Abstract
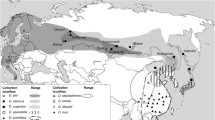
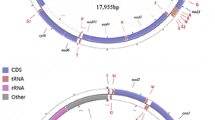
In China, there are two subspecies of the chestnut tiger butterfly, Parantica sita (Kollar) (Lepidoptera: Nymphalidae), P. s. sita in southwest China and P. s. niphonica on Taiwan Island. Our previous study found substantial genetic divergence between both subspecies. This paper reports a novel haplotype of the mitochondrial genome from the specimens in Leshan, Sichuan Province, China (P. sita LS), which is located within the distribution range of P. s. sita. The length of the complete mitochondrial genome is 15,202 bp, and its general characteristics match those published before. However, genetically, P. sita LS diverges substantially from both P. s. sita and P. s. niphonica. First, the COI gene of P. sita LS differs from the COI genes of P. s. sita and P. s. niphonica by 4.7% and 2.5%, respectively. Second, consistent with the difference between P. s. sita and P. s. niphonica, the AT-rich region of P. sita LS shows a fragment deletion. Third, these three haplotypes of P. sita are not forming a monophyly in phylogenetic trees. Instead, P. luzonensis, a related species, mixes in the cluster of P. sita. Based on these results, this paper proposes the division of P. sita into three independent species, which needs to be corroborated by further evidence.




Similar content being viewed by others
References
Ackery PR, Vane-Wright RI (1984) Milkweed butterflies, their cladistics and biology, being an account of the natural history of the Danainae, a subfamily of the Lepidoptera, Nymphalidae. British Museum (Natural History), London
Ballard JWO, Whitlock MC (2004) The incomplete natural history of mitochondria. Mol Ecol 13:729–744. https://doi.org/10.1046/j.1365-294X.2003.02063.x
Basset Y, Eastwood R, Sam L, Lohman DJ, Novotny V, Treuer T, Miller SE, Weiblen GD, Pierce NE, Bunyavejchewin S, Sakchoowong W, Kongnoo P, Osorio-Arenas MA (2011) Comparison of rainforest butterfly assemblages across three biogeographical regions using standardized protocols. J Res Lepid 44:17–28
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Boore JL (1999) Animal mitochondrial genomes. Nucleic Acids Res 27:1767–1780. https://doi.org/10.1093/nar/27.8.1767
Borchers TE, Marcus JM (2014) Genetic population structure of buckeye butterflies (Junonia) from Argentina. Syst Ent 39:242–255. https://doi.org/10.1111/syen.12053
Brower AVZ, Wahlberg N, Ogawa JR, Boppre M, Vane-Wright RI (2010) Phylogenetic relationships among genera of danaine butterflies (Lepidoptera: Nymphalidae) as implied by morphology and DNA sequences. Syst Biodivers 8:75–89. https://doi.org/10.1080/14772001003626814
Cameron SL (2014) Insect mitochondrial genomics: implications for evolution and phylogeny. Annu Rev Entomol 59:95–117. https://doi.org/10.1146/annurev-ento-011613-162007
Cameron SL, Whiting MF (2008) The complete mitochondrial genome of the tobacco hornworm, Manduca sexta, (Insecta: Lepidoptera: Sphingidae), and an examination of mitochondrial gene variability within butterflies and moths. Gene 408:112–123. https://doi.org/10.1016/j.gene.2007.10.023
Chen L, Wahlberg N, Liao CQ, Wang CB, Ma FZ, Huang GH (2020) Fourteen complete mitochondrial genomes of butterflies from the genus Lethe (Lepidoptera, Nymphalidae, Satyrinae) with mitogenome-based phylogenetic analysis. Genomics 112:4435–4441. https://doi.org/10.1016/j.ygeno.2020.07.042
