Abstract
Carnobacterium maltaromaticum is a species of lactic acid bacteria (LAB) that has been isolated from various natural environments. It is well-known for producing a diverse spectrum of bacteriocins with potential biotechnological applications. In the present study, a new psychrotolerant strain of C. maltaromaticum CM22 is reported, isolated from a salmon gut sample and producing a variant of the bacteriocin piscicolin 126 that has been named piscicolin CM22. After identification by 16S rRNA gene, this strain has been genomically characterized by sequencing and assembling its complete genome. Moreover, its bacteriocin was purified and characterized. In vitro tests demonstrated that both the strain and its bacteriocin possess antimicrobial activity against several Gram-positive bacteria of interest in human and animal health, such as Listeria monocytogenes, Clostridium perfringens, or Enterococcus faecalis. However, this bacteriocin did not produce any antimicrobial effect on Gram-negative species. The study of its genome showed the genetic structure of the gene cluster that encodes the bacteriocin, showing a high degree of homology to the gene cluster of piscicolin 126 described in other C. maltaromaticum. Although more studies are necessary concerning its functional properties, this new psychrotolerant strain C. maltaromaticum CM22 and its bacteriocin could be considered an interesting candidate with potential application in agri-food industry.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, European consumers have changed their consumption habits and dietary models towards natural foods, locally produced foods, less industrially processed foods, and foods free of synthetic additives [1]. In addition, there is a growing concern about sustainable animal production and consumption of antibiotic-free animal products [2, 3]. The use of probiotics or their bacteriocins could be a good alternative to the use of synthetic compounds for the livestock and food industry, increasing animal welfare and food safety [4].
Bacteriocins represent a promising alternative to the use of antibiotics or chemical additives; however, their commercial use has not yet expanded significantly, mainly due to regulatory aspects [5]. Bacteriocins usually alter the membrane integrity of susceptible cells [6] and cause the leakage of intracellular solutes, ultimately leading to cell death. Furthermore, bacteriocins usually have a narrow inhibition spectrum that targets genera or species closely related to the producing strain [7]. These compounds are small ribosomally synthesized peptides typically produced by Gram-positive bacteria. They primarily inhibit the growth or cause the demise of other Gram-positive bacteria [8].
Bacteriocins of lactic acid bacteria (LAB) are the most studied ones due to the GRAS (generally recognized as safe) status that most of these species have. Thus, their bacteriocins can be safely used as natural preservatives in food [9]. However, nisin is currently the only bacteriocin that can be used as an authorized additive. Nevertheless, pediocin is being considered for its forthcoming commercial application in food preservation purposes, and its use is safeguarded by several patents [10, 11]. Therefore, there is a big niche for new broad-spectrum molecules in this market [12]. These molecules are of great interest in the agro-food industry, since they can be more easily authorized compared to other synthetically produced chemical compounds. Their aim is to enhance food safety, improve the quality, and extend the shelf life of various food products.
Carnobacterium is a genus of LAB which produces several bacteriocins belonging to classes IIa, IIc, and cyclic bacteriocins such as carnobacteriocin BM1, carnobacteriocin B2, carnobacteriocin A, carnocyclin A, piscicolin 126, and piscicocin CS526 [13]. The recent characterization of these bacteriocins has revealed interesting activity against several food-borne microorganisms, such as Enterococcus faecalis or Listeria monocytogenes [14]. This suggests that studying them could provide novel biopreservation alternatives. Finally, different strains of LAB, including Carnobacterium species with probiosis properties [15,16,17], have been described. The use of these bacteriocinogenic strains is an interesting tool for the agri-food sector. For instance, some LAB have shown an increase in productive parameters in farm animals [18, 19], an improvement in animal welfare by reducing infections [20, 21], and an increase in food safety from the source to the consumer [22, 23].
The aim of this work to explore the antibiosis properties of Carnobacterium maltaromaticum CM22, a bacteriocinogenic strain isolated from salmon. For this purpose, we have sequenced its genome, characterized this strain, and purified and identified its bacteriocins.
