Abstract
Strategies targeting nucleolin have enabled a significant improvement in intracellular bioavailability of their encapsulated payloads. In this respect, assessment of the impact of target cell heterogeneity and nucleolin homology across species (structurally and functionally) is of major importance. This work also aimed at mathematically modelling the nucleolin expression levels at the cell membrane, binding and internalization of pH-sensitive pegylated liposomes encapsulating doxorubicin and functionalized with the nucleolin-binding F3 peptide (PEGASEMP), and resulting cytotoxicity against cancer cells from mouse, rat, canine, and human origin. Herein, it was shown that nucleolin expression levels were not a limitation on the continuous internalization of F3 peptide-targeted liposomes, despite the saturable nature of the binding mechanism. Modeling enabled the prediction of nucleolin-mediated total doxorubicin exposure provided by the experimental settings of the assessment of PEGASEMP’s impact on cell death. The former increased proportionally with nucleolin-binding sites, a measure relevant for patient stratification. This pattern of variation was observed for the resulting cell death in nonsaturating conditions, depending on the cancer cell sensitivity to doxorubicin. This approach differs from standard determination of cytotoxic concentrations, which normally report values of incubation doses rather than the actual intracellular bioactive drug exposure. Importantly, in the context of development of nucleolin-based targeted drug delivery, the structural nucleolin homology (higher than 84%) and functional similarity across species presented herein, emphasized the potential to use toxicological data and other metrics from lower species to infer the dose for a first-in-human trial.
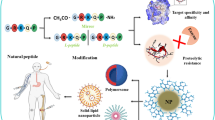
Graphical abstract





Similar content being viewed by others
Availability of data and materials
All data generated or analyzed during this study are included in this published article [and its supplementary information files].
References
Fonseca NA, Moura V, Colelli F, Pesce D, Cardile F, Pisano C, et al. Targeting nucleolin with doxorubicin-containing nanoparticle induces a significant tumor growth inhibition in an orthotopic animal model of standard of care-resistant mesothelioma. Cancer Res. 2017;77:5155.
Fonseca NA, Gregorio AC, Valerio-Fernandes A, Simoes S, Moreira JN. Bridging cancer biology and the patients’ needs with nanotechnology-based approaches. Cancer Treat Rev. 2014;40:626–35.
Srinivasarao M, Low PS. Ligand-targeted drug delivery. Chem Rev. 2017;117:12133–64.
Hendriks BS, Klinz SG, Reynolds JG, Espelin CW, Gaddy DF, Wickham TJ. Impact of tumor HER2/ERBB2 expression level on HER2-targeted liposomal doxorubicin-mediated drug delivery: multiple low-affinity interactions lead to a threshold effect. Mol Cancer Ther. 2013;12:1816–28.
Munster P, Krop IE, LoRusso P, Ma C, Siegel BA, Shields AF, et al. Safety and pharmacokinetics of MM-302, a HER2-targeted antibody-liposomal doxorubicin conjugate, in patients with advanced HER2-positive breast cancer: a phase 1 dose-escalation study. Br J Cancer. 2018;119:1086–93.
Gregorio AC, Lacerda M, Figueiredo P, Simoes S, Dias S, Moreira JN. Meeting the needs of breast cancer: a nucleolin’s perspective. Crit Rev Oncol Hematol. 2018;125:89–101.
Fonseca NA, Gregório AC, Mendes VM, Lopes R, Abreu T, Gonçalves N, et al. GMP-grade nanoparticle targeted to nucleolin downregulates tumor molecular signature, blocking growth and invasion, at low systemic exposure. Nano Today. 2021;37.
Srivastava M, Pollard HB. Molecular dissection of nucleolin’s role in growth and cell proliferation: new insights. FASEB J. 1999;13:1911–22.
Gaume X, Tassin AM, Ugrinova I, Mongelard F, Monier K, Bouvet P. Centrosomal nucleolin is required for microtubule network organization. Cell Cycle. 2015;14:902–19.
Ugrinova I, Monier K, Ivaldi C, Thiry M, Storck S, Mongelard F, et al. Inactivation of nucleolin leads to nucleolar disruption, cell cycle arrest and defects in centrosome duplication. BMC Mol Biol. 2007;8:66.