Chou I (1994) Monographia rhopalocerorum sinensium (monograph of Chinese butterflies). Henan Scientific and Technological Press, Zhengzhou, pp 276–277
Dierckxsens N, Mardulyn P, Smits G (2017) NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res. https://doi.org/10.1093/nar/gkw955
Donath A, Juhling F, Al-Arab M, Bernhart SH, Reinhardt F, Stadler PF, Middendorf M, Bernt M (2019) Improved annotation of protein-coding genes boundaries in metazoan mitochondrial genomes. Nucleic Acids Res 47:10543–10552. https://doi.org/10.1093/nar/gkz833
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Garey JR, Wolstenholme DR (1989) Platyhelminth mitochondrial-DNA: evidence for early evolutionary origin of a tRNA (serAGN) that contains a dihydrouridine arm replacement loop, and of serine-specifying AGA and AGG codons. J Mol Evol 28:374–387. https://doi.org/10.1007/bf02603072
Gonçalves R, Freitas AI, Jesus J, De la Rua P, Brehm A (2015) Structure and genetic variation of the mitochondrial control region in the honey bee Apis mellifera. Apidologie 46:515–526. https://doi.org/10.1007/s13592-014-0341-y
Hebert PDN, Ratnasingham S, deWaard JR (2003) Barcoding animal life: cytochrome c oxidase subunit 1 divergences among closely related species. P Roy Soc B-Biol Sci 270:S96–S99. https://doi.org/10.1098/rsbl.2003.0025
Hoang DT, Chernomor O, von Haeseler A, Minh BQ, Vinh LS (2018) UFBoot2: Improving the ultrafast bootstrap approximation. Mol Biol Evol 35:518–522. https://doi.org/10.1093/molbev/msx281
Hu P, Wang R (2019) The complete mitochondrial genome of Parantica sita sita (Lepidoptera: Nymphalidae: Danainae) revealing substantial genetic divergence from its sibling subspecies P. s. niphonica. Gene 686:76–84. https://doi.org/10.1016/j.gene.2018.10.088
Joshi J, Prakash A, Kunte K (2017) Evolutionary assembly of communities in butterfly mimicry rings. Am Nat 189:E58–E76. https://doi.org/10.1086/690907
Kanazawa I, Cheng WWW, Pun HSF, Sakiyama Y, Doi H (2015) First migration record of chestnut tiger butterfly, Parantica sita niphonica (Moore, 1883) (Lepidoptera: Nymphalidae: Danainae) from Japan to Hong Kong and longest recorded movement by the species. Pan-Pac Entomol 91:91–97. https://doi.org/10.3956/2014-91.1.091
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Kim MI, Wan X, Kim MJ, Jeong HC, Ahn NH, Kim KG, Han YS, Kim I (2010) Phylogenetic relationships of true butterflies (Lepidoptera: Papilionoidea) inferred from COI, 16S rRNA and EF-1 alpha sequences. Mol Cells 30:409–425. https://doi.org/10.1007/s10059-010-0141-9
Lam-Tung N, Schmidt HA, von Haeseler A, Bui Quang M (2015) IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Lanfear R, Frandsen PB, Wright AM, Senfeld T, Calcott B (2017) PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol Biol Evol 34:772–773. https://doi.org/10.1093/molbev/msw260
Letunic I, Bork P (2021) Interactive tree of life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 49:W293–W293. https://doi.org/10.1093/nar/gkab301
Li X, Wang Y, Rong W, Liu X, Li R (2020) A new record of complete mitochondrial genome of Parantica aglea (Lepidoptera: Danainae) and its phylogeny. Mitochondrial DNA B Resour 5:1257–1258. https://doi.org/10.1080/23802359.2020.1731367
Nikolaou C, Almirantis Y (2005) A study on the correlation of nucleotide skews and the positioning of the origin of replication: different modes of replication in bacterial species. Nucleic Acids Res 33:6816–6822. https://doi.org/10.1093/nar/gki988