Material and Methods
Bacterial Strains and Culture Conditions
Different bacterial strains were used to test antimicrobial activity of C. maltaromaticum CM22, or as positive/negative controls for the specific characteristics investigated (Table 1). The microorganisms used in this study were obtained from the CECT (Spanish Collection of Type Cultures) and wild isolates from our collection (Table 1). All strains were routinely cultivated on either trypticase soy broth (TSB, Scharlau, Barcelona, Spain) at 37 °C and stored, or 4 °C on the respective agar slants.
Screening and Isolation Procedure
The strain C. maltaromaticum CM22 was isolated during a LAB biodiversity study from a fish farm in Puerto Montt (Chile). A salmon (Salmo salar) that weighed 2 kg was hermetically packed in pre-sterile polyethylene bag. The fish was washed with sterile distilled water and then, the surface was cleaned with 70% alcohol. The fish gut was removed under aseptic conditions and washed with 0.85% saline solution. The gut sample was placed in a sterile bag for homogenization in a digester (Stomacher, VWR International, Radnor, Pennsylvania, USA) with sterile buffered peptone water (Scharlau). The digestion product was plated on MRS agar (Scharlau) and incubated anaerobically at 5 °C, 10 °C, and 30 °C for 2–7 days. Colonies were then randomly selected, replicated on MRS agar plates (Scharlau), and incubated for 72 h. Screening was carried out on 180 psychrotolerant strains searching for antimicrobial activity. This activity was examined for antibiosis production by overlaying the strains with 6 mL of soft agar inoculated with 2% stationary-phase cultures of E. faecalis JH2-2 and L. innocua CECT 4030 (Table 1). After incubation at 30 °C, strains that show antibiosis would inhibit the growth of both indicator strains, showing halos around its colony. In this sense, C. maltaromaticum CM22 strain was selected for further analysis due to its highest antibacterial activity against both strains.
Spectrum of Antibiosis
The antimicrobial activity of C. maltaromaticum CM22 strain was studied against some of the most common human and animal pathogens and food spoilage bacteria (both Gram-negative and Gram-positive species) (Table 1) by the agar diffusion test described by Alonso et al. [25]. Target bacteria were grown in Brain Heart Infusion (Scharlau), C. maltaromaticum CM22 strain was grown in MRS (Scharlau), and both cultures were incubated overnight at 30 °C. Target bacteria were spread in Brain Heart Infusion Agar (BHA) Petri dishes using a bacterial suspension adjusted to form a bacterial lawn, and once the plate was dried, drops (15 µL) of a stationary-phase culture of the C. maltaromaticum CM22 strain were dispensed onto the plates. After incubation at 30 °C for 24 h, the plates were examined for the absence/presence of inhibition zones and the results were interpreted as positive ( +) or negative ( −). Tests against each indicator strain were performed in duplicate.
In addition, a well-diffusion technique [26] was carried out with the aim of detecting antibiosis in liquid medium. Therefore, sterile stainless-steel towers 8 mm in diameter × 10 mm in height (Oxford Towers, Scharlab, Barcelona, Spain) were deposited on plates with a base layer of tryptic soy agar (Scharlau). An overlay of tryptic soy agar (0.8% agar) was then melted, inoculated with the target strains, and poured evenly onto the plate. After solidification of the overcoating layer, the cylinders were removed. Then, holes were filled individually with 100 μL of C. maltaromaticum CM22 culture supernatant previously filtered through a 0.22-µm polyethersulfone (PES) syringe-driven filters (Merck Millipore, Carrigtwohill, Ireland). Finally, the plates were incubated aerobically at 37 °C for 24–48 h. After incubation, the clear zones were measured using a calliper. Tests against each indicator strain were performed in duplicate.
Carnobacterium maltaromaticum CM22 Strain Identification and Genomic Characterization
Preliminary identification was based upon phenotypic characteristics, including cell morphology and Gram staining, catalase activity, API50 system (BioMérieux, Craponne, France), and ability to grow at 10 and 45 °C and in the presence of 6.5% (w/v) NaCl.
