androgenetic alopecia (p = 0.01457). This discovery contributes to comprehending the underlying mechanisms involved in the pathogenesis of androgenetic alopecia. Additionally, this particular SNP presents implications for treatment methods, particularly in the context of latanoprost, which acts as an agonist of this receptor [37].
Prostanoids, including prostaglandins, are prominent lipid mediators derived from a serial reaction involving phospholipase A2 and prostaglandin endoperoxide synthases. This subsequently transforms the endoperoxide prostaglandin into specific prostaglandins or thromboxanes. The prostaglandin F2α receptor (FP) is a specific binding site for PGF2α, one of the prostanoids. These prostanoids exert influence over various biological activities, such as regulating cell growth and inflammatory responses, primarily acting as autocrine/paracrine hormones utilising specific G-protein coupled receptors (GPCRs) for downstream signalling. Past research has demonstrated that the metabolism of PGE2 and PGF2α is evident in human hair follicles, suggesting their association with hair growth and differentiation[38]. Thus, the discovered SNP rs10782665 on the prostaglandin F receptor offers valuable insights into the pathogenesis of androgenetic alopecia and prospective treatment strategies [39, 40].
Study Limitation
The current study correlates androgenetic alopecia pathogenesis to canditate genetic variations on genes potentially asociated to the pharmacotherapy of hair loss. The main limitation is related to the amount of genomic targets in the study, we were only looking at specific genes while a further study covering a larger section of the genome should be conducted to find more targets.
Conclusions
The analysis of a range of SNPs in relation to the diagnosis of androgenetic alopecia provides valuable insights that enhance our understanding of the disease’s pathogenesis and prospective therapeutic approaches. The SNP rs1800012 (COL1A1), gene encoding a type I collagen chain [41,42,43], is related to the structure of collagen fibre. Additionally, the SNP rs12724719, linked to CRABP2, which is associated with Vitamin A deficiency, has been identified [44]. Both these SNPs have revealed statistically significant correlations to alopecia and might represent potential targets for therapy. Variations in the vasodilatory system and the metabolism of testosterone, such as the ones identified for the genes PTGES2, ACE and SRD5A2, offer insights into the underlying mechanisms of the disease and, further, the identification of the SNPs linked to prostaglandin receptors, as is the case for PTGFR, expands our understanding of potential genetic influences in the therapeutic response to alopecia [28, 29, 37, 38].
The association of multiple SNPs with the disease underscores the increasing complexity of the genetic network that contributes to androgenetic alopecia. The findings in this study augment the growing body of evidence, suggesting that a more nuanced appreciation of genetic contributions may offer a promising path to understanding the mechanisms underlying alopecia’s pathogenesis. Furthermore, such an understanding could guide more tailored therapies, such as those targeting the prostaglandin D2 receptor or the prostaglandin F receptor. The genetic variation identified within these pathways may contribute to the varying responsiveness to treatments observed in clinical practice [32,33,34,35, 38, 39].
The prospect of employing genetic information to guide therapeutic strategies is an emerging field in the treatment of alopecia. The results indicate that specific genetic markers can influence the pathogenesis of androgenetic alopecia and potentially the responsiveness to treatment. However, this preliminary analysis should be considered a stepping stone in exploring this complex disease. Further studies are necessary to validate the potential for genetic evaluations in guiding therapy for alopecia. While we are still far from a comprehensive understanding of the interplay of genetic and environmental factors in the pathogenesis and treatment of alopecia, our findings provide information on the potential benefits of employing genetic information in the pharmacotherapy of androgenetic alopecia.
References
Tosti A, Piraccini BM, Iorizzo M, Voudouris S. The natural history of androgenetic alopecia. J Cosmet Dermatol. 2005;4:41–3.
Devjani S, Ezemma O, Kelley KJ, Stratton E, Senna M. Androgenetic alopecia: therapy update. Drugs. 2023;83:701–15.
Adil A, Godwin M. The effectiveness of treatments for androgenetic alopecia: a systematic review and meta-analysis. J Am Acad Dermatol. 2017;77:136-141.e5.
Dominguez-Santas M, Diaz-Guimaraens B, Saceda-Corralo D, Hermosa-Gelbard A, Muñoz-Moreno Arrones O, Pindado-Ortega C, et al. The state-of-the-art in the management of androgenetic alopecia: a review of new therapies and treatment algorithms. JEADV Clin Pract. 2022;1:176–85.
Rundegren J. A one-year observational study with minoxidil 5% solution in Germany: results of independent efficacy evaluation by physicians and patients. J Am Acad Dermatol. 2004;50:P91.
Lolli F, Pallotti F, Rossi A, Fortuna MC, Caro G, Lenzi A, et al. Androgenetic alopecia: a review. Endocrine. 2017;57:9–17.
Mella JM, Perret MC, Manzotti M, Catalano HN, Guyatt G. Efficacy and safety of finasteride therapy for androgenetic alopecia. Arch Dermatol. 2010;146:1141–50.
