Abstract
A common problem in the cultivation and breeding of cauliflower (Brassica oleracea L. var. botrytis) is the occurrence of aneuploids in offspring families. To reveal the chromosomal cause of such numerical variants, it was necessary to develop karyotype tools with which chromosomes can be easily identified. Since mitotic chromosomes in this crop are morphologically similar and lack differentiating banding patterns, we tested two Fluorescent in situ Hybridization (FISH) procedures for chromosome identification: (1) FISH painting with diagnostic repetitive DNA patterns and (2) cross-species chromosome painting. The first method consists of a five-colour FISH with 5s rDNA, 45S rDNA, and two Brassica rapa centromere-specific repeats, and a B. rapa BAC (KBrH092N02) containing a dispersed repeat of an unknown class. The second method is an advanced FISH technology based on hybridising DNA probes of a related species under adapted stringency conditions to identify their homoeologous loci. To this end, we applied four pools of BACs from Arabidopsis thaliana in a multicolour FISH for a banding pattern on the chromosomes of cauliflower (Brassica oleracea L. var. botrytis). Due to the genome triplication and various chromosome rearrangements of Brassica oleracea compared to Arabidopsis, we used MUMmer whole-genome alignment plot information to select Arabidopsis BAC pools with which all cauliflower chromosomes could be identified. In a sample of 21 plants with aberrant phenotypes, we demonstrated primary trisomy for chromosomes 1–6 and 8, and telo-trisomy for chromosomes 7 and 9. Finally, we discuss the advantages and drawbacks of the two painting methods and eventual alternatives for demonstrating numerical aberrations in the cauliflower populations.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cytogenetic analyses of eukaryotic organisms generally begin with the morphological description of their chromosome set. Methods to describe relative lengths, centromere positions and banding patterns are usually referred to as chromosome portraying or karyotype analysis and were obtained for numerous species, including human [59], dog [78], rice [9, 57], wheat [20, 22], tomato [60] and soybean [16]. Karyotype analysis of an organism is associated with its physical map; it helps the assembly of the genomic DNA sequences, supports genetic mapping, and is helpful for phylogenetic and evolutionary studies [1, 7, 12, 28, 51, 65, 67, 71]. Several strategies have been used for karyotype analysis. In the early days, it included measurements of relative chromosome length and long/short arm ratios [40, 60]. However, this strategy fails to identify individual chromosomes in many species, even with newer, sophisticated digital measurements.
Chromosome banding protocols based on chemical or physical treatments of the air-dried chromosome spreads can provide new markers, given that unique banding patterns for different chromosomes are obtained. For several plant species, especially with large chromosomes, banding technologies were developed, such as C-banding, N-banding and modified C-banding [20, 22, 33, 53, 64]. More recent improvements in epifluorescent microscopy, molecular tools and advanced labelling strategies led to the development of Fluorescent in situ hybridisation (FISH) and allow the detection of chromosomal targets directly or indirectly with different fluorophores, ultimately giving rise to multi-colour banding patterns, with which all chromosomes can be identified unequivocally. FISH technology is more informative because it identifies known sequence regions and is versatile and more reliable than simple karyotype analysis, with or without chromosome banding.
FISH has been developed into different technologies, each with specific strength and application. Genomic in situ hybridisation allows the distinction of parental chromosomes in interspecific hybrids using probed genomic DNA of one of the parents with a second different probe or no labelling of the other genomic DNA if the lack of sufficient divergence of the genomes so requires. This genome painting has shown a powerful tool to distinguish parental chromosomes in sexual or somatic hybrids [52, 66]. The second class involves repetitive sequences, either the ubiquitous ribosomal DNA (rDNA) and telomere repeats, satellite and tandem repeats, or dispersed retrotransposons. Such repeats show fluorescent foci on one or several pairs of chromosomes and are used in most crop and model species, including Arabidopsis thaliana [34], maize [31, 39], Pinus species [25] and Norway spruce [73]. The most used repeats for chromosome identification and karyotype analysis through fluorescent in situ hybridisation are rDNA, centromere-specific, and telomere repeats [31].
The third class of DNA sequences comprises unique and single-copy sequences, which are explored in Chromosome in situ Suppression hybridisation (CISS), where probe DNAs were micro-dissected from chromosome spread preparations or flow-sorted from a large number of metaphase chromosomes, then amplified and labelled with Degenerate Oligonucleotide-Primed (DOP)-PCR [70] or Long and Accurate (LA)-PCR [29]. However, applied to plant chromosomes, this strategy was unsuccessful, as plant genomes contain many more dispersed repetitive sequences [61] than animal genomes, thus making them unsuitable for efficient blocking. Besides the abundant occurrence of repetitive sequences, plant genomes also have very similar AT: CG ratio on different chromosomes compared to animal genomes, yet again a presumed consequence of repeat homogenisation along the chromosomes that complicates applying CISS painting [63].
As an alternative strategy to Chromosome In Situ Suppression (CISS) hybridisation, plant cytogeneticists developed a painting technology based on repeat-poor probes from isolated vector DNA. Fuchs et al. [19] utilised yeast artificial chromosomes (YACs), which contained large chromosome contig inserts as probes for hybridisation on tomato and potato chromosomes. Later, contigs or super-contigs of bacterial artificial chromosomes (BAC) clones became much more popular as probes for FISH and were successfully applied in rice and potato species [15, 28]. Lysak et al. [45] used chromosome painting extensively on all five Arabidopsis chromosomes with contigs of adjacent BAC clones, making this model species the first with chromosome-specific painting. Later, Lysak et al. [46, 48,49,50] and Mandáková and Lysak [51] applied the so-called Cross-species Chromosome Painting (CCP) with probes of Arabidopsis BAC contigs on Brassicaceae species under adapted stringency conditions. This CCP was also successful in chromosome studies of potato, tomato and other Solanaceae species [68, 69], sorghum and maize [2]. The CCP studies from Lysak [47, 49] and Ziolkowski [79] revealed a unique evolutionary genome triplication, i.e., as a result of which single Arabidopsis contigs always showed three copies on two or three pairs of chromosomes in diploid Brassica species, including Brassica rapa, Brassica. nigra and Brassica. oleracea [37, 38, 47, 49, 79]. To further elaborate on this genome triplication, Kaczmarek et al. [30] compared the genomes of A. thaliana and B. oleracea with their genetic linkage maps and showed three or four copies of Arabidopsis probes represented in the Brassica genome. A comparable analysis by Parkin et al. [58] demonstrated four to seven copies of A. thaliana in the allotetraploid B. napus. Wang et al. [75], who presented the draft genome sequence of Chinese cabbage (B. rapa), established segmental collinearity of the genomes of B. rapa and A. thaliana, thus confirming the almost complete triplication of the B. rapa genome relative to A. thaliana.
With the triplication of B. rapa in mind and the assumption that B. oleracea demonstrates a similar genome structure, we hypothesised that CCP with probes from Arabidopsis BACs would display three pairs of fluorescent signals on the cauliflower chromosome complement. By selecting specific combinations of pooled Arabidopsis BACs representing contig blocks with known genomic positions on the B. oleracea chromosomes, we expected to create unique multicolour FISH patterns where all individual Brassica chromosomes could be identified.