For the genotypic characterization, genomic DNA was extracted from a pure liquid culture according to Martín-Platero et al. [27]. This genomic DNA was used as a template for 16S rDNA amplification using the WO12 and WO1 primers according to Ogier et al. [28]. PCR reactions were performed in a total volume of 50 µL containing 5 µL of 10X Taq reaction buffer, 1.5 mM of MgCl2, 400 μM of dNTPs, 0.4 μM of the primers WO1 and WO12, 1 U of Taq DNA polymerase (IBIAN Technologies, Zaragoza, Spain), and 1 µL (20–50 ng) of template DNA. The amplification program consisted of an initial denaturing step at 94 °C for 4 min followed by 30 cycles at 94 °C for 30 s, 50 °C for 30 s, and 72 °C for 60 s and a final extension of 72 °C for 2 min. A 700-bp fragment of the 16S rDNA gene containing the V1–V4 variable regions was obtained, purified with a Perfectprep Gel Cleanup kit (Eppendorf, Hamburg, Germany).
The genome was sequenced with the Illumina HiSeq4000 platform by STAB VIDA (Caparica, Portugal). Sequencing libraries were constructed with an insert size of 300 bp, sequencing 150 bp from each end. The assembly of the readings was carried out with SPAdes 3.13.0 [29]. The scaffold was made with MeDuSa 1.6 [30]. GapFiller 1.10 was used to close the scaffold gaps [31]. In addition, the annotation was made with Prokka 1.13.3 [32]. To perform the comparative genomic analysis, Artemis [33] and BLAST [34] were used. Finally, to determine if the C. maltaromaticum CM22 genome was associated with the structural gene of any bacteriocin, a TBLASTN (version 2.10.1 +) was run with a 10−6 e-value threshold [34, 35] between our genomes and bacteriocins from the BACTIBASE databases [36].
Bacteriocin Production, Purification, and Identification
A flask containing 1 L of Brain Heart Infusion (BHI, Scharlab) adjusted to pH 6.5 was inoculated at 2% with a stationary-phase culture of C. maltaromaticum CM22 strain. The cultured flask (1 L) was incubated at 28 °C overnight and centrifuged for 20 min at 4 °C and 3724 rpf (Beckman Coulter, California, USA), collecting the supernatant that was tested by the well-diffusion technique [26].
The bacteriocin present in the medium was recovered by Carboxymethyl-Sephadex CM-25 (Merck Life Science S.L.U. Madrid, Spain) cation exchange chromatography. The recovery of the bacteriocins was carried out using the method described by Abriouel et al. [37] with some modifications. The supernatants at pH 7.0 were mixed with 1 N NaOH, 200 mL of Carboxymethyl Sephadex CM-25 (GE Healthcare, Madrid, Spain) and left stirring for 30 min. Afterwards, the supernatants were removed, and the CM-25 gel was transferred to a cylindrical filtering funnel with a plate porosity of 100–160 microns (Pobel, Madrid, Spain). The gel was washed with 3 volumes of distilled water, followed by 3 volumes of 1 M NaCl and 3 volumes of 1.5 M to elute the adsorbed bacteriocin. During the process, 50-mL fractions were collected and filtered (0.22 μm PES, Merck Millipore, Cork, Ireland), and its activity was measured using the afore mentioned well-diffusion technique.
