Abstract
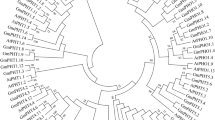
Nitrate transporters (NRTs) are important channel proteins facilitating cross-membrane movement of small molecules like NO3− which is a critical nutrient for all life. However, the classification and evolution of nitrate transporters in the legume plants are still elusive. In this study, we surveyed the wild soybean (G. soja) genomic databases and identified 120 GsNRT1 and 5 GsNRT2 encoding genes. Phylogenetic analyses show that GsNRT1 subfamily is consisted of eight clades (NPF1 to NPF8), while GsNRT2 subfamily has only one clade. Gene chromosomal location and evolutionary historic analyses indicate that GsNRT genes are unevenly distributed on 19 out of 20 G. soja chromosomes and segmental duplications may take a major part in the expansion of GsNRT family. Investigations of gene structure and protein motif compositions suggest that GsNRT family members are highly conserved in structures of both gene and protein levels. In addition, we analyzed the spatial expression patterns of representative GsNRT genes and their responses to exogenous nitrogen and carbon supplies and different abiotic stresses. The qRT-PCR data indicated that 16 selected GsNRT genes showed various expression levels in the roots, stems, leaves, and pods of young G. soja plants, and these genes were regulated by not only nitrogen and carbohydrate nutrients but also NaCl, NaHCO3, abscisic acid (ABA), and salicylic acid (SA). These results suggest that GsNRT genes may be involved in the regulation of plant growth, development, and adaptation to environmental stresses, and the study will shed light on functional dissection of plant nitrate transporter proteins in the future.








Similar content being viewed by others
Abbreviations
- CDS:
-
Coding sequence
- CHL1:
-
Chlorate resistant mutant 1
- NRT:
-
Nitrate transporter
- MFS:
-
Major facilitator superfamily
- HMM:
-
Hidden Markov model
- TD:
-
Tandem duplicated
- SD:
-
Segmental duplicated
- Chr:
-
Chromosome
References
Amarasinghe BH, de Bruxelles GL, Braddon M, Onyeocha I, Forde BG, Udvardi MK (1998) Regulation of GmNRT2 expression and nitrate transport activity in roots of soybean. (Glycine max). Planta 206(1):44–52 https://doi.org/10.1007/s004250050372
Anwar A, Li Y, He C, Yu X (2019) 24-Epibrassinolide promotes NO3- and NH4+ ion flux rate and NRT1 gene expression in cucumber under suboptimal root zone temperature. BMC Plant Biol 19(1):225. https://doi.org/10.1186/s12870-019-1838-3
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME Suite: tools for motif discovery and searching. Nucleic Acids Res 37(suppl_2):W202–W208. https://doi.org/10.1093/nar/gkp335
Bi C, Xu Y, Ye Q, Yin T, Ye N (2016) Genome-wide identification and characterization of WRKY gene family in Salix suchowensis. Peer J 4(9):e2437. https://doi.org/10.7717/peerj.2437
Crawford NM (1995) Nitrate: nutrient and signal for plant growth. Plant Cell 7(7):859–868. https://doi.org/10.1105/tpc.7.7.859
Fan X, Naz M, Fan X, Xuan W, Miller AJ, Xu G (2017) Plant nitrate transporters: from gene function to application. J Exp Bot 68(10):2463–2475. https://doi.org/10.1093/jxb/erx011
Glass AD, Shaff JE, Kochian LV (1992) Studies of the uptake of nitrate in barley : IV. Electrophysiology. Plant Physiol 99(2):456–463. https://doi.org/10.1104/pp.99.2.456
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59(3):307–321. https://doi.org/10.1093/sysbio/syq010
Guo Q, Turnbull MH, Song J, Roche J, Novak O, Spath J, Jameson PE, Love J (2017) Depletion of carbohydrate reserves limits nitrate uptake during early regrowth in Lolium perenne L. J Exp Bot 68(7):1569–1583. https://doi.org/10.1093/jxb/erx056
He QH, Qiao DR, Zhang QL, Li Y, Xu H, Wei L, Gu Y, Cao Y (2004) Cloning and expression study of a putative high-affinity nitrate transporter gene from Dunaliella salina. J Appl Phycol 16(5):395–400. https://doi.org/10.1023/B:JAPH.0000047950.76549.ce
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G (2014) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31(8):1296. https://doi.org/10.1093/bioinformatics/btu817
Kanno Y, Hanada A, Chiba Y, Ichikawa T, Nakazawa M, Matsui M, Koshiba T, Kamiya Y, Seo M (2012) Identification of an abscisic acid transporter by functional screening using the receptor complex as a sensor. Proc Natl Acad Sci U S A 109(24):9653–9658. https://doi.org/10.1073/pnas.1203567109]
Kazutaka K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molecular Biology & Evolution 30(4):772–780. https://doi.org/10.1093/molbev/mst010
Kiba T, Feria-Bourrellier AB, Lafouge F, Lezhneva L, Boutet-Mercey S, Orsel M, Brehaut V, Miller A, Daniel-Vedele F, Sakakibara H, Krapp A (2012) The Arabidopsis nitrate transporter NRT2.4 plays a double role in roots and shoots of nitrogen-starved plants. Plant Cell 24(1):245–258. https://doi.org/10.1105/tpc.111.092221
Kim MY, Lee S, Van K, Kim T-H, Jeong S-C, Choi I-Y, Kim D-S, Lee Y-S, Park D, Ma J (2010) Whole-genome sequencing and intensive analysis of the undomesticated soybean (Glycine soja Sieb. and Zucc.) genome. Proc Natl Acad Sci U S A 107(51):22032–22037. https://doi.org/10.1073/pnas.1009526107
Komarova NY, Thor K, Gubler A, Meier S, Dietrich D, Weichert A, Suter Grotemeyer M, Tegeder M, Rentsch D (2008) AtPTR1 and AtPTR5 transport dipeptides in planta. Plant Physiol 148(2):856–869. https://doi.org/10.1104/pp.108.123844
Lefort V, Longueville JE, Gascuel O (2017) SMS: smart model selection in PhyML. Mol Biol Evol 34(9):2422–2424. https://doi.org/10.1093/molbev/msx149
Léran S, Varala K, Boyer JC, Chiurazzi M, Crawford N, Daniel-Vedele F, David L, Dickstein R, Fernandez E, Forde B (2014) A unified nomenclature of nitrate transporter 1/peptide transporter family members in plants. Trends Plant Sci 19(1):5–9. https://doi.org/10.1016/j.tplants.2013.08.008
Li Z, Zhang C, Guo Y, Niu W, Wang Y, Xu Y (2017) Evolution and expression analysis reveal the potential role of the HD-Zip gene family in regulation of embryo abortion in grapes (Vitis vinifera L.). BMC Genomics 18(1):744. https://doi.org/10.1186/s12864-017-4110-y
Little DY, Rao H, Oliva S, Daniel-Vedele F, Krapp A, Malamy JE (2005) The putative high-affinity nitrate transporter NRT2.1 represses lateral root initiation in response to nutritional cues. Proc Natl Acad Sci U S A 102(38):13693–13698. https://doi.org/10.1073/pnas.0504219102
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Migocka M, Warzybok A, Kłobus G (2013) The genomic organization and transcriptional pattern of genes encoding nitrate transporters 1 (NRT1) in cucumber. Plant Soil 364(1–2):245–260. https://doi.org/10.1007/s11104-012-1345-x
Mikita S, David T, Peer B (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Research 34(Web Server issue):W609. https://doi.org/10.1093/nar/gkl315
Pellizzaro A, Clochard T, Cukier C, Bourdin C, Juchaux M, Montrichard F, Thany S, Raymond V, Planchet E, Limami AM, Morere-Le Paven MC (2014) The nitrate transporter MtNPF6.8 (MtNRT1.3) transports abscisic acid and mediates nitrate regulation of primary root growth in Medicago truncatula. Plant Physiol 166(4):2152–2165. https://doi.org/10.1104/pp.114.250811
Pellizzaro A, Alibert B, Planchet E, Limami AM, Morere-Le Paven MC (2017) Nitrate transporters: an overview in legumes. Planta 246(4):585–595. https://doi.org/10.1007/s00425-017-2724-6
Song Y, Zhang H, You H, Liu Y, Chen C, Feng X, Yu X, Wu S, Wang L, Zhong S, Li Q, Zhu Y, Ding X (2019) Identification of novel interactors and potential phosphorylation substrates of GsSnRK1 from wild soybean (Glycine soja). Plant Cell Environ 42(1):145–157. https://doi.org/10.1111/pce.13217
Stitt M (1999) Nitrate regulation of metabolism and growth. Curr Opin Plant Biol 2(3):178–186. https://doi.org/10.1016/S1369-5266(99)80033-8
Taulemesse F, Le Gouis J, Gouache D, Gibon Y, Allard V (2015) Post-flowering nitrate uptake in wheat is controlled by N status at flowering, with a putative major role of root nitrate transporter NRT2.1. PLoS One 10(3):e0120291. https://doi.org/10.1371/journal.pone.0120291
Tong Y, Zhou JJ, Li ZS, Miller AJ (2005) A two-component high-affinity nitrate uptake system in barley. Plant J l 41(3):442–450. https://doi.org/10.1111/j.1365-313X.2004.02310.x
Tsay YF, Schroeder JI, Feldmann KA, Crawford NM (1993) The herbicide sensitivity gene CHL1 of Arabidopsis encodes a nitrate-inducible nitrate transporter. Cell 72(5):705–713. https://doi.org/10.1016/0092-8674(93)90399-B
Tsay YF, Chiu CC, Tsai CB, Ho CH, Hsu PK (2007) Nitrate transporters and peptide transporters. FEBS Lett 581(12):2290–2300. https://doi.org/10.1016/j.febslet.2007.04.047
Wang YY, Tsay YF (2011) Arabidopsis nitrate transporter NRT1.9 is important in phloem nitrate transport. Plant Cell 23(5):1945–1957. https://doi.org/10.1105/tpc.111.083618
Wang YY, Hsu PK, Tsay YF (2012) Uptake, allocation and signaling of nitrate (vol 17, pg 458, 2012). Trends Plant Sci 17(10):624–624. https://doi.org/10.1016/j.tplants.2012.08.007
Wang L, Hu W, Sun J, Liang X, Yang X, Wei S, Wang X, Zhou Y, Xiao Q, Yang G (2015) Genome-wide analysis of SnRK gene family in Brachypodium distachyon and functional characterization of BdSnRK2.9. Plant Sci 237:33–45. https://doi.org/10.1016/j.plantsci.2015.05.008
Xie M, Chung CY, Li MW, Wong FL, Wang X, Liu A, Wang Z, Leung AK, Wong TH, Tong SW, Xiao Z, Fan K, Ng MS, Qi X, Yang L, Deng T, He L, Chen L, Fu A, Ding Q, He J, Chung G, Isobe S, Tanabata T, Valliyodan B, Nguyen HT, Cannon SB, Foyer CH, Chan TF, Lam HM (2019) A reference-grade wild soybean genome. Nat Commun 10(1):1216. https://doi.org/10.1038/s41467-019-09142-9
Xu G, Guo C, Shan H, Kong H (2012) Divergence of duplicate genes in exon-intron structure. Proc Natl Acad Sci U S A 109(4):1187–1192. https://doi.org/10.1073/pnas.1109047109
Xu L, Shen Z-L, Chen W, Si G-Y, Meng Y, Guo N, Sun X, Cai Y-P, Lin Y, Gao J-S (2019) Phylogenetic analysis of upland cotton MATE gene family reveals a conserved subfamily involved in transport of proanthocyanidins. Mol Biol Rep 46(1):161–175. https://doi.org/10.1007/s11033-018-4457-4
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Molecular Biology & Evolution 24(8):1586–1591. https://doi.org/10.1093/molbev/msm088
Zhang H, Jennings A, Barlow PW, Forde BG (1999) Dual pathways for regulation of root branching by nitrate. Proc Natl Acad Sci U S A 96(11):6529–6534. https://doi.org/10.1073/pnas.96.11.6529
Zhang GB, Yi HY, Gong JM (2014) The Arabidopsis ethylene/jasmonic acid-NRT signaling module coordinates nitrate reallocation and the trade-off between growth and environmental adaptation. Plant Cell 26(10):3984–3998. https://doi.org/10.1105/tpc.114.129296
Zhou JJ, Theodoulou FL, Muldin I, Ingemarsson B, Miller AJ (1998) Cloning and functional characterization of a Brassica napus transporter that is able to transport nitrate and histidine. J Biol Chem 273(20):12017–12023. https://doi.org/10.1074/jbc.273.20.12017
Acknowledgments
We thank Dr. Victor Manon for critical reading of this manuscript.
Funding
This work was financially supported by the National Natural Science Foundation of China (31670272 to X.D.), Natural Science Foundation of Heilongjiang Province (C2017014 to X.D.), Study Abroad for Young Scholar Program of China Scholarship Council (to Q.L.), and the Starting Fund of Northeast Agricultural University (to X.D.).
Author information
Authors and Affiliations
Contributions
Experiment design: Q.L., X.D., Y.L. Experiment performance and data analyses: Y.L., H.Y., H.L., P.Z., T.N.M., Q.L. Data interpretation: X.D., W.L., J.X. Manuscript drafting: Q.L., X.D., Y.L.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Research involving human participants and/or animals
Not Applicable
Informed consent
Not Applicable
Additional information
Communicated by: Izabela Pawłowicz
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Fig. S1
The Python scripts for peptide length, molecular weight_and pI calculation (PDF 18052 kb)
Fig. S2
Distribution of intron numbers in different GsNRT genes. (PDF 3178 kb)
Fig. S3
The details of conserved motifs for the GsNRT family. (PDF 3275 kb)
Fig. S4
Transcriptomic analyses of GsNRT genes in different G. soja tissues. (PDF 19933 kb)
ESM 1
(DOC 317 kb)
Rights and permissions
About this article
Cite this article
You, H., Liu, Y., Minh, T.N. et al. Genome-wide identification and expression analyses of nitrate transporter family genes in wild soybean (Glycine soja). J Appl Genetics 61, 489–501 (2020). https://doi.org/10.1007/s13353-020-00571-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-020-00571-7