This study focuses on chromosome identification in cauliflower (Brassica oleracea var. botrytis) because this crop shows high aneuploidy rates in offspring families. Most of these aneuploids are trisomics (unpublished results) and display low yield and aberrant plants and curds, which make them unacceptable for cauliflower cultivation. Aneuploids are a severe problem in most Brassica oleracea crops, both in pure lines and in open-pollinated and F1 hybrid varieties, with aneuploidy rates exceeding 5% in some lines [5, 6]. The aberrant morphology is heritable, but simple Mendelian segregation cannot explain its transmission [6]. Even more, different environmental conditions were suggested to influence the occurrence of abnormal plant shapes [43]; however, solid experimental support still needs to be improved. Chable et al. [6] proposed global DNA methylation as the cause for the aberrant phenotypes in cauliflower, although observations did not support the explanation. More recently, Ji et al. [27] described highly desynaptic male meiosis, which may explain the high rate of unbalanced gametes and aneuploid offspring. Which chromosomes are involved was unknown; therefore, we focussed on identifying the extra chromosomes in the aneuploid cauliflowers. Two multicolour FISH labelling strategies were tested to karyotype cauliflower chromosomes in the cell complement, identifying the aneuploidy based on chromosome painting.
Materials and methods
Plant materials
B. oleracea L. var. botrytis lines and derived diploid and aneuploid progeny plants were obtained from Rijk Zwaan R&D Fijnaart, the Netherlands. The Brassica we used for genome comparison came from the sequenced “walking stick kale” (B. oleracea var. longata) from Tenerife, Canary Islands, clearly distinguishable from the Jersy kale from the Isle of Jersey.
Repetitive DNA sequence probes
Four repetitive DNA sequences were selected for chromosome identification: (1) the 45S rDNA was isolated from the pTa71 plasmid [21] using the High plasmid purification kit (Roche, REF: 11754785001); (2) the 5S rDNA plasmid (pCT 4.2, see Campell et al. [4]) was amplified by PCR reaction with the following primers: 5′-GATCCCATCAGAACTTC-3′ (forward) and 5′-GGTGCTTTAGTGCTGGTAT-3′ (reverse) [32]; (3) CentBr1 and (4) CentBr2 are repetitive centromere sequences of B. rapa and are known to paint the centromere region of B. oleracea [41, 77]. These centromere-specific repeats were amplified with the primer sets given below, as described by Xiong and Pires [77]:
Cent Br1 forward primer 5′-GAATAGCACAGCTTCATCGTCGTTCC-3′
reverse primer 5′-CTGGGAAACTGTAATCACCTGATCTGAAA-3′
CentBr2 forward primer 5′-GGGAATATGACACCTTCTTTGTCATTCT-3′
reverse primer 5′-CAGGAAAACTGGGATCACCTGATTTAAAT-3′
BAC-FISH probes containing repetitive sequences
We also used the KBrH092N2 BACs from the B. rapa Chiifu 401 genotype library [58], which contain repetitive sequences on all chromosome pairs in cauliflower (B. oleracea L. var. botrytis)[77].
Sequence comparison
B. oleracea var. longata was sequenced with Dr Frederic Lens (Naturalis Biodiversity Centre, Leiden, the Netherlands). The referenced assembly was done by Patrick Edger at the University of Missouri using the “iTO1000” genome as a reference (unpublished). The B. rapa genome v1.0 was obtained from the NCBI FTP site, while the A. thaliana genome v10 was obtained from the TAIR database. The relation between the three genomes was studied using MUMmer v3.23 [36]. This software exports only the best alignments ordered so that they create the longest diagonal, assuming a one-to-one relationship between the sequences. After that, the result is filtered to leave only the most extended consistent alignment between the genomes, permitting rearrangements and excluding alignments smaller than 5 Kbp or alignments with sequence identity or sequence uniqueness smaller than 10%.
BAC-FISH probes contain Arabidopsis BACs
To design a painting scheme to identify B. oleracea chromosomes using A. thaliana BACs, we selected forty Arabidopsis BACs from chromosomes 1, 2 and 3 (Table 1 and Results section). Each BAC pool was labelled in one colour. These BACs were provided by the Arabidopsis Biological Resource Center (Columbus, OH, USA, https://www.arabidopsis.org/). BAC DNA was isolated with a high-purification kit (Roche, Germany) and amplified with REPLI-g kit (Qiagen, Germany). (Supplementary data, Table 1).
Slide preparation
We used young flower buds of cauliflower, fixed in freshly prepared ethanol – acetic acid fixative (60% pure ethanol, 30% chloroform and 10% glacial acetic acid) for at least 1 h, followed by storing in ethanol 70%. Anthers were dissected from the flower buds, washed in Milli-Q and 10 mM Na-citrate buffer (pH 4.5), respectively, and then mildly digested in a pectolytic enzyme mix (1% of cellulase RS (Yakult Pharmaceutical IND.CO, LTD, Tokyo, Japan, Yakult 203,033), pectolyase Y23 (pectolyase from Aspergillus japonicus, Sigma Aldrich, St. Louis, MO, USA, P-3026) and cytohelicase (Cyto-helicase from Helix pomatia, Sigma Aldrich, St. Louis, MO, USA, C8274) as a stock solution, then dilute to 0.3% final concentration each in the Na-citrate buffer for 3 h at 37 °C. After removing the enzyme solution by two washing steps in Milli-Q water, one fragile anther was carefully transferred to a clean slide and dissected with fine needles to make a cell mix in 15 µL Milli-Q water. Cells were spread on the slide with 25 µL 50% acetic acid for 4 min on a 50 °C hot plate. Then, the slides were fixed with 50 µL ethanol–acetic acid fixative. We precipitated and dehydrated the cells and let them air dry for a few minutes. We selected only the best slides with well-spread cells, showing chromosomes without cytoplasm for fluorescent in situ hybridisation using a phase contrast microscope equipped with 40 × or 64 × no-cover glass optics.
Fluorescent in situ hybridisation
BAC DNA and selected repetitive sequences were labelled with (either) Cy3.5-dCTP (GE Healthcare Life Science, Amersham, UK, REF: PA53521), Cy3-dUTP (ENZO, REF: ENZ-42501), DEAC-5-dUTP (Perkin Elmer Life Sciences, Boston, MA, REF: NEL-455001EA) and simultaneously with Anti-Digoxigenin-Fluorescein (Roche Applied Science, REF: 11207741910) and Rabbit-anti-fluorescein (FITC) (Jackson (bio-connect), REF: 313-096-003) or biotinylated-anti-streptavidin (Vector Lab, CA, REF: BA-0500) and streptavidin, Alexa Fluor 647 (Cy5) (Invitrogen, S21374) with Dig-Nick-translation (Roche Applied Science, REF: 11725816,910) or Biotin-Nick-translation (Roche Applied Science, REF: 11745824910, 11725816910). The probes of two BAC and of other repetitive sequences hybridised into various slides of the same plant. We pre-treated the cell spreads with 1% formaldehyde for extra fixation (10 min at 20 °C), followed by RNAase (100 μg/mL DNase-free ribonuclease A stock solution, AppliChem, St. Louis, MO, USA, diluted as 1:100 in 2 × SSC (Saline-sodium citrate buffer, pH 7) at 37 °C for 1 h, and washed again with 2 × SSC for 3 × 5 min. The slides were then treated with pepsin (Sigma Aldrich, St. Louis, MO, USA) diluted from a stock solution of 100 mg/mL in 10 mM HCl in a ratio of 1:100 in 10 mM HCl) to remove cytoplasm covering the chromosomes as much as possible. Subsequently, slides were fixed with 1% formaldehyde at room temperature for 10 min. After the fixation, the slides were washed with 2 × SSC and dehydrated in an ethanol series (70%, 90%, and 100% for three minutes each). After dehydration and air drying, we checked the slides in the phase contrast microscope to see if the cytoplasm was sufficiently removed.
The DNA probes (10 µL) were then added to a 10 µl hybridisation mixture containing 50% formamide, and 20% dextran sulphate, followed by denaturation in boiling water for 10 min and put on ice before being added onto the slides. We added 20 µL probe mixture to each slide, which was then transferred to an 80 °C hot plate for a 3-min denaturation step (denaturing chromosomes spread on the slide), followed by hybridisation overnight in a humid chamber at 37 °C. After hybridisation, slides were washed at 42 °C in 50% formamide/2 × SSC for 3 × 5 min, followed by 3 × 5 min of flows in 2 × SSC. For the detection step, we amplified the probe with 500 μg/mL biotinylated-anti-streptavidin (Cy-5) (Vector Laboratories, BA-0500, stock solution 1:200 diluted in TNB buffer (0.1M Tris-HCl pH7.5, 0.15M NaCl, 0.5% blocking reagent, Roche Applied Science, REF: 11096176001) or 200 μg/mL Anti-digoxigenin-fluorescein (FITC) (Roche, 11207741910) a stock solution diluted as 1:200 in TNB buffer) and signal amplification (Cy-5) (200 μg/mL streptavidin, Alexa Fluor 647 (Invitrogen, S21374) a stock solution diluted as 1:800 in TNB buffer or 200 μg/mL Rabbit-anti-sheep-fluorescein (FITC) (Jackson, Bio-connect; 313-096-003, a stock solution diluted as 1:800 in TNB buffer). After the detection steps, slides were dehydrated through an ethanol series (70%, 90%, and 100%, each for three minutes). The air-dried slides were counterstained with 12 μL DAPI (4′, 6-diamidino-2-phenylindole, Sigma Aldrich, St. Louis, MO, USA, D-1388, dissolved in Milli-Q water, stock solution 100 μg/mL, then dilute as 1:20 in Vectashield (50 µL/mL)), and covered with a 24 × 50-glass cover-slip.
The CCP is similar to the repetitive painting described above, with some adaptations. Probe hybridisation on slides was carried out in 36–48 h. After hybridisation, slides were washed three times at 42 °C in 20% formamide/2 × SSC to get a higher hybridisation ratio. The cells were examined with a Zeiss Axioplan 2 imaging photomicroscope. It contained the epi-fluorescence illumination and was equipped with filter sets for DAPI, FITC, Cy3, Cy5, DEAC, and Cy3.5 fluorescence. The images were processed with Genus Image Analysis Workstation software (Applied Imaging), and the selected images were captured by a Photometrics Sensys monochrome 1305 × 1024-pixel CCD camera. Different fluorescent signals were captured and combined using the Genus software's multicolour channel mode. If needed, we further improved brightness and contrast with Adobe Photoshop CS 6.
Results
FISH painting with repetitive DNA sequence
Nature of the repeats
The two Brassica centromere repeats (CentBr1 and CentBr2) contain a 176 bp repeat motif [41] and are present in different Brassica rapa BACs: KBrH001P13 (CentBr1) and KBrH015B20 (CentBr2). Lim [41] grouped the 176 bp motif into two classes CentBr1 and CentBr2, based on sequence similarity, CentBr1 and CentBr2 share around 82% of their sequences, while the members within the two classes share over 90% sequence similarity.
Localisation of different repetitive sequences on B. oleracea
We hybridised all repetitive sequences on the cauliflower slides in a single experiment. Figure 1 displays the painting scheme of the used probes. The 45S rDNA (blue DEAC fluorescence), 5S rDNA (orange Cy3), CentBr1 (far-red Cy5), CentBr2 (green FITC) and the KBrH092NO2 BAC repeat (red Cy 3.5). The 45S rDNA has two loci located on the distal part of the short arms of chromosomes 7 and 8. Chromosome 4 has a locus of 5S rDNA on the long arm. CentBr1 signals were observed on the centromeres and pericentromeres of chromosomes 1, 2, 4, 5, 6, 7 and 9. CentBr2 has more loci than CentBr2 and is also located on chromosomes 3 and 8 but lacks on chromosome 2. Five pairs of the B. oleracea chromosomes could be identified with these four classes of repetitive sequences.
Chromosome sets and ideograms based on multicolour FISH with repetitive sequences as probes on a Brassica oleracea mitotic cell complement. The chromosome identification was based on a labelling scheme of five fluorescent signals of a 45S rDNA (DEAC, blue) and 5s rDNA (Cy3, yellow), b CentBr2 (FITC, green), c KBrH092N02 (Cy 3.5, red), d CentBr1 (Cy5, purple) on top of the DAPI counterstained chromosomes (grey). The multicolour FISH identified all chromosomes and the short arm telosome of chromosome 3 (t3). The magnification bars in the figures equal 5 µm (colour figure online)
Xiong and Pires [77] described two B. rapa BACs, KBrH092N02 and KBrB072L17, that should allow the identification of the remaining two chromosomes. In our hands, only KBrH092N02 gave fluorescent signals, which were present on the pericentromeres of all B. oleracea chromosomes. This suggests that this BAC contains one or more pericentromeric-specific repeats; since the BAC painting size and region are different on the nine chromosome sets of B. oleracea, the BAC could be used as a chromosome marker. Together with this marker, we created a standard set for the karyotype of B. oleracea (Fig. 1). According to the painting result, six chromosome sets (1, 2, 3, 5, 6, 9) are identical with showing signals of BAC KBrH092N02, CentBr1 and CentBr2, but could be distinguished as follows:
Chromosome 1 shows a large region of KBrH092N02 signals on the short arm and the centromere region, which is different from the other chromosome sets.
Chromosome 2 is unique in lacking the presence of signal for CentBr2.
Chromosome 3 lacks the signal of CentBr1, which is different from other chromosomes.
Chromosome 4 contains 5S rDNA on the long arm, which is unique from the other eight chromosomes.
Chromosome 5 shows a large region of KBrH092N02 signals on the long arm and a small region on the short arm. The signal for CentBr2 is weaker than it is on other chromosomes.
Chromosome 6, KBrH092N02 paints the centromere region and the entire short arm, which the same painting does not present in other chromosomes.
Chromosome 7, has 45S rDNA at the distal end of the short arm and N2 at the centromere region. Although it has both CentBr1 and CentBr2 foci, the signal for CentBr2 is weaker than on other chromosomes.
Chromosome 8, is similar to chromosome 7, with 45S rDNA at the distal of the short arm and N2 at the centromere region. It lacks the presence of a signal for CentBr1.
Chromosome 9, CentBr1 displays a strong signal compared with other chromosome sets and does not entirely overlap with CentBr2.
Multicolour cross-species chromosome painting
Comparison of A. thaliana, B. oleracea and B. rapa
To develop a proper painting scheme for identifying all B. oleracea chromosomes, several comparative genome analyses of homoeologous segments between the two species had to be carried out. To this end, we displayed a whole genome alignment by revealing segmental collinearity using the Maximum Unique Matcher (MUMmer 3.23) software. The parameter settings were first tested for B. rapa (Supplementary Fig. 1S) and then compared with the previously published MUMmer plot of the same species [74]. This revealed that our MUMmer plot perfectly matches that in the study of Wang et al. [74]. The plot confirms the same genome triplication and numerous larger and smaller chromosome rearrangements (inversions and translocations). We then constructed a similar MUMmer plot for the five chromosomes of A. thaliana versus the nine chromosomes of B. oleracea (Supplementary Figs. 2S), and that for B. oleracea versus B. rapa (Supplementary Fig. 3S).