The fractions that showed antibacterial activity from cation exchange chromatography were repurified by reversed-phase extraction on a C-18 solid support. For this, 5 g of C-18 resin (Waters) was used in a plastic column (2.5 × 10 cm). The C-18 column was washed with 10 mL of isopropanol to acetonitrile 2:1 added with 0.1% trifluoroacetic acid (solvent B) for each gram of resin for equilibration. Subsequently, it was washed with 25 mL of 0.1% trifluoroacetic acid in MilliQ water (solvent A). The active fractions were passed through the column and another wash was carried out with 15 mL of solvent A. Finally, the samples were eluted with increasing concentration of solvent B. Thus, 15 mL 30% solvent B, 15 mL 60% solvent B, and 15 mL solvent B were collected in separate tubes and tested for antimicrobial activity [26]. Those fractions that showed antimicrobial activity were lyophilized and resuspended in solvent A for repurification by reversed-phase high-performance liquid chromatography (RP-HPLC). Purification was performed using a Zorbax Eclipse XBD C18 5 µm particle 4.6 × 150 mm column (Agilent) and an Agilent 1100 analytical HPLC. The machine was operated at 1 mL/min flow rate using a protocol consisting of an equilibration step of 5 min at 5% solvent B, 1 min 5–40% solvent B, 5 min 40% solvent B, and a separation step of 40–95% solvent B gradient for 20 min.
Finally, the peptide nature of the purified bacteriocin was determined by proteolysis with pancreatic trypsin (10 mg/mL in 50 mM Tris–HCl, pH 7.2) (Merck) using a 1:1 enzyme to substrate (v:v) ratio at 37 °C for 2 h. After enzymatic treatment, loss of activity was monitored by well-diffusion against L. innocua.
Nisin was purified from commercial Nisaplin as indicated by Slootweg and coworkers [38] rendering HPLC-grade nisin.
Determination of the Minimal Inhibitory Concentration of Nisin and Piscicolin CM22 in Solid Media
HPLC-purified nisin and piscicolin CM22 were solubilized in 0.05% acetic acid, and their concentrations were measured using the Bio-Rad protein assay. Both peptides were adjusted to a final concentration of 0.6 g/L. Twofold serial dilutions of each peptide were done. Subsequently, 5 µL of each dilution was spotted onto a 7-mL overlay of soft BHI 0.7% agar inoculated with 100 µL of a stationary-phase culture of the susceptible strain L. monocytogenes CECT4032. The minimal inhibitory concentration was determined as the last concentration at which clear inhibition was observed.
Determination of the Molecular Weight and MALDI-TOF Identification
The molecular weight of the purified bacteriocin was determined by MALDI-TOF. For this purpose, a tricine sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) (Sigma-Aldrich, Madrid, Spain) system was used. Electrophoresis was carried out using the Criterion™ Cell (300 V, 20 min). The gel was washed with sterile distilled water and fragments were fixed with 25% (v/v) isopropanol and 10% (v/v) glacial acetic acid (Sigma). One of the fixed gels was stained with Coomassie blue (Sigma) overnight with constant shaking, and then, a solution of water and methanol was used to attenuate the gel. Bands were revealed by a gel documentation system. For the antimicrobial activity test, the other gel fragment without fixation process was washed [39], placed in a sterile Petri dish, and then covered with Brain Heart Infusion agar containing the indicator strain L. innocua CECT4030. Finally, the sample of fixed gel was stored in sterile distilled H2O for identification by matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS) using a Voyager-DE PRO spectrometer from Applied Biosystems.
Results
Isolation and Identification of Carnobacterium maltaromaticum CM22 Strain
The C. maltaromaticum CM22 strain was selected during a study on bacteriocin production among LABs isolated from a Chilean salmon fish farm. This psychrotolerant strain, capable of growing within a temperature range of 5 to 30 °C, exhibited a notable inhibitory spectrum against both target bacteria, E. faecalis JH2-2 and L. innocua CECT 4030. Initially, this strain was identified as C. maltaromaticum based on the API50 system. Afterward, a similarity analysis of the 16S rDNA sequence obtained by PCR amplification confirmed the C. maltaromaticum species identity with a similarity of 925/928 bp (99.7%) compared to the reference sequence M58825 of C. maltaromaticum DSM 20342.