Ohtsuki T, Kawai G, Watanabe K (2002) The minimal tRNA: unique structure of Ascaris suum mitochondrial tRNA(UCU)(Ser) having a short T arm and lacking the entire D arm. FEBS Lett 514:37–43. https://doi.org/10.1016/s0014-5793(02)02328-1
Paradis E, Schliep K (2019) ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35:526–528. https://doi.org/10.1093/bioinformatics/bty633
R Core Team (2020) R: a language and environment for statistical computing. https://www.R-project.org. Accessed 10 Oct 2020
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Hohna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) Mrbayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Sperling FAH, Hickey DA (1994) Mitochondrial-DNA sequence variation in the spruce budworm species complex (Choristoneura, Lepidoptera). Mol Biol Evol 11:656–665. https://doi.org/10.1093/oxfordjournals.molbev.a040144
Sperling F, Byers R, Hickey D (1996) Mitochondrial DNA sequence variation among pheromotypes of the dingy cutworm, Feltia jaculifera (Gn) (Lepidoptera:Noctuidae). Can J Zool 74:2109–2117. https://doi.org/10.1139/z96-240
Taylor MFJ, McKechnie SW, Pierce N, Kreitman M (1993) The Lepidopteran mitochondrial control region: structure and evolution. Mol Biol Evol 10:1259–1272. https://doi.org/10.1093/oxfordjournals.molbev.a040075
Wahlberg N, Leneveu J, Kodandaramaiah U, Pena C, Nylin S, Freitas AVL, Brower AVZ (2009) Nymphalid butterflies diversify following near demise at the Cretaceous/Tertiary boundary. P Roy Soc B-Biol Sci 276:4295–4302. https://doi.org/10.1098/rspb.2009.1303
Wang X, Chen ZM, Gu XS, Wang M, Huang GH, Zwick A (2019) Phylogenetic relationships among Bombycidae s.l. (Lepidoptera) based on analyses of complete mitochondrial genomes. Syst Entomol 44:490–498. https://doi.org/10.1111/syen.12337
Whinnett A, Brower AVZ, Lee MM, Willmott KR, Mallet J (2005) Phylogenetic utility of Tektin, a novel region for inferring systematic relationships among Lepidoptera. Ann Entomol Soc Am 98:873–886. https://doi.org/10.1603/0013-8746(2005)098[0873:Puotan]2.0.Co;2
Wilson JJ, Sing KW, Sofian-Azirun M (2013) Building a DNA barcode reference library for the true butterflies (Lepidoptera) of Peninsula Malaysia: what about the subspecies? PLoS One. https://doi.org/10.1371/journal.pone.0079969
Wolstenholme DR (1992) Animal mitochondrial-DNA: structure and evolution. Int Rev Cytol 141:173–216. https://doi.org/10.1016/s0074-7696(08)62066-5
Wu L, Lin L, Lees DC, Hsu Y (2014) Mitogenomic sequences effectively recover relationships within brush-footed butterflies (Lepidoptera: Nymphalidae). BMC Genomics. https://doi.org/10.1186/1471-2164-15-468
Zhang DX, Hewitt GM (1997) Insect mitochondrial control region: a review of its structure, evolution and usefulness in evolutionary studies. Biochem Syst Ecol 25:99–120. https://doi.org/10.1016/s0305-1978(96)00042-7
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 31772446). Sample collection in the field was supported by Mr. Fei Xiong. We especially thank Dr. Wei Zhang from Peking University for assistance with sequencing and data analysis, and Yuping Meng for routine laboratory work.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhang, Y., Lu, L., Hu, P. et al. A novel mitochondrial genome haplotype in Parantica sita sita (Lepidoptera: Nymphalidae: Danainae) indicates substantial intraspecific genetic divergence. Appl Entomol Zool 57, 109–118 (2022). https://doi.org/10.1007/s13355-022-00772-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13355-022-00772-y