Xu JY, Lu S, Xu XY, Hu SL, Li B, Li WX, et al. Prognostic significance of nuclear or cytoplasmic nucleolin expression in human non-small cell lung cancer and its relationship with DNA-PKcs. Tumour Biol: the journal of the International Society for Oncodevelopmental Biology and Medicine. 2016;37:10349–56.
Jain N, Zhu H, Khashab T, Ye Q, George B, Mathur R, et al. Targeting nucleolin for better survival in diffuse large B-cell lymphoma. Leukemia. 2018;32:663–74.
Qiu W, Zhou F, Zhang Q, Sun X, Shi X, Liang Y, et al. Overexpression of nucleolin and different expression sites both related to the prognosis of gastric cancer. APMIS. 2013;121:919–25.
Chen C, Chen L, Yao Y, Qin Z, Chen H. Nucleolin overexpression is associated with an unfavorable outcome for ependymoma: a multifactorial analysis of 176 patients. J Neurooncol. 2016;127:43–52.
Marcel V, Catez F, Berger CM, Perrial E, Plesa A, Thomas X, et al. Expression profiling of ribosome biogenesis factors reveals nucleolin as a novel potential marker to predict outcome in AML patients. PLoS ONE. 2017;12:e0170160.
Berger CM, Gaume X, Bouvet P. The roles of nucleolin subcellular localization in cancer. Biochimie. 2015;113:78–85.
Christian S, Pilch J, Akerman ME, Porkka K, Laakkonen P, Ruoslahti E. Nucleolin expressed at the cell surface is a marker of endothelial cells in angiogenic blood vessels. J Cell Biol. 2003;163:871–8.
Fonseca NA, Cruz AF, Moura V, Simoes S, Moreira JN. The cancer stem cell phenotype as a determinant factor of the heterotypic nature of breast tumors. Crit Rev Oncol Hematol. 2017;113:111–21.
Hovanessian AG, Soundaramourty C, Khoury DE, Nondier I, Svab J, Krust B. Surface expressed nucleolin is constantly induced in tumor cells to mediate calcium-dependent ligand internalization. PLoS ONE. 2010;5:e15787.
Porkka K, Laakkonen P, Hoffman JA, Bernasconi M, Ruoslahti E. A fragment of the HMGN2 protein homes to the nuclei of tumor cells and tumor endothelial cells in vivo. Proc Natl Acad Sci U S A. 2002;99:7444–9.
Lauffenburger DA, Linderman J, Berkowitz L. Analysis of mammalian cell growth factor receptor dynamics. Ann N Y Acad Sci. 1987;506:147–62.
Rae JM, Creighton CJ, Meck JM, Haddad BR, Johnson MD. MDA-MB-435 cells are derived from M14 melanoma cells–a loss for breast cancer, but a boon for melanoma research. Breast Cancer Res Treat. 2007;104:13–9.
Sellappan S, Grijalva R, Zhou X, Yang W, Eli MB, Mills GB, et al. Lineage infidelity of MDA-MB-435 cells: expression of melanocyte proteins in a breast cancer cell line. Cancer Res. 2004;64:3479–85.
Fonseca NA, Rodrigues AS, Rodrigues-Santos P, Alves V, Gregorio AC, Valerio-Fernandes A, et al. Nucleolin overexpression in breast cancer cell sub-populations with different stem-like phenotype enables targeted intracellular delivery of synergistic drug combination. Biomaterials. 2015;69:76–88.
Xiong Q, Wilson WK, Pang J. The Liebermann-Burchard reaction: sulfonation, desaturation, and rearrangment of cholesterol in acid. Lipids. 2007;42:87–96.
Moura V, Lacerda M, Figueiredo P, Corvo ML, Cruz ME, Soares R, et al. Targeted and intracellular triggered delivery of therapeutics to cancer cells and the tumor microenvironment: impact on the treatment of breast cancer. Breast Cancer Res Treat. 2012;133:61–73.
Fonseca NA, Gomes-da-Silva LC, Moura V, Simoes S, Moreira JN. Simultaneous active intracellular delivery of doxorubicin and C6-ceramide shifts the additive/antagonistic drug interaction of non-encapsulated combination. J Control Release. 2014;196:122–31.
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol. 2011;7:539.
Said EA, Courty J, Svab J, Delbe J, Krust B, Hovanessian AG. Pleiotrophin inhibits HIV infection by binding the cell surface-expressed nucleolin. FEBS J. 2005;272:4646–59.
Said EA, Krust B, Nisole S, Svab J, Briand JP, Hovanessian AG. The anti-HIV cytokine midkine binds the cell surface-expressed nucleolin as a low affinity receptor. J Biol Chem. 2002;277:37492–502.
Nisole S, Krust B, Hovanessian AG. Anchorage of HIV on permissive cells leads to coaggregation of viral particles with surface nucleolin at membrane raft microdomains. Exp Cell Res. 2002;276:155–73.
Cherry RJ, Smith PR, Morrison IEG, Fernandez N. Mobility of cell surface receptors: a re-evaluation. FEBS Lett. 1998;430:88–91.
Lauffenburger DA, Linderman JJ. Receptors. Models for binding, trafficking, and signalling. The International Journal of Biochemistry & Cell Biology. 1996;28.
Thoumine O, Saint-Michel E, Dequidt C, Falk J, Rudge R, Galli T, et al. Weak effect of membrane diffusion on the rate of receptor accumulation at adhesive contacts. Biophys J. 2005;89:L40–2.
Laouini A, Jaafar-Maalej C, Limayem-Blouza I, Sfar S, Charcosset C, Fessi H. Preparation, characterization and applications of liposomes: state of the art. J Colloid Sci Biotechnol. 2012;1:147–68.
Bulbake U, Doppalapudi S, Kommineni N, Khan W. Liposomal Formulations in Clinical Use: An Updated Review. Pharmaceutics. 2017;9.
Perelson AS. Receptor clustering on a cell surface. III. theory of receptor cross-linking by multivalent ligands: description by ligand states. Mathematical Biosciences. 1981;53:1–39.
Nir S, Peled R, Lee K-D. Analysis of particle uptake by cells: Binding to several receptors, equilibration time, endocytosis. Colloids Surf, A. 1994;89:45–57.
Xu H, Shaw DE. A simple model of multivalent adhesion and its application to influenza infection. Biophys J. 2016;110:218–33.
Straubinger RM, Hong K, Friend DS, Papahadjopoulos D. Endocytosis of liposomes and intracellular fate of encapsulated molecules: Encounter with a low pH compartment after internalization in coated vesicles. Cell. 1983;32:1069–79.
Daleke DL, Hong K, Papahadjopoulos D. Endocytosis of liposomes by macrophages: binding, acidification and leakage of liposomes monitored by a new fluorescence assay. Biochimica et Biophysica Acta (BBA) - Biomembranes. 1990;1024:352–66.
Düzgüneş N, Nir S. Mechanisms and kinetics of liposome–cell interactions. Adv Drug Deliver Rev. 1999;40:3–18.
Bareford LM, Swaan PW. Endocytic mechanisms for targeted drug delivery. Adv Drug Deliv Rev. 2007;59:748–58.
Zhu J, Miao Q, Tang J, Wang X, Dong D, Liu T, et al. Nucleolin mediates the internalization of rabbit hemorrhagic disease virus through clathrin-dependent endocytosis. PLoS Pathog. 2018;14:e1007383.
Caldeira De Moura VLDN, De Almeida Moreira JNS, De Magalhaes SSP, Pedroso De Lima MDCM, inventorsTargeted delivery to human diseases and disorders. PT2012.
Birtwistle MR, Kholodenko BN. Endocytosis and signalling: a meeting with mathematics. Mol Oncol. 2009;3:308–20.
Thorn CF, Oshiro C, Marsh S, Hernandez-Boussard T, McLeod H, Klein TE, et al. Doxorubicin pathways: pharmacodynamics and adverse effects. Pharmacogenet Genomics. 2011;21:440–6.
Šimůnek T, Štěrba M, Popelová O, Adamcová M, Hrdina R, Geršl V. Anthracycline-induced cardiotoxicity: overview of studies examining the roles of oxidative stress and free cellular iron. Pharmacol Rep. 2009;61:154–71.
Brown MS, Anderson RGW, Goldstein JL. Recycling receptors: The round-trip itinerary of migrant membrane proteins. Cell. 1983;32:663–7.
Moehren G, Markevich N, Demin O, Kiyatkin A, Goryanin I, Hoek JB, et al. Temperature dependence of the epidermal growth factor receptor signaling network can be accounted for by a kinetic model. Biochemistry. 2002;41:306–20.
Tomoda H, Kishimoto Y, Lee YC. Temperature effect on endocytosis and exocytosis by rabbit alveolar macrophages. J Biol Chem. 1989;264:15445–50.
Goutelle S, Maurin M, Rougier F, Barbaut X, Bourguignon L, Ducher M, et al. The Hill equation: a review of its capabilities in pharmacological modelling. Fundam Clin Pharmacol. 2008;22:633–48.
Eliaz RE, Nir S, Marty C, Szoka FC Jr. Determination and modeling of kinetics of cancer cell killing by doxorubicin and doxorubicin encapsulated in targeted liposomes. Cancer Res. 2004;64:711–8.
Waters CM, Oberg KC, Carpenter G, Overholser KA. Rate constants for binding, dissociation, and internalization of EGF: effect of receptor occupancy and ligand concentration. Biochemistry. 1990;29:3563–9.
Bourbon H-M, Lapeyre B, Amalric F. Structure of the mouse nucleolin gene. J Mol Biol. 1988;200:627–38.
Bourbon H-M, Amalric F. Nucleolin gene organization in rodents: highly conserved sequenced within three of the 13 introns. Gene. 1990;88:187–96.
Srivastava M, Fleming PJ, Pollard HB, Burns AL. Cloning and sequencing of the human nucleolin cDNA. FEBS Lett. 1989;250:99–105.
Lindblad-Toh K, Wade CM, Mikkelsen TS, Karlsson EK, Jaffe DB, Kamal M, et al. Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature. 2005;438:803–19.
Serin G, Joseph G, Faucher C, Ghisolfi L, Bouche G, Amalric F, et al. Localization of nucleolin binding sites on human and mouse pre-ribosomal RNA. Biochimie. 1996;78:530–8.
González-Camacho F, Medina FJ. Nucleolins from different model organisms have conserved sequences reflecting the conservation of key cellular functions through evolution. J Appl Biomed. 2004;2:151–61.
Ginisty H, Sicard H, Roger B, Bouvet P. Structure and functions of nucleolin. J Cell Sci. 1999;112(Pt 6):761–72.
Mongelard F, Bouvet P. Nucleolin: a multiFACeTed protein. Trends Cell Biol. 2007;17:80–6.
Tajrishi MM, Tuteja R, Tuteja N. Nucleolin: the most abundant multifunctional phosphoprotein of nucleolus. Commun Integr Biol. 2011;4:267–75.
Epstein H, Afergan E, Moise T, Richter Y, Rudich Y, Golomb G. Number-concentration of nanoparticles in liposomal and polymeric multiparticulate preparations: empirical and calculation methods. Biomaterials. 2006;27:651–9.
Gianni L, Vigano L, Locatelli A, Capri G, Giani A, Tarenzi E, et al. Human pharmacokinetic characterization and in vitro study of the interaction between doxorubicin and paclitaxel in patients with breast cancer. J Clin Oncol. 1997;15:1906–15.
Pinto AC, Moreira JN, Simoes S. Ciprofloxacin sensitizes hormone-refractory prostate cancer cell lines to doxorubicin and docetaxel treatment on a schedule-dependent manner. Cancer Chemother Pharmacol. 2009;64:445–54.
Peer D, Karp JM, Hong S, Farokhzad OC, Margalit R, Langer R. Nanocarriers as an emerging platform for cancer therapy. Nat Nanotechnol. 2007;2:751–60.
Pereira PMR, Sharma SK, Carter LM, Edwards KJ, Pourat J, Ragupathi A, et al. Caveolin-1 mediates cellular distribution of HER2 and affects trastuzumab binding and therapeutic efficacy. Nat Commun. 2018;9:5137.
Leahy DJ. Structure and function of the epidermal growth factor (EGF/ErbB) family of receptors. Adv Protein Chem. 2004;68:1–27.
Austin CD, De Maziere AM, Pisacane PI, van Dijk SM, Eigenbrot C, Sliwkowski MX, et al. Endocytosis and sorting of ErbB2 and the site of action of cancer therapeutics trastuzumab and geldanamycin. Mol Biol Cell. 2004;15:5268–82.