The plot for the genomes of A. thaliana versus B. oleracea showed a highly identical pattern of triplication as that for the genomes A. thaliana versus B. rapa, with only a few sequences inverted between the two plots. Sequences of A. thaliana chromosome 1 aligned with B. oleracea chromosome 8, while the identical sequences of A. thaliana are inverted on the genome of B. rapa chromosome 9 (Supplementary Figs. 1S and 2S). The plot for B. oleracea versus B. rapa explains the high similarity of Figs. 1S and 2S.
The plot of the B. oleracea and B. rapa genomes demonstrated the expected high similarity between the two homoeologous genomes (Supplementary Fig. 3S). All B. oleracea contigs have at least one copy in the B. rapa genome, and some contigs have two or three copies. Chromosomes C01, C02 and C03 from B. oleracea were almost completely collinear with chromosomes A01, A02 and A03 in B. rapa. Contig 15, 16, and 17 of B. oleracea chromosome 4 were collinear with B. rapa chromosome 4. We also found some inverted regions between the two genomes, corresponding to and corroborating the above-explained differences found when A. thaliana was compared with both species.
Selection of BAC pools for cross-species FISH
Using the MUMmer plot of A. thaliana and B. oleracea (Supplementary Fig. 2S) we selected several contigs to paint B. oleracea chromosomes. These contigs, derived from Arabidopsis chromosomes 1, 2, and 3, are shown in Fig. 2, with the resulting colour scheme on B. oleracea. We also demonstrated this colour scheme in meiotic complements. In Fig. 3, we show the patterns of the pooled BAC clones on a spread pollen mother cell at diakinesis.
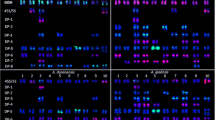
MUMmer plot of the genomes Arabidopsis thaliana (Y-axis) versus Brassica oleracea (X-axis), which was used to construct a painting scheme for multicolour cross-species FISH on B. oleracea cell complements. The genomic relations between A. thaliana and B. oleracea were used to select pools of repeat poor Arabidopsis BACs and were probed with FITC (green), Cy3 (yellow), red (Cy3.5), purple (Cy5). Chromosome identification was based on a combinatorial labelling scheme of the five colours: a BACs from Arabidopsis group 1b (contig 8, green), b BACs from Arabidopsis group 2 (contig 12 and 13, yellow), c BACs from Arabidopsis group 1a (contig 1, red), d BACs from Arabidopsis group 3 (contig 14, purple). Figure e displays the combination of the probes (colour figure online)
Multicolour chromosome painting based on selected pools of BACs of Arabidopsis chromosomes 1, 2 and 3 (cf. Figure 2) and 45S rDNA (DEAC, blue) on a cauliflower chromosome complement at diakinesis from a spread pollen mother cell in which nine chromosome pairs can be discerned. The fluorescent signals follow the labelling scheme for the Arabidopsis BACs in Fig. 2 (colour figure online)
We used a list of selected Arabidopsis BAC clones with a relatively low amount of repetitive sequences (Table 1, Dr Mandáková, pers. comm.). Four contigs of a total of 39 BAC clones of Arabidopsis chromosome regions were now used, together with 45S rDNA for the NOR chromosomes, identifying all nine pairs of B. oleracea chromosomes. The BAC sets of each contig were labelled with different colours for chromosome identification.
The complete list of BAC pools used in our experiments is given hereafter (Table 2).
Arabidopsis chromosome 1 Contig 1 (Arabidopsis group 1b) aligned on B. oleracea chromosomes 5 and 8; the contig was inverted on chromosome 8. Contig 8 (Arabidopsis group 1a) is aligned on B. oleracea chromosomes 2 and 6 and is inverted on chromosome 6.
Arabidopsis chromosome 2 Contig 12 and 13 (Arabidopsis group 2) aligned on B. oleracea chromosomes 3 and 4, the two contigs both aligned and were in inverted orientation on chromosome 4
Arabidopsis chromosome 3 Contig 14 (Arabidopsis group 3) is aligned on B. oleracea chromosome 3 and is inverted on chromosomes 1 and 5.
Karyotype of Arabidopsis BAC on Brassica oleracea
Using Arabidopsis BAC contigs of chromosomes 1, 2 and 3 and 45S rDNA, we could identify nine sets of B. oleracea chromosomes on mitosis or meiosis (Figs. 2 and 3). Colours follow Fig. 2, and chromosomes can be characterised as follows (Table 2):
Chromosome C01 Contig 14 (Arabidopsis group 3) of Arabidopsis chromosome 3 localises on the distal end of the long arm.
Chromosome C02 Contig 8 (Arabidopsis group 1a) of Arabidopsis chromosome 1 is on the pericentromere region of the short arm.
Chromosome C03 Contigs 13 (Arabidopsis group 2) and 14 (Arabidopsis group 3) from Arabidopsis chromosomes 2 and 3 are next to each other on the short arm. Contig 14 localises between the centromere and Contig 13.
Chromosome C04 Contig 13 (Arabidopsis group 2) of Arabidopsis chromosome 2 localizes on the distal region of both arms.
Chromosome C05 Contig 1 (Arabidopsis group 1b) of Arabidopsis chromosome 1 localises at the end of the short arm, while Contig 14 (Arabidopsis group 3) of Arabidopsis chromosome 3 localises at the end of the long arm.
Chromosome C06 Contig 8 (Arabidopsis group 1a) has two copies that localise next to each other on the distal region of the short arm.
Chromosome C07 45s rDNA is at the telomere region of the short arm.
Chromosome C08 45s rDNA is located on the distal region of the short arm and contig 1 (Arabidopsis group 1b) has two copies next to each other on the distal region of the long arm.
Chromosome C09 no contig or 45s rDNA is localised on this chromosome.
Identification of aneuploid cauliflowers with aberrant phenotype
With cross-species chromosome painting and repeat-based karyotyping, we could distinguish all the chromosomes of B. oleracea and identify different trisomics among cauliflower offspring (Figs. 3 and 4).
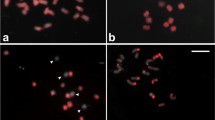
Chromosome identification of an aneuploid B. oleracea based on a simple FISH painting of one repeat probe. a Hybridisation with the 5S rDNA probe reveals trisomy for chromosome 4. The blue signal is the 45S rDNA probe, which identifies chromosome pairs 7 and 8. b FISH with the polled BACs from Arabidopsis chromosome 2 (orange) demonstrated in this complement three copies of the chromosome 3 (trisomy 3), which have a single interstitial signal, and two for chromosome 4, which are characterised by clear distal signals on the short and the long arms (colour figure online)
For cross-species painting with the four pools of Arabidopsis BACs having three copies on the B. oleracea genome, we see six foci for each BAC pool on normal diploid cauliflower chromosomes (Fig. 2). In the case of a primary trisomic containing three copies of one of the chromosomes, the cross-species painting produces seven or eight pairs of foci if the extra chromosome has two copies of the same contig. Figure 4b shows that a cell at mitotic prometaphase has seven orange fluorescing foci, identifying the extra chromosome as C03.
With the repeat-based karyotype, we studied the chromosome complements of various cauliflower plants with the aberrant phenotype (Figs. 1 and 4). As several repeats partly overlap in the multicolour FISH, we analysed the signals in a multi-channel mode of the image analysis software by switching the signals of the different repeats before combining them. Among the results obtained, we not only observed primary trisomics, of which the extra chromosome was identical to one of the nine chromosome pairs, but also observed two plants containing an extra telosome (chromosome arm) to the cell complement (Fig. 1, Table 1).
Discussion
Power of the FISH karyotyping for chromosome identification
This study on chromosome-specific FISH karyotyping in cauliflower aneuploids with molecular markers has shown that repetitive and single-copy probes can successfully identify individual chromosomes in aneuploid cauliflower. However, both technologies have advantages and drawbacks. Repetitive sequences are extensively applied as FISH probes for chromosome identification in plants. With this method, it is possible to reveal unique repeat banding patterns in chromosomes that, without such patterns, appear morphologically similar. The technology can also allow to distinguish between different homoeologous genomes, alien chromosomes from related species and introgressed regions [17, 72]. In our study, with two classes of Brassica centromere repeats, 45S rDNA, 5S rDNA and repetitive sequences of BAC KBrH092N02, we were able to identify the nine chromosomes of B. oleracea, as well as parts of chromosomes (3t and 9t). Repeat polymorphism for the rDNA repeats is obvious if one compares our data with that of Howell [26], who showed three pairs of 45S rDNA foci on the B. oleracea chromosomes 2, 4 and 7, and one 45S rDNA focus on the chromosome that also contains the 5S rDNA. Lim [42] described two chromosomes containing 45S loci in B. oleracea, one co-localized with CentBr1 and CentBr2 and the other with only CentBr2, which is different from our results with two chromosomes containing 45S loci, of which one co-localized with CentBr1 and CentBr2 and the other co-localized only with CentBr1. Xiong and Pires [77] demonstrated that chromosomes 4 and 9 of B. oleracea do not have CentBr1 loci, and chromosome 7 has no CentBr2 locus, whereas B. rapa BAC KBrB072L17 has several loci on B. oleracea while B. rapa BAC KBrH092N02 has only locus on one major pair of chromosomes. Although our chromosome study uses the same probe set as Xiong and Pires [77], our painting with KBrB072L17 did not hybridise on several loci of B. oleracea. In contrast, KBrH092N02 hybridised on all B. oleracea chromosomes.
Chromosome-specific BAC FISH painting in plants is complex because large vector DNAs may contain abundant genomic repetitive sequences. Even if the probe DNA is obtained from a related species, repetitive sequences may obscure a clear FISH pattern from the single copy sequences as repetitive elements in the probe DNA hybridise throughout the target genome. It is helpful to align the genomes of the two species involved in detecting single-copy regions. Recent molecular biosystematic studies convincingly demonstrated ancient whole genome duplications in most plant families [13, 62]. Sequence information indicated that 60% of the Arabidopsis sequences were duplicated in 24 segments in Brassica [48]. Other studies revealed that some genome contigs of Brassica species are triplicated compared to the Arabidopsis thaliana genome [8, 47]. The MUMmer genome comparison between A. thaliana, B. rapa and B. oleracea genome sequences clearly showed that most B. rapa and B. oleracea genome contigs have three copies from A. thaliana genome, and few contigs have one, two or four copies (Supplementary Figs. 1S and 2S). The advantages of this technology are obvious compared to other chromosome banding and painting strategies. Firstly, there is an almost endless choice of Arabidopsis BAC pools for identifying individual chromosomes. Still, such probes are also eminent for identifying minor chromosome domains and are most appropriate for demonstrating large chromosome rearrangements. Secondly, single-copy sequences in BACs directly point at the homoeologous region in the Brassica genome, making it easier to correlate genetic and physical maps. Thirdly, the Arabidopsis probe DNA and target B. oleracea DNA differ in their repeat content. So this painting method outperforms traditional BAC FISH painting because repetitive elements do not hybridise to the target genome and do disturb the specific signals of the single copy sequences (cf. the supplementary Figs. 1S, 2S and 3S). Besides those advantages, there are also some drawbacks. When the genomes of the related species are highly similar, the cross-species FISH specificity may affected by repeat elements. Cheung et al. [10] compared BACs from B. oleracea and B. rapa and concluded a close similarity between their sequences. A similar impression is obvious when comparing the MUMmer plots from the same species. The similar alignment patterns suggested that B. rapa and B. oleracea genomes have largely colinear contigs (Supplementary Fig. 3S). According to the MUMmerplot result, most B. oleracea genome contigs have one copy on the B. rapa genome, while some have two or three copies. In other words, using B. rapa BACs as probes on B. oleracea chromosomes will likely be problematic. When the aneuploids existing on chromosome parts including a centromere; these might not be distinguished with BAC FISH only; for these a centromeric probe is needed (see Fig. 4).
Trisomy identification with karyotyping
The repetitive and cross-species chromosome painting we presented in this study convincingly demonstrated the power of karyotype analysis and chromosome identification in cauliflower. With this technique, we can establish primary trisomics in the progeny of cauliflower breeding material with high rates of aneuploid offspring. Such painting sets enable us to diagnose fast and precisely not only numerical aberrations but also large inversions and translocations in and between the cauliflower chromosomes. As to the molecular marker technology, false positives may occur in the case of telotrisomics if just one marker per chromosome is used, and this marker is located on the disomic chromosome arm (Fig. 1). Telotrisomics originate from univalents that break at their centromere during first meiotic segregation, and their transmission through the gametes can be higher than that of primary trisomics telo-trisomics [18, 35]. This, however, needs to explain the discrepancies between the results of the molecular marker assay and the karyotype analysis (Table 1). The only plausible explanation is that mitotic instability in plants leads to mosaicism of cell lines with different chromosome sets [14].
Chromosome identification allows researchers to identify chromosome segments, indi- vidual chromosomes and whole chromosome sets in a cell complement, and so establish numerical and structural chromosome changes for karyotype evolution or aneuploid di- agnosis. Our study showed two methods to distinguish chromosomes in a crop and to identify extra chromosomes in aneuploid organisms. Either has its advantages and drawbacks, but they are, above all, time-consuming and not suitable for high throughput analyses. To demonstrate aneuploidy in larger populations, array-comparative genomic hybridisation (aCGH) and chromosomal microarray technology (CMA) are good alternatives to FISH-based karyotype technologies. However, a complete physical map and sufficient genomic markers are required. Simple sequence repeat (SSR) and single nucleotide polymorphism (SNP) markers are considered to be reliable for aneuploid detection. Henry et al. [24] presented a novel technology for whole genome karyotyping by quantitative fluorescence-polymerase chain reaction (QF-PCR) on heterozygous aneuploid Arabidopsis. The method determines deviating dosages (allelic ratios) for different chromosome markers. However, the QF-PCR method is restricted to heterozygous genotypes. To detect the frequency of heterozygous genotypes, many codominant SSR markers and multiple types of fluorescent markers may be required. Wen et al. [76] developed a method using SSR markers combined with quantitative PCR to detect aneuploid of loquat; the method could describe the complete molecular karyotype of aneuploid and did not require a large number of SSR markers and not restrict to heterozygous genotype. Kompetitive Allele Specific PCR (KASP) is one of the SNP genotyping platforms which could be applied for aneuploid detection. SNPs are considered an alternative to SSR markers in crop species since SNPs are low-cost and have the potential for high-throughput analysis compared with SSR markers. SSR markers and SNPs are required for whole genome sequence information, which does not apply to uncommon crops and model plants lacking the assembled genome sequence information. Cross-species Chromosome Painting (CCP) or Oligonucleotides fluorescence in situ hybridisation (Oligo-FISH) is an option for aneuploid detection if there is whole genome sequence information available of related model species [44].
Conclusion
Our study demonstrated the power of karyotype analysis in cauliflower based on repetitive and cross-species chromosome painting. Both methods could identify extra copies of the nine chromosomes and demonstrate additional telosomes in cell complements.
The methods that we describe here do have a few drawbacks. Firstly, the method is time-consuming, and one set of microscopic slides requires a full week for the complete slide preparation, FISH and microscopic analysis. Secondly, the repeats that we used can be polymorphic between different B. oleracea varieties and genotypes, so these repeats have to be tested for any repeat polymorphism in new material. Besides trisomy identification, chromosome karyotype is also an excellent method to help understand species differentiation as in B. oleracea species, repetitive sequences are polymorphic in different varieties [3, 25, 74], describing the different parental genomes in polyploid species [11, 55], or can be used to determine homeologous chromosome pairing in interspecific hybrids [23, 54, 56]. An alternative technique to reveal numerical variants in cauliflower populations involves using co-dominant molecular markers for each of the 18 chromosome arms of the crop species.
Data availability
All the data collected during the study have been provided in the manuscript.
Abbreviations
- aCGH:
-
Array-comparative genomic hybridisation
- BAC:
-
Bacterial artificial chromosome
- CCP:
-
Cross-species chromosome painting
- CISS:
-
Chromosome in situ Suppression hybridisation
- CMA:
-
Chromosomal MicroArray technology
- Cy3:
-
Cyanine-3 fluorochrome
- Cy3.5:
-
Cyanine-3.5 fluorochrome
- Cy5:
-
Cyanine-5 fluorochrome
- DAPI:
-
4′,6-Diamidino-2-phenylindole
- DEAC:
-
7-Diethylaminocoumarin-3-carboxylic acid
- DNA:
-
Deoxyribonucleic Acid
- DOP-PCR:
-
Degenerate oligonucleotide-primed polymerase chain reaction
- FISH:
-
Fluorescent in situ hybridisation
- FITC:
-
Fluorescein isothiocyanate
- KASP:
-
Kompetitive allele specific polymerase chain react
- LA-PCR:
-
Long and Accurate polymerase chain reaction
- MUMmer:
-
Maximum unique matcher
- NOR:
-
Nucleolar organizer region
- Oligo-FISH:
-
Oligonucleotides fluorescence in situ hybridisation
- QF-PCR:
-
Quantitative fluorescence-polymerase chain reaction
- SNP:
-
Single nucleotide polymorphism
- SSC:
-
Standard saline citrate
- SSR:
-
Simple sequence repeat
- TAIR:
-
The Arabidopsis information resource
- TNB:
-
Tris–HCl pH7.5, 0.15 M NaCl, 0.5% blocking reagent
- YAC:
-
Yeast artificial chromosome
References
Adams SP, Leitch IJ, Bennett MD, Chase MW, Leitch AR. Ribosomal DNA evolution and phylogeny in aloe (Asphodelaceae). Am J Bot. 2000;87(11):1578–83.
Amarillo FIE, Bass HW. A transgenomic cytogenetic sorghum (Sorghum propinquum) bacterial artificial chromosome fluorescence in situ hybridization map of maize (Zea mays L.) pachytene chromosome 9, evidence for regions of genome hyperexpansion. Genetics. 2007;177:1509–26.
Armstrong SJ, Fransz P, Marshall DF, Jones GH. Physical mapping of DNA repetitive sequences to mitotic and meiotic chromosomes of Brassica oleracea var. alboglabra by fluorescence in situ hybridization. Heredity. 1998;81:666–7.
Campell BR, Song Y, Posch TE, Cullis CA, Town CD. Sequence and organization of 5S ribosomal RNA-encoding genes of Arabidopsis thaliana. Gene. 1992;112(2):225–8.
Chable V, Rival A, Cadot V, Boulineau F, Salmon A, Bellis H, Manzanares-Dauleux M. “Aberrant” plants in cauliflower: 1 phenotype and heredity. Euphytica. 2008;164:325–37.
Chable V, Rival A, Beulé T, Jahier J, Eber F, Cadot V, Boulineau F, Salmon A, Bellis H, Manzanares-Dauleux M. ‘“Aberrant”’ plants in cauliflower: 2 aneuploidy and global DNA methylation. Euphytica. 2009;170:275–87.
Chamala S, Chanderbali AS, Der JP, Lan T, Walts B, Albert VA, de Pamphilis CW, Leebens-Mack J, Rounsley S, Schuster SC, Wing RA, Xiao N, Moore R, Soltis PS, Soltis DE, Barbazuk WB. Assembly and validation of the genome of the nonmodel basal angiosperm Amborella. Science. 2013;342(6165):1516–7. https://doi.org/10.1126/science.1241130.
Cheng F, Wu J, Wang X. Genome triplication drove the diversification of Brassica plants. Hortic Res. 2014;1:14024. https://doi.org/10.1038/hortres.2014.24.
Cheng Z, Buell CR, Wing RA, Gu M, Jiang J. Toward a cytological characterization of the rice genome. Genome Res. 2001;11:2133–41.
Cheung F, Trick M, Drou N, Lim YP, Park JY, Kwon SJ, et al. Comparative analysis between homoeologous genome segments of Brassica napus and its progenitor species reveals extensive sequence-level divergence. Plant Cell. 2009;21(7):1912–28. https://doi.org/10.1105/tpc.108.060376.
Danilova TV, Friebe B, Gill BS. Development of a wheat single gene FISH map for analysing homoeologous relationship and chromosomal rearrangements within the triticeae. Theor Appl Genet. 2014;127(3):715–30. https://doi.org/10.1007/s00122-013-2253-z.
de Fátima RC, VanzelaI ALL, Santos MO, Fregonezi JN, Ruas PM, Matzenbacher NI, de Aguiar-Perecin MLR. Chromosomal organization and phylogenetic relationships in Hypochaeris species (Asteraceae) from Brazil. Genet Mol Biol. 2005;28(1):129–39. https://doi.org/10.1590/S1415-47572005000100023.
Debodt S, Maere S, van de Peer Y. Genome duplication and the origin of angiosperms. Trends Ecol Evol. 2005;20(11):591–7. https://doi.org/10.1016/j.tree.2005.07.008.
Deng M, Chen J, Henny RJ, Li Q. Chromosome number and karyotype variation in Codiaeum variegatum cultivars. HortScience. 2010;45(4):538–40.
Dong F, Song J, Naess SK, Helgeson JP, Gebhardt C, Jiang J. Development and applications of a set of chromosome-specific cytogenetic DNA markers in potato. Theor Appl Genet. 2000;101:1001–7.
Findley SD, Cannon S, Varala K, Du J, Ma J, Hudson ME, Birchler JA, Stacey G. A fluorescence in situ hybridization system for karyotyping soybean. Genetics. 2010;185:727–44.
Fradkin M, Ferrari MR, Espert SM, Ferreira V. Differentiation of triticale cultivars through FISH karyotyping of their rye chromosomes. Genome. 2013;56:267–72. https://doi.org/10.1139/gen-2012-0117.
Friebe B, Zhang P, Linc G, Gill BS. Robertsonian translocations in wheat arise by centric misdivision of univalents at anaphase I and rejoining of broken centromeres during interkinesis of meiosis II. Cytogenet Genome Res. 2005;109(1–3):293–7.
Fuchs J, Kloos D, Ganal MW, Schubert I. In situ localization of yeast artificial chromosome sequences on tomato and potato metaphase chromosomes. Chromosome Res. 1996;4(4):277–81.
Gerlach WL. N-banded karyotypes of wheat species. Chromosoma (Berl). 1977;62:49–56.
Gerlach WL, Bedbrook JR. Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acid Res. 1979;7:1869–85.
Gill BS, Friebe B, Endo TR. Standard karyotype and nomenclature system for description of chromosome bands and structural aberrations in wheat (Triticum aestivum). Genome. 1991;34:830–9.
Hao M, Luo J, Yang M, Zhang L, Yan Z, Yuan Z, Zheng Y, Zhang H, Liu D. Comparison of homoeologous chromosome pairing between hybrids of wheat genotypes Chinese spring ph1b and Kaixian-luohanmai with rye. Genome. 2011;54:1–6.
Henry IM, Dilkes BP, Comai L. Molecular karyotyping and aneuploidy detection in Arabidopsis thaliana using quantitative fluorescent polymerase chain reaction. Plant J. 2006;48(2):307–19.
Hizume M, Shibata F, Matsusaki Y, Garajova Z. Chromosome identification and comparative karyotypic analyses of four Pinus species. Theor Appl Genet. 2002;105:491–7.
Howell EC, Barker GC, Jones GH, Kearsey MJ, King GJ, Kop EP, Ryder CD, Teakle GR, Vicente JG, Armstrong SJ. Integration of the cytogenetic and genetic linkage maps of Brassica oleracea. Genetics. 2002;161:1225–34.
Ji X, Lelivelt C, Wijnker E, de Jong H. Is partial desynapsis in cauliflower (Brassica oleracea L. var. botrytis) pollen mother cells linked to aneuploidy in the crop? Euphytica. 2022;218:79.
Jiang J, Gill BS, Wang G, Ronald PC, Ward DC. Metaphase and interphase fluorescence in situ hybridization mapping of the rice genome with bacterial artificial chromosomes. Proc Natl Acad Sci USA. 1995;92:4487–91.
Johnson DH. Molecular cloning of DNA from specific chromosomal regions by microdissection and sequence-independent amplification of DNA. Genomics. 1990;6:243–51.
Kaczmarek M, Koczyk G, Ziolkowski PA, Babula-Skowronska D, Sadowski J. Comparative analysis of the Brassica oleracea genetic map and the Arabidopsis thaliana genome. Genome. 2009;52:620–33.
Kato A, Vega JM, Han F, Lamb JC, Birchler JA. Advances in plant chromosome identification and cytogenetic techniques. Curr Opin Plant Biol. 2005;8:148–54.
Koo DH, Hur Y, Jin DC, Bang JW. Karyotype analysis of a Korean cucumber cultivar (Cucumis sativus L. cv. Winter Long) using C-banding and bicolor fluorescence in situ hybridization. Mol Cells. 2002;13(3):413–8.
Koo DH, Choi HW, Cho J, Hur Y, Bang JW. A high-resolution karyotype of cucumber (Cucumis sativus L. ‘Winter Long’) revealed by C-banding, pachytene analysis, and RAPD-aided fluorescence in situ hybridization. Genome. 2005;48:534–40.
Koornneef M, Fransz P, de Jong H. Cytogenetic tools for Arabidopsis thaliana. Chromosome Res. 2003;11:183–94.
Koornneef M, Van der Veen JH. Trisomics in Arabidopsis thaliana and the location of linkage groups. Genetica. 1983;61(1):41–6.
Kurtz S, Phillippy A, Delcher AL, Smoot M, Shumway M, Antonescu C, Salzberg SL. Versatile and open software for comparing large genomes. Genome Biol. 2004;5:12.
Lagercrantz U, Lydiate DJ. Comparative genome mapping in Brassica. Genetics. 1996;144:1903–10.
Lagercrantz U, Putterill J, Coupland G, Lydiate D. Comparative mapping in Arabidopsis and Brassica, fine scale genome collinearity and congruence of genes controlling flowering time. Plant J. 1996;9:13–20.
Lamb JC, Birchler JA. Retroelement genome painting: cytological visualization of retroelement expansions in the genera Zea and Tripsacum. Genetics. 2006;173:1007–21.
Levan A, Fredga K, Sandberg AA. Nomenclature for centromeric position on chromosomes. Hereditas. 1964;52(2):201–20.
Lim KB, de Jong H, Yang TJ, Park JY, Kwon SJ, Kim JS, Lim MH, Kim JA, Jin M, Jin YM, Kim SH, Lim YP, Bang JW, Kim HI, Park BS. Characterization of rDNAs and tandem repeats in the heterochromatin of Brassica rapa. Mol Cells. 2005;19(3):436–44.
Lim KB, Yang TJ, Hwang YJ, Kim JS, Park JY, Kwon SJ, Kim J, Choi BS, Lim MH, Jin M, Kim HI, de Jong H, Bancroft I, Lim Y, Park BS. Characterization of the centromere and peri-centromere retrotransposons in Brassica rapa and their distribution in related Brassica species. Plant J. 2007;49(2):173–83.
Lindemann-Zutz K, Fricke A, Stützel H. Prediction of time to harvest and its variability in broccoli (Brassica oleracea var. italica) part I. Plant developmental variation and forecast of time to head induction. Sci Hortic. 2016;198:424–33.
Liu G, Zhang T. Single copy oligonucleotide fluorescence in situ hybridization probe design platforms: development, application and evaluation. Int J Mol Sci. 2021;22(13):7124.
Lysak MA, Fransz PF, Ali HBM, Schubert I. Chromosome painting in Arabidopsis thaliana. Plant J. 2001;28(6):689–97.
Lysak MA, Pecinka A, Schubert I. Recent progress in chromosome painting of Arabidopsis and related species. Chromosome Res. 2003;11(3):195–204.
Lysak MA, Koch M, Pecinka A, Schubert I. Chromosome triplication found across the tribe brassiceae. Genome Res. 2005;15:516–25.
Lysak MA, Berr A, Pecinka A, Schmidt R, McBreenm K, Schubert I. Mechanisms of chromosome number reduction in Arabidopsis thaliana and related Brassicaceae species. PNAS. 2006;103(13):5224–9.
Lysak MA, Cheung K, Kitschke M, Bures P. Ancestral chromosomal blocks are triplicated in Brassiceae species with varying chromosome number and genome size. Plant Physiol. 2007;145(2):402–10.
Lysak MA, Mandáková T, Lacombe E. Reciprocal and multi-species chromosome bac painting in crucifers (Brassicaceae). Cytogenet Genome Res. 2010;129(1–3):184–9. https://doi.org/10.1159/000312951.
Mandáková T, Lysak MA. Chromosomal phylogeny and karyotype evolution in x=7 crucifer species (Brassicaceae). Plant Cell. 2008;20(10):2559–70.
Markova M, Vyskot B. New horizons of genomic in situ hybridization. Cytogenet Genome Res. 2009;126(4):368–75. https://doi.org/10.1159/000275796.
