Abstract
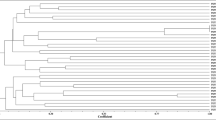
The diversity of rhizobia that establish symbiosis with Lotus corniculatus has scarcely been studied. Several species of Mesorhizobium are endosymbionts of this legume, including Mesorhizobium loti, the type species of this genus. We analysed the genetic diversity of strains nodulating Lotus corniculatus in Northwest Spain and ten different RAPD patterns were identified among 22 isolates. The phylogenetic analysis of the 16S rRNA gene showed that the isolated strains belong to four divergent phylogenetic groups within the genus Mesorhizobium. These phylogenetic groups are widely distributed worldwide and the strains nodulate L. corniculatus in several countries of Europe, America and Asia. Three of the groups include the currently described Mesorhizobium species M. loti, M. erdmanii and M. jarvisii which are L. corniculatus endosymbionts. An analysis of the recA and atpD genes showed that our strains belong to several clusters, one of them very closely related to M. jarvisii and the remanining ones phylogenetically divergent from all currently described Mesorhizobium species. Some of these clusters include L. corniculatus nodulating strains isolated in Europe, America and Asia, although the recA and atpD genes have been sequenced in only a few L. corniculatus endosymbionts. The results of this study revealed great phylogenetic diversity of strains nodulating L. corniculatus, allowing us to predict that even more diversity will be discovered as further ecosystems are investigated.




Similar content being viewed by others
References
Ampomah OY, Huss-Danell K (2011) Genetic diversity of root nodule bacteria nodulating Lotus corniculatus and Anthyllis vulneraria in Sweden. Syst Appl Microbiol 34:267–275
Armas-Capote N, Pérez-Yépez J, Martínez-Hidalgo P, Garzón-Machado V, del Arco-Aguilar M, Velázquez E, León-Barrios M (2014) Core and symbiotic genes reveal nine Mesorhizobium genospecies and three symbiotic lineages among the rhizobia nodulating Cicer canariense in its natural habitat (La palma, Canary Islands). Syst Appl Microbiol 37:140–148
Binde DR, Menna P, Bangel EV, Barcellos FG, Hungria M (2009) Rep-PCR fingerprinting and taxonomy based on the sequencing of the 16S rRNA gene of 54 elite commercial rhizobial strains. Appl Microbiol Biotechnol 83:897–908
De Meyer SE, Van Hoordea K, Vekeman B, Braeckmana T, Willems A (2011) Genetic diversity of rhizobia associated with indigenous legumes in different regions of Flanders (Belgium). Soil Biol Biochem 43:2384–2396
Escaray FJ, Menéndez AB, Garriz A, Pieckenstain FL, Estrella MJ, Castagno LN, Carrasco P, Sanjuán J, Ruiz OA (2012) Ecological and agronomic importance of the plant genus Lotus. Its application in grassland sustainability and the amelioration of constrained and contaminated soils. Plant Sci 182:121–133
Gaunt MW, Turner SL, Rigottier-Gois L, Lloyd-Macgilp SA, Young JWP (2001) Phylogenies of atpD and recA support the small subunit rRNA-based classification of rhizobia. Int J Syst Evol Microbiol 51:2037–2048
Gossmann JA, Markmann K, Brachmann A, Rose LR, Parniske M (2012) Polymorphic infection and organogenesis patterns induced by a Rhizobium leguminosarum isolate from Lotus root nodules are determined by the host genotype. New Phytol 196:561–573
Grant WF, Small E (1996) The origin of the Lotus corniculatus (fabaceae) complex: a synthesis of diverse evidence. Can J Bot 74:975–989
Jarvis BDW, Pankhurst CE, Patel JJ (1982) Rhizobium loti, a new species of legume root nodule bacteria. Int J Syst Bacteriol 32:378–380
Jarvis BDW, van Berkum P, Chen WX, Nour SM, Fernandez MP, Cleyet-Marel JC, Gillis M (1997) Transfer of Rhizobium loti, Rhizobium huakuii, Rhizobium ciceri, Rhizobium mediterraneum, and Rhizobium tianshanense to Mesorhizobium gen. nov. Int J Syst Bacteriol 47:895–898
Kaneko T, Nakamura Y, Sato S, Asamizu E, Kato T, Sasamoto S, Watanabe A, Idesawa K, Ishikawa A, Kawashima K, Kimura T, Kishida Y, Kiyokawa C, Kohara M, Matsumoto M, Matsuno A, Mochizuki Y, Nakayama S, Nakazaki N, Shimpo S, Sugimoto M, Takeuchi C, Yamada M, Tabata S (2000) Complete genome structure of the nitrogen-fixing symbiotic bacterium Mesorhizobium loti. DNA Res 7:331–338
Kelly S, Sullivan J, Ronson C, Tian R, Bräu L, Munk C, Goodwin L, Han C, Woyke T, Reddy T, Huntemann M, Pati A, Mavromatis K, Markowitz V, Ivanova N, Kyrpides N, Reeve W (2014) Genome sequence of the Lotus spp. microsymbiont Mesorhizobium loti strain R7A. Stand Genomic Sci 9:6
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Lorite MJ, Muñoz S, Olivares J, Soto MJ, Sanjuán J (2010) Characterization of strains unlike Mesorhizobium loti that nodulate Lotus spp. in saline soils of Granada, Spain. Appl Environ Microbiol 76:4019–4026
Martínez-Hidalgo P, Ramírez-Bahena MH, Flores-Félix JD, Rivas R, Igual JM, Mateos PF, Martínez-Molina E, León-Barrios M, Peix A, Velázquez E (2015) Revision of the taxonomic status of type strains of Mesorhizobium loti and reclassification of strain USDA 3471T as Mesorhizobium erdmanii sp. nov. and ATCC 33669T Mesorhizobium jarvisii sp. nov. Int J Syst Evol Microbiol 65:1703–1708
Qian J, Parker MA (2002) Contrasting nifD and ribosomal gene relationships among Mesorhizobium from Lotus oroboides in northern Mexico. Syst Appl Microbiol 25:68–73
Ramírez-Restrepo CA, Barry TN (2005) Alternative temperate forages containing secondary compounds for improving sustainable productivity in grazing ruminants. Anim Feed Sci Technol 120:179–201
Reeve W, Sullivan J, Ronson C, Tian R, Bräu L, Davenport K, Goodwin L, Chain P, Woyke T, Lobos E, Huntemann M, Pati A, Mavromatis K, Markowitz V, Ivanova N, Kyrpides N (2014) Genome sequence of the Lotus corniculatus microsymbiont Mesorhizobium loti strain R88B. Stand Genomic Sci 9:3
Rivas R, Velázquez E, Valverde A, Mateos PF, Martínez‐Molina E (2001) A two primers random amplified polymorphic DNA procedure to obtain polymerase chain reaction fingerprints of bacterial species. Electrophoresis 22:1086–1089
Rivas R, Peix A, Mateos PF, Trujillo ME, Martínez-Molina E, Velázquez E (2006) Biodiversity of populations of phosphate solubilizing rhizobia that nodulates chickpea in different Spanish soils. Plant Soil 287:23–33
Rivas R, García-Fraile P, Mateos PF, Martínez-Molina E, Velázquez E (2007) Characterization of xylanolytic bacteria present in the bract phyllosphere of the date palm Phoenix dactylifera. Lett Appl Microbiol 44:181–187
Robledo M, Jiménez-Zurdo JI, Velázquez E, Trujillo ME, Zurdo-Piñeiro JL, Ramírez-Bahena MH, Ramos B, Díaz-Mínguez JM, Dazzo F, Martínez-Molina E, Mateos PF (2008) Rhizobium cellulase CelC2 is essential for primary symbiotic infection of legume host roots. PNAS, USA 105:7064–7069
Saitou N, Nei M (1987) A neighbour-joining method: a new method for reconstructing phylogenetics trees. Mol Biol Evol 4:406–425
Sotelo M, Irisarri P, Lorite MJ, Casaretto E, Rebuffo M, Sanjuán J, Monza J (2011) Diversity of rhizobia nodulating Lotus corniculatus grown in northern and southern regions of Uruguay. Appl Soil Ecol 49:197–207
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The clustal_X windows interface: flexible strategies for multiple sequence alignement aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Vincent JM (1970) The cultivation, isolation and maintenance of rhizobia. In: Vincent JM (ed) A manual for the practical study of root-nodule. Blackwell Scientific, Oxford, pp. 1–13
Acknowledgments
This work was funded by the “Junta de Castilla y León” (Regional Government, Grant SA169U14. MM thanks for PhD fellowship from the “Miguel Casado San José” foundation (Spain) and XCG acknowledges a research grant from the “Kinesis” foundation (Puerto Rico, US).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Marcos-García, M., Menéndez, E., Cruz-González, X. et al. The high diversity of Lotus corniculatus endosymbionts in soils of northwest Spain. Symbiosis 67, 11–20 (2015). https://doi.org/10.1007/s13199-015-0368-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13199-015-0368-5