Kaufman KD, Olsen EA, Whiting D, Savin R, DeVillez R, Bergfeld W, et al. Finasteride in the treatment of men with androgenetic alopecia. J Am Acad Dermatol. 1998;39:578–89.
Trüeb RM. Molecular mechanisms of androgenetic alopecia. Exp Gerontol. 2002;37:981–90.
Tcherepanova I, Puigserver P, Norris JD, Spiegelman BM, McDonnell DP. Modulation of estrogen receptor-α transcriptional activity by the coactivator PGC-1. J Biol Chem. 2000;275:16302–8.
Ho BS-Y, Vaz C, Ramasamy S, Chew EGY, Mohamed JS, Jaffar H, et al. Progressive expression of PPARGC1α is associated with hair miniaturization in androgenetic alopecia. Sci Rep. 2019;9:8771.
Saha AK, Persons K, Safer JD, Luo Z, Holick MF, Ruderman NB. AMPK regulation of the growth of cultured human keratinocytes. Biochem Biophys Res Commun. 2006;349:519–24.
Lee MJ, Cha HJ, Lim KM, Lee O-K, Bae S, Kim C-H, et al. Analysis of the microRNA expression profile of normal human dermal papilla cells treated with 5α-dihydrotestosterone. Mol Med Rep. 2015;12:1205–12.
Karnik P, Tekeste Z, McCormick TS, Gilliam AC, Price VH, Cooper KD, et al. Hair follicle stem cell-specific PPARγ deletion causes scarring alopecia. J Investig Dermatol. 2009;129:1243–57.
Heilmann S, Kiefer AK, Fricker N, Drichel D, Hillmer AM, Herold C, et al. Androgenetic alopecia: identification of four genetic risk loci and evidence for the contribution of WNT signaling to its etiology. J Investig Dermatol. 2013;133:1489–96.
Li R, Brockschmidt FF, Kiefer AK, Stefansson H, Nyholt DR, Song K, et al. Six novel susceptibility loci for early-onset androgenetic alopecia and their unexpected association with common diseases. PLoS Genet. 2012;8: e1002746.
González JR, Armengol L, Solé X, Guinó E, Mercader JM, Estivill X, et al. SNPassoc: an R package to perform whole genome association studies. Bioinformatics. 2007;23:654–5.
Aukerman EL, Jafferany M. The psychological consequences of androgenetic alopecia: a systematic review. J Cosmet Dermatol. 2023;22:89–95.
Salman KE, Altunay IK, Kucukunal NA, Cerman AA. Frequency, severity and related factors of androgenetic alopecia in dermatology outpatient clinic: hospital-based cross-sectional study in Turkey. An Bras Dermatol. 2017;92:35–40.
Niu W, Qi Y, Gao P, Zhu D. Review: Association between angiotensin converting enzyme G2350A polymorphism and hypertension risk: a meta-analysis. J Renin Angiotensin Aldosterone Syst. 2011;12:8–14.
Fischer A, Grallert H, Böhme M, Gieger C, Boomgaarden I, Heid I, et al. Association analysis between the prostaglandin E synthase 2 R298H polymorphism and body mass index in 8079 participants of the KORA study cohort. Genet Test Mol Biomark. 2009;13:223–6.
Zhang Y. From gene variants to novel therapies. Is the prostaglandin E2 pathway in primary graft dysfunction ready for prime time? Am J Respir Crit Care Med. 2014;189:507–8.
Messenger AG, Rundegren J. Minoxidil: mechanisms of action on hair growth. Br J Dermatol. 2004;150:186–94.
Suchonwanit P, Thammarucha S, Leerunyakul K. Minoxidil and its use in hair disorders: a review. Drug Des Dev Ther. 2019;13:2777–86.
Chen X, Liu B, Li Y, Han L, Tang X, Deng W, et al. Dihydrotestosterone regulates hair growth through the Wnt/β-catenin pathway in C57BL/6 mice and in vitro organ culture. Front Pharmacol. 2020;10.
Ustuner ET. Cause of androgenic alopecia. Plast Reconstr Surg Glob Open. 2013;1: e64.
Urysiak-Czubatka I, Kmieć ML, Broniarczyk-Dyła G. Assessment of the usefulness of dihydrotestosterone in the diagnostics of patients with androgenetic alopecia. Adv Dermatol Allergol. 2014;4:207–15.
Xiao Q, Wang L, Supekar S, Shen T, Liu H, Ye F, et al. Structure of human steroid 5α-reductase 2 with the anti-androgen drug finasteride. Nat Commun. 2020;11:5430.
Li X, Huang Y, Fu X, Chen C, Zhang D, Yan L, et al. Meta-analysis of three polymorphisms in the steroid-5-alpha-reductase, alpha polypeptide 2 gene (SRD5A2) and risk of prostate cancer. Mutagenesis. 2011;26:371–83.
Hayes VM, Severi G, Padilla EJD, Morris HA, Tilley WD, Southey MC, et al. 5α-Reductase type 2 gene variant associations with prostate cancer risk, circulating hormone levels and androgenetic alopecia. Int J Cancer. 2007;120:776–80.
Zeng X-T, Su X-J, Li S, Weng H, Liu T-Z, Wang X-H. Association between SRD5A2 rs523349 and rs9282858 polymorphisms and risk of benign prostatic hyperplasia: a meta-analysis. Front Physiol. 2017;8.
Garza LA, Liu Y, Yang Z, Alagesan B, Lawson JA, Norberg SM, et al. Prostaglandin D2 inhibits hair growth and is elevated in bald scalp of men with androgenetic alopecia. Sci Transl Med. 2012;4.
Nieves A, Garza LA. Does prostaglandin D2 hold the cure to male pattern baldness? Exp Dermatol. 2014;23:224–7.
Chen X, Xiang H, Yang M. Topical cetirizine for treating androgenetic alopecia: a systematic review. J Cosmet Dermatol. 2022;21:5519–26.
Hossein Mostafa D, Samadi A, Niknam S, Nasrollahi SA, Guishard A, Firooz A. Efficacy of cetirizine 1% versus minoxidil 5% topical solution in the treatment of male alopecia: a randomized, single-blind controlled study. J Pharm Pharm Sci. 2021;24:191–9.
Rossi A, Priolo L, Iorio A, Vescarelli E, Gerardi M, Campo D, et al. Evaluation of a therapeutic alternative for telogen effluvium: a pilot study. J Cosmet Dermatol Sci Appl. 2013;03:9–16.
Cordeiro MF, Gandolfi S, Gugleta K, Normando EM, Oddone F. How latanoprost changed glaucoma management. Acta Ophthalmol. 2023;e140–55.
Shin DW. The physiological and pharmacological roles of prostaglandins in hair growth. Korean J Physiol Pharmacol. 2022;26:405–13.
Rafati M, Mahmoudian R, Golpour M, Kazeminejad A, Saeedi M, Nekoukar Z. The effect of latanoprost 0.005% solution in the management of scalp alopecia areata, a randomized double-blind placebo-controlled trial. Dermatol Ther. 2022;35:e15450.
Bloch LD, Escudeiro CC, Sarruf FD, Valente NYS. Latanoprosta e minoxidil: Estudo duplocego comparativo, placebo-controlado no tratamento da queda de cabelos. Surg Cosmet Dermatol. 2018;10:41–5.
Cross DS, Ivacic LC, McCarty CA. Development of a fingerprinting panel using medically relevant polymorphisms. BMC Med Genom. 2009;2:17.
Long J-R, Zhao L-J, Liu P-Y, Lu Y, Dvornyk V, Shen H, et al. Patterns of linkage disequilibrium and haplotype distribution in disease candidate genes. BMC Genet. 2004;5:11.
Rojano-Mejía D, Coral-Vázquez RM, Espinosa LC, López-Medina G, Aguirre-García MC, Coronel A, et al. JAG1 and COL1A1 polymorphisms and haplotypes in relation to bone mineral density variations in postmenopausal Mexican-Mestizo Women. Age (Omaha). 2013;35:471–8.
Manolescu DC, El-Kares R, Lakhal-Chaieb L, Montpetit A, Bhat PV, Goodyer P. Newborn serum retinoic acid level is associated with variants of genes in the retinol metabolism pathway. Pediatr Res. 2010;67:598–602.
Medical Writing and Editorial Assistance
The authors did not use any medical writing or editorial assistance for this article.
Author Contributions
M.P.F. was responsible for the data organisation and study idealisation. L.V.-V. was involved in the manuscript writing. V.R. was responsible for evaluating genotype data and quality control. H.C.P. was responsible for manuscript writing and data analysis. G.T.S. was responsible data cleaning, statistical analysis, bioinformatics, manuscript writing and scientific conceptualisation.
Funding
There was no direct funding for the study. Fagron BV is responsible for the payment of the journal’s Rapid Service Fee.
Data Availability
The datasets generated and/or analysed during the current study are composed of fully anonymised data and are available upon request.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
M.P.F., L.V.-V., V.R., H.C.P. and G.T.S. declare no conflicts of interest.
Ethical Approval
Ethics committee approval was not required to perform the current study as all the data derive from medical records generated as an industry by-product and the whole dataset was anonymised as required by the Helsinki Declaration. Permission to access and use the database was granted by the owner of the database, and the patients grant access to analyse and store their data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Francès, M.P., Vila-Vecilla, L., Russo, V. et al. Utilising SNP Association Analysis as a Prospective Approach for Personalising Androgenetic Alopecia Treatment. Dermatol Ther (Heidelb) 14, 971–981 (2024). https://doi.org/10.1007/s13555-024-01142-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13555-024-01142-y