Martin J, Hesemann CU. Cytogenetic investigations in wheat, rye and triticale. I. Evaluation of improved Giemsa C- and fluorochrome banding techniques in rye chromosomes. Heredity. 1988;61:459–67.
Miller TE, Reader SM, Purdie KA, King IP. Fluorescent in situ hybridization-a useful aid to the introduction of alien genetic variation into wheat. Euphytica. 1996;89:113–9.
Mukai Y, Nakahara Y, Yamamoto M. Simultaneous discrimination of the three genomes in hexaploid wheat by multicolour fluorescence in situ hybridization using total genomic and highly repeated DNA as probes. Genome. 1993;36(3):489–94.
Nicolas SD, Le Mignon G, Eber F, Coriton O, Monod H, Clouet V, Huteau V, Lostanlen A, Delourme R, Chalhoub B, Ryder CD, Chèvre AM, Jenczewski E. Homeologous recombination plays a major role in chromosome rearrangements that occur during meiosis of Brassica napus haploids. Genetics. 2007;175(2):487–503.
Ohmido N, Fukui K. Cytological studies of African cultivated rice. Oryza glaberrima Theor Appl Genet. 1995;91:212–7.
Parkin IAP, Gulden SM, Sharpe AG, Lukens L, Trick M, Osborn TC, Lydiate DJ. Segmental Structure of the Brassica napus genome Based on comparative analysis with Arabidopsis thaliana. Genetics. 2005;171(2):765–81.
Pearson P. The use of new staining techniques for human chromosome identification. J Med Genet. 1972;9(3):264–75.
Ramanna MS, Prakken R. Structure of and homology between pachytene and somatic metaphase chromosomes of the tomato. Genetica. 1967;38(1):115–33.
Schmidt T, Heslop-Harrison JS. Genomes, genes and junk: the large-scale organization of plant chromosomes. Trends Plant Sci. 1998;3:195–9.
Schranz ME, Mohammadin S, Edger PP. Ancient whole genome duplications, novelty and diversification: the WGD radiation lag-time model. Curr Opin Plant Biol. 2012;15(2):147–53. https://doi.org/10.1016/j.pbi.2012.03.011.
Schwarzacher T, Wang ML, Leitch AR, Miller N, Moore G, Heslop-Harrison JS. Flow cytometric analysis of the chromosomes and stability of a wheat cell-culture line. Theor Appl Genet. 1997;94:91–7.
Seal AG, Bennett MD. Preferential C-banding of wheat or rye chromosomes. Theor Appl Genet. 1982;63(3):227–33.
Shearer LA, Anderson LK, de Jong H, Smit S, Goicoechea JL, Roe BA, Hua A, Giovannoni JJ, Stack SM. Fluorescence in situ hybridization and optical mapping to correct scaffold arrangement in the tomato genome. G3 Genes, Genomes, Genet. 2014;4(8):1395–405.
Silva GS, Souza MM. Genomic in situ hybridization in plants. Genet Mol Res. 2013;12(3):2953–65. https://doi.org/10.4238/2013.August.12.11.
Stack SM, Royer SM, Shearer LA, Chang SB, Giovannoni JJ, Westfall DH, White RA, Anderson LK. Role of fluorescence in situ hybridization in sequencing the tomato genome. Cytogenet Genome Res. 2009;124(3–4):339–50. https://doi.org/10.1159/000218137.
Szinay D, Wijnker E, van den Berg R, Visser RGF, de Jong H, Bai Y. Chromosome evolution in Solanum traced by cross-species BAC-FISH. New Phytol. 2012;195(3):688–98. https://doi.org/10.1111/j.1469-8137.2012.04195.x.
Tang X, Szinay D, Lang C, Ramanna MS, van der Vossen EAG, Datema E, Lankhorst RK, de Boer J, Peters SA, Bachem C, Stiekema W, Visser RGF, de Jong H, Bai Y. Cross-species BAC-FISH painting of the tomato and potato chromosome 6 reveals undescribed chromosomal rearrangements. Genetics. 2008;180:1319–28.
Telenius H, Carter NP, Bebb CE, Nordenskjöld M, Ponder BAJ, Tunnacliffe A. Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics. 1992;13(3):718–25.
The Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature. 2012; vol. 485, p. 635-641.
Tsujimoto H, Mukai Y, Akagawa K, Nagaki K, Fujigaki J, Yamamoto M, Sasakuma T. Identification of individual barley chromosomes based on repetitive sequences: conservative distribution of Afa-family repetitive sequences on the chromosomes of barley and wheat. Genes Genet Syst. 1997;72(5):303–9.
Vischi M, Jurman I, Bianchi G, Morgante M. Karyotype of Norway spruce by multicolor FISH. Theor Appl Genet. 2003;107:591–7.
Wang J, Lydiate DJ, Parkin IAP, Falentin C, Delourme R, Carion PWC, King GJ. Integration of linkage maps for the amphidiploid Brassica napus and comparative mapping with Arabidopsis and Brassica rapa. BMC Genomics. 2011;12:101.
Wang X, et al. The genome of the mesopolyploid crop species Brassica rapa. Nat Genet. 2011;43(10):1035–9.
Wen G, Dang J, Xie Z, Wang J, Jiang P, Guo Qand Liang G. Molecular karyotypes of loquat (Eriobotrya japonica) aneuploids can be detected by using SSR markers combined with quantitative PCR irrespective of heterozygosity. Plant Methods. 2020;16(1):22.
Xiong Z, Pires JC. Karyotype and identification of all homoeologous chromosomes of Allopolyploid Brassica napus and its diploid progenitors. Genetics. 2011;187(1):37–49.
Yang F, Graphodatsky AS, O’Brien PC, Colabella A, Solanky N, Squire M, Sargan DR, Ferguson-Smith MA. Reciprocal chromosome painting illuminates the history of the genome evolution of the domestic cat, dog and human. Chromosome Res. 2000;8:393–404.
Ziolkowski PA, Kaczmarek M, Babula D, Sadowski J. Genome evolution in Arabidopsis/Brassica: conservation and divergence of ancient rearranged segments and their breakpoints. Plant J. 2006;47(1):63–74.
Acknowledgements
The paper is a compilation of studies previously published in two chapters of the PhD thesis of the first author (Xianwen Ji: Numerical and structural chromosome aberrations in cauliflower (Brassica oleracea var. botrytis) and Arabidopsis thaliana. PhD thesis, Wageningen University, 2014, ISBN 978-94-6257-160-0). Data in these chapters were merged, and the text was critically reviewed.
Funding
Rijk Zwaan R&D Fijnaart, the Netherlands, provided funding for the PhD study of Xianwen Ji.
Author information
Authors and Affiliations
Contributions
XJ and HdJ designed the experimental work and wrote the manuscript. SA, SP and MES provided genome sequence information and bioinformatics analyses. JP and AV were helpful in the experimental work. SP, CL and HdJ supervised the project and critically reviewed the manuscript. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no potential conflict of interest in this paper.
Ethical approval
No approval was required for the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Corrrsponding Editor: Maki Yamamoto; Reviewer: Y. Mukai.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ji, X., Aflitos, S.A., Peters, S. et al. FISH painting for chromosome identification of aneuploid cauliflower (Brassica oleracea L. var. botrytis). Nucleus 66, 273–288 (2023). https://doi.org/10.1007/s13237-023-00449-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13237-023-00449-z