Chen X, Kube DM, Cooper MJ, Davis PB. Cell surface nucleolin serves as receptor for DNA nanoparticles composed of pegylated polylysine and DNA. Mol Ther. 2008;16:333–42.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Rye IH, Trinh A, Saetersdal AB, Nebdal D, Lingjaerde OC, Almendro V, et al. Intratumor heterogeneity defines treatment-resistant HER2+ breast tumors. Mol Oncol. 2018;12:1838–55.
Paschalis A, Sheehan B, Riisnaes R, Rodrigues DN, Gurel B, Bertan C, et al. Prostate-specific membrane antigen heterogeneity and DNA repair defects in prostate cancer. Eur Urol. 2019;76:469–78.
Wilhelm S, Tavares AJ, Dai Q, Ohta S, Audet J, Dvorak HF, et al. Analysis of nanoparticle delivery to tumours. Nat Rev Mater. 2016;1:16014.
Gedye CA, Hussain A, Paterson J, Smrke A, Saini H, Sirskyj D, et al. Cell surface profiling using high-throughput flow cytometry: a platform for biomarker discovery and analysis of cellular heterogeneity. PLoS ONE. 2014;9:e105602.
Bates PJ, Reyes-Reyes EM, Malik MT, Murphy EM, O'Toole MG, Trent JO. G-quadruplex oligonucleotide AS1411 as a cancer-targeting agent: Uses and mechanisms. Biochimica et Biophysica Acta (BBA) - General Subjects. 2017;1861:1414–28.
Hu Q, Gu G, Liu Z, Jiang M, Kang T, Miao D, et al. F3 peptide-functionalized PEG-PLA nanoparticles co-administrated with tLyp-1 peptide for anti-glioma drug delivery. Biomaterials. 2013;34:1135–45.
Winer I, Wang S, Lee YE, Fan W, Gong Y, Burgos-Ojeda D, et al. F3-targeted cisplatin-hydrogel nanoparticles as an effective therapeutic that targets both murine and human ovarian tumor endothelial cells in vivo. Cancer Res. 2010;70:8674–83.
Acknowledgements
MIT-Portugal Program.
Funding
This work was supported by the following projects: QREN/FEDER MultiNanoMed (Ref: 23240). This work was also financed by the European Regional Development Fund (ERDF), through the Centro 2020 Regional Operational Program under project CENTRO-01-0247-FEDER-017646 (ODD4PEGASEMP), and through the COMPETE 2020—Operational Program for Competitiveness and Internationalisation and Portuguese national funds via FCT – Fundação para a Ciência e a Tecnologia, I.P., under projects POCI-01-0145-FEDER-016390 (CancelStem), Euronanomed (FCT reference ENMed/0005/2015), CENTRO-01-0145-FEDER-000012-HealthyAging2020 and CIBB (FCT reference UIDB/04539/2020).
Author information
Authors and Affiliations
Contributions
R.L., K.S., and N.A.F. contributed equally for this work. R.L. designed and performed experiments, analyzed data, prepared figures, wrote and edited the manuscript. K.S. designed and performed the mathematical modeling, prepared figures, wrote and edited the manuscript. N.A.F. designed experiments, performed data interpretation, prepared figures, wrote, and edited the manuscript. A.G. provided support on the canine cell line and edited the manuscript. J.S.R. performed the conservation analysis and edited the manuscript. L.A. contributed to the design of the experiments. V.M. and S.S. supervised all the components of the study and edited the manuscript. B.T. supervised the mathematical modeling and edited the manuscript. J.N.M. supervised all components of this study, designed experiments, wrote and edited the manuscript.
Corresponding author
Ethics declarations
Consent for publication
All authors agree with manuscript publication.
Competing interests
V.M. and N.A.F. were former employees at TREAT U, SA. S.S., L.A. and J.N.M. are share-holders of TREAT U, SA. The remaining authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lopes, R., Shi, K., Fonseca, N.A. et al. Modelling the impact of nucleolin expression level on the activity of F3 peptide-targeted pH-sensitive pegylated liposomes containing doxorubicin. Drug Deliv. and Transl. Res. 12, 629–646 (2022). https://doi.org/10.1007/s13346-021-00972-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13346-021-00972-z