Assessment of Antimicrobial Potential
The antibacterial effect was assayed using agar diffusion tests, and C. maltaromaticum CM22 strain was found to inhibit the growth of 9 out of the 13 tested species (Table 2). The target strains included representative members of food-spoilage bacteria and potential pathogens for animals and humans. C. maltaromaticum CM22 showed antibacterial activity both in solid and in liquid media against all the Gram-positive bacteria tested, except against S. aureus. The antibacterial activity demonstrated against Listeria strains, which presented inhibitory zones of up to 33 mm in diameter using the well-diffusion method, was noteworthy (Supplementary Fig. 1S; Table 2). C. maltaromaticum CM22 did not show antimicrobial activity against any of the Gram-negative bacteria tested. These results suggest the ability of C. maltaromaticum CM22 to produce specific extracellular agents against other Gram-positive strains.
In addition, to establish an initial approximation of the potency of C. maltaromaticum CM22, a comparison was conducted against other described bacteriocins producers known for their potent antilisterial activity (Supplementary Fig. 2S). Arbitrary units per milliliter (AU/mL) were quantified against nisin and pediocin from Lactococcus lactis and Pediococcus acidilactici, respectively (Sigma-Aldrich), using L. monocytogenes CECT 4032 as the target strain. The results showed 640 AU/mL for nisin and 320 AU/mL for pediocin. This suggests, pending further investigation into its spectrum of action, that antimicrobial potency of C. maltaromaticum CM22 could be comparable to that of nisin and slightly superior to that of pediocin.
Purification and Characterization of Antimicrobial Agents
After purification using cation exchange chromatography, both the NaCl eluates and the initial supernatant showed antimicrobial activity, with inhibition halos ranging between 15 and 20 mm in diameter against L. innocua CECT 4030. Samples that showed activity after chromatography with CM-25 were then subsequently pooled and further purified by reversed-phase chromatography with C-18. The different eluted fractions were assayed, showing inhibition halos of 19–20 mm in diameter against both E. faecalis and L. innocua, respectively.
The C18 fractions with the highest antimicrobial activity were lyophilized, resuspended in solvent A, then mixed, and HPLC purified. The different peaks in the chromatogram were collected separately, and only the peak with 18-min retention time showed antimicrobial activity and was further characterized (Supplementary Fig. 3S).
The molecular mass of the different eluted fractions from C-18 was estimated by polyacrylamide gel electrophoresis under denaturing conditions (tricine SDS-PAGE) and subsequent antimicrobial activity. Only those major fractions with antimicrobial activity were analyzed. Polyacrylamide gel electrophoresis showed a single band corresponding to a substance with a molecular weight lower than 6 kDa (Supplementary Fig. 4S). Furthermore, the antimicrobial susceptibility test, using the overcoat technique with L. innocua CECT 4030 as an indicator strain corroborated the presence of an inhibitory substance of peptide nature,with a low molecular weight.
The determination of the molecular mass of the active fractions obtained by RP-HPLC (at 18 min) was performed by MALDI-TOF mass spectrometry. A well-defined peak with a mass of 4428.1 Da (Fig. 1) was observed, which presents a great similarity with literature data piscicolin 126 (4416.6 Da). Finally, in order to confirm the nature of the molecule, the treatment of the filtered supernatant with a protease (pancreatic trypsin, Merck) in proportion (1:1) (v:v) resulted in the total loss of antimicrobial activity (data not shown).
The purified fraction maintains the activity against the sensor strains included in Table 2. Furthermore,the potency of purified piscicolin CM22 was assayed against L. monocytogenes and compared to that of the reference bacteriocin nisin. For this, both compounds were solubilized at the same concentration and serial dilutions were spotted onto the susceptible strain. L. monocytogenes was inhibited by both compounds, with piscicolin CM22 remaining active up to 4.7 mg/L and 2.3 mg/L against the strains CECT4032 and CECT5366, respectively, and nisin at 18.7 mg/L under these conditions. Additionally, we conducted the same test on the intermediate susceptible strain L. aquatica obtaining a minimal inhibitory concentration of 300 mg/L and 37,5 mg/L of piscicolin CM22 and nisin, respectively. These data are consistent with the higher potency of type IIa bacteriocins against Listeria strains as reported in literature.
Genomic Analysis of Carnobacterium maltaromaticum CM22
After sequencing, assembly, and annotation, C. maltaromaticum CM22 genome showed a size of 4.08 Mb, with a GC content of 33.59%. Additionally, 3797 protein-coding DNA sequences (CDS), 62 tRNA genes, and 10 rRNA were found in its genome.
Gene Ontology (GO) analysis of the protein-coding genes assigned 5585 GO terms to 2031 (61%) genes. Of the total GO terms, 605 (10.8%), 3209 (57.5%), and 1771 (31.7%) were assigned to cellular components, molecular functions, and biological processes respectively. Membrane components, DNA binding, and regulation of DNA-template transcription were the most abundant terms among each category (Fig. 2A–C). InterProScan detected 3329 proteins from C. maltaromaticum CM22 genome. A total of 10,610 families were assigned to 3125 proteins (93.9%, 3125/3329), where P-loop containing nucleoside triphosphate hydrolase family (IPR027417) obtained the highest assignation, followed by winged helix-like DNA-binding domain superfamily (IPR036388) and the AAA + ATPase domain (IPR003593) (Fig. 2D).
Results of TBLASTN against the BACTIBASE database showed that genes for piscicolin 126 variant, namely piscicolin CM22, and carnobacteriocin BM1 bacteriocins were present in C. maltaromaticum CM22 genome. The class IIa bacteriocin carnobacteriocin BM1 (6572 Da) gene cluster was formed by two open reading frames (ORFs), cbnBM1 and cbiBM1 (Fig. 3A). cbnBM1 corresponded to the structural gene and would be responsible for the production of a protein with 61 amino acid residues, while cbiBM1 encoded an immunity protein [14, 40,41,42].
Genetic organization of carnobacteriocin BM1 and piscicolin CM22 bacteriocins. A Two identified open reading frames (ORFs) are depicted in the carnobacteriocin BM1 gene cluster: cbnBM1 (dark blue arrow) corresponds to the structural gene while cbiBM1 (light blue) encodes an immunity protein. B Gene cluster of piscicolin 126. A total of 8 ORFs are depicted: pisI (violet arrow) is the putative accessory protein; pisA (pink arrow) is the structural gene for the bacteriocin; pisN (red arrow) codes for a putative induction peptide; pisK and pisR (light and dark orange arrows) encode an histidine kinase and a response regulator respectively; pisT (light green arrow) is the ABC transporter, and pisE (dark green arrow) encodes a putative transport accessory protein
Piscicolin CM22 gene cluster contained at least eight ORFs (Fig. 3B). The structural gene (pisA) encoded a protein of 62 amino acid residues of which the first 21 corresponded to a signal peptide. pisN encoded a putative induction peptide (IP) that might be involved in regulating piscicolin 126 production. pisK and pisR were part of a two-component system where pisK encoded a histidine kinase and pisR encoded for a response regulator. pisT encoded for the bacteriocin ABC transporter, pisE encoded for the putative transport accessory protein, and pisI encoded for the immunity protein. Despite the presence of the genetic sequence responsible for the synthesis of carnobacteriocin BM1 in its genome, MALDI-TOF analysis of the active fraction did not detect the presence of this bacteriocin. This fact may be attributed to the absence of BM1 production or its exclusive synthesis in specific circumstances that are unknown at the moment, such as the different temperature conditions [43].
The genomic analysis of C. maltaromaticum CM22 revealed a genome size slightly larger than those previously described for this genus, which typically ranges from 3.33 to 3.87 [44, 45]. The increased chromosomal size of C. maltaromaticum CM22 may contribute to its ability to colonize multiple and diverse habitats compared to other carnobacterial species [46].
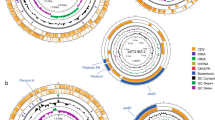
Gursky et al. [42] described and analyzed the gene clusters responsible for the production of piscicolin 126 in the C. maltaromaticum UAL126 strain (accession number AY812745.1) and in the C. piscicola JG126 strain (accession number AF275938.1), respectively. A comparison between the piscicolin gene clusters of C. maltaromaticum CM22, C. maltