Abstract
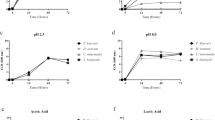
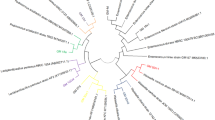
This study aimed to isolate and characterize potential probiotic yeasts from Ethiopian injera sourdough and the study was conducted by collecting samples from Gondar and Bahir Dar cities, Ethiopia. The potential yeasts were isolated and identified using morphological, physiological, biochemical and molecular based analysis. Promising isolates were selected to further investigate their in vitro probiotic properties, including survival at different temperatures (25, 30, 37, and 42 °C), acidic pH (2, 3, 4 and 5), bile salt (0.1, 0.3, and 0.5%), and osmotolerance (20, 30, 40, and 50% glucose concentration), antimicrobial activities, proteolytic and lipolytic activities as well as resistance to four antibiotics. From 20 samples, 38 isolates were obtained. Among these, 10 produced low or non-hydrogen sulfide and were selected for further work. Further screening tests revealed that five isolates (G1N1, G2N4, G3N1, G8N1, and B6N3) were able tolerate and grow at 37 °C, with harsh conditions of the human digestive tract like low pH, bile salt, and higher osmotic effect. The maximum growth OD values were recorded at 37 °C by isolate G4N1 (OD value (0.6667), while G3N1 exhibited a maximum growth OD value of 0.4227 at pH 2. On the other hand, G2N4 gave a maximum OD value of 0.8800 at 0.3% bile salt concentration. The promising isolates were sequenced and identified to species level. Based on phylogenetic tree analysis, all the five probiotic yeast isolates had one common ancestor and belonging to Saccharomyces cerevisiae (G1N1 and G2N4), Candida humilis (G3N1 and B6N3), and Pichia kudriavzevii (G8N1). This study revealed that Ethiopian injera sourdough could be potential source of different probiotic yeast strains. Strong emphasis should be given about the use of probiotic yeasts that are isolated from Ethiopian fermented foods.





Similar content being viewed by others
Data availability
The data used to support the findings of this study are included within the article. The nucleotide sequence data are available in the GenBank (NCBI) databases under the accession numbers of OP942415 (for B6N3), OP942416 (for G1N1), OP942417 (for G2N4), OP942418 (for G3N1), and OP942419 (for G8N1).
References
Antia UE, Akan OD, Stephen NU, Eno-Ibanga CK, Akpan NG (2018) Isolation and screening of yeast isolates indigenous palm wine for ethanol production. Philip J Sci 147(3):411–417
Barnett JA, Payne RW, Yarrow D (2000) Yeasts: characteristics and identification, 3rd edn. Cambridge University Press, Cambridge
Barretto R, Buenavista RM, Rivera JL, Wang S, Prasad PV, Siliveru K (2021) Teff (Eragrostis teff) processing, utilization and future opportunities: a review. Int J Food Sci Technol 56(7):3125–3137
Bhukya B, Marrivada SR, Devi TA, Reddy YR, Rao LV (2010) Screening and characterization of stress tolerant Saccharomyces cerevisiae isolated from brewery effluents for animal probiotic applications. Institute Integrat Omics Appl Biotechnol J 1(4):32–39
Chen LS, Ma Y, Maubois JL, He SH, Chen LJ, Li HM (2010) Screening for the potential probiotic yeast strains from raw milk to assimilate cholesterol. Dairy Sci Technol 90(5):537–548
Cocolin L, Bisson LF, Mills DA (2000) Direct profiling of the yeast dynamics in wine fermentations. FEMS Microbiol Lett 189:81–87
Corbo MR, Sinigaglia M, Bevilacqua A (2017) Biotechnological application of yeasts in food science: starter cultures, probiotics and enzyme production. J Appl Microbiol 123:1360–1372
Czerucka D, Piche T, Rampal P (2007) Review article: yeast as probiotics—Saccharomyces boulardii. Aliment Pharmacol Ther 26:767–778
Foulquie MR, Callewaert R, Devreese B, Van Beeumen J, De Vuyst L (2003) Isolation and biochemical characterization of enterocins produced by enterococci from different sources. J Appl Microbiol 94(2):214–229
Genetie AA (2020) Assessment of downriver pollution profile of Gondar city wastewater and its influence on Keha river. Appl J Environ Eng Sci 6(3):291–309
Islam T (2016) Isolation of amylase producing bacteria from soil and identification by 16S rRNA gene sequencing and characterization of amylase. Dissertation, BRAC University
Karki TB, Timilsina PM, Yadav A, Pandey GR, Joshi Y, Bhujel S, Neupane K (2017) Selection and characterization of potential baker’s yeast from indigenous resources of Nepal. Biotechnol Res Int 2:2
Katarzyna R, Alina KS (2010) Probiotic properties of yeasts isolated from chicken feces and kefirs. J Microbiol 59(4):257–263
Khagwal N, Sharma DC, Sharma PK (2019) Isolation and characterization of potential probiotic strains isolated from traditional Indian fermented foods. Int J Curr Microbiol Appl Sci 8(3):680–689
Khidhr KO (2014) Isolation and identification of Saccharomes cerevisiae var boulardii and its uses as a probiotic (in vitro). Rafidain J Sci 25(1):1–11
Kindu G (2019) Efficiency of locally available spices to improve shelf life and sensory attributes of teff injera. World Sci News 136:159–172
Kıvanc M, Yapici E (2015) Kefir as a probiotic dairy beverage: determination lactic acid bacteria and yeast. Int J Food Eng 1(1):55–60
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Kunyeit L, Ka AA, Rao RP (2020) Application of probiotic yeasts on Candida species associated infection. J Fungi 6(4):189
Kurtzman CP, Robnett CJ (1998) Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Antonie Leeuwenhoek 72:331–371
Lourens A, Viljoen BC (2001) Growth and survival of a probiotic yeast in dairy products. Food Res Int 134:791–796
Mezemir S (2015) Probiotic potential and nutritional importance of teff (Eragrostistef (Zucc) Trotter) enjerra—a review. Afr J Food Agr Nut Dev 15(2):9964–9981
Mohamed Z, Feuilloley MGJ, Connil N (2020) Update of probiotics in human world: A nonstop source of benefactions till the end of time. Microorganisms 8(12):1–33
Moradi R, Nosrati R, Zare H, Tahmasebi T, Saderi H, Owlia P (2018) Screening and characterization of in-vitro probiotic criteria of Saccharomyces and Kluyveromyces strains. Iran J Microbiol 10(2):123–131
Plou FJ, Ferrer M, Nuero OM, Calvo MV, Alcalde M, Reyes F, Ballesteros A (1998) Analysis of tween 80 as an esterase/lipase substrate for lipolytic activity assay. Biotechnol Tech 12(3):183–186
Prasad M (2014) Characterization of amylase gene in Bacillus species isolated from different soil samples. Int J Current Microbiol Appl Sci 3(9):891–896
Ragavan ML, Das N (2017) Molecular identification of probiotic yeast strains and their characterization. Asian J PharmaClin Res 10(4):339–343
Rajkowska K, Kunicka SA (2010) Probiotic properties of yeasts isolated from chicken feces and kefirs. Polish J Microbiol 59(4):257
Salminen SJ, Gueimonde M, Isolauri E (2005) Probiotics that modify disease risk. J Nutr 135(5):1294–1298
Sarwar A, Aziz T, Al-Dalali S, Zhao X, Zhang J, Udin J, Chen C, Cao Y, Yang Z (2019) Physicochemical and microbiological properties of synbiotic yogurt made with probiotic yeast Saccharomyces boulardii in combination with inulin. Foods 8:468
Satheesh N, Solomon WF (2020) Injera (an ethnic, traditional staple food of ethiopia): a review on traditional practice to scientific developments. J Ethnic Foods 7(1):1–15
Staniszewski A, Kordowska-Wiater M (2021) Probiotic and potentially probiotic yeasts—characteristics and food application. Foods 10(6):1306
Teramoto Y, Sato R, Ueda S (2005) Characteristics of fermentation yeast isolated from traditional Ethiopian honey wine. Afr J Biotechnol 4(2):160–163
Tripathi MK, Giri SK (2014) Probiotic functional foods: survival of probiotics during processing and storage. J Funct Foods 9:225–241
Truong HTD, Saeys W (2021) Fluorescence-based discrimination of vegetative cells of bacillus strains from Escherichia coli and Saccharomyces cerevisiae. Biosys Eng 209:232–245
Tsegaye Z, Tefera G, Gizaw B, Abatenh E (2018) Characterization of yeast species isolated from local fruits used for bakery industrial application. J Appl Microbiol Res 1(1):21–26
Walker GM, White NA (2017) Introduction to fungal physiology. In: Kavangh K (ed) Fungi: biology and applications. JohnWiley and Sons, New Jersey, pp 1–58
Yuma YS (2020) Isolation and characterization of yeast as potential probiotics from fermented cereals dough. Vet Med Res 7(6):1–7
Acknowledgements
Nigus Muche would like to acknowledge University of Gondar for sponsoring his MSc study. All authors are delighted to extend their heartfelt thanks to hotels and households of Gondar and Bahir Dar cities for their generous permission in collecting injera sourdough samples.
Funding
This research work received no funding.
Author information
Authors and Affiliations
Contributions
NM performed the experiments as part of his master’s thesis work. All this work was carried out under the supervision of TG and TMJ. TMJ helped in editing the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflicts of interest.
Research involving human participants and/or animals
We confirm that the research does not involve human participants and/or animals.
Informed consent
Since the study does not involve any human participants, informed consent is not applicable for the study.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Muche, N., Geremew, T. & Jiru, T.M. Isolation and characterization of potential probiotic yeasts from Ethiopian injera sourdough. 3 Biotech 13, 300 (2023). https://doi.org/10.1007/s13205-023-03729-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-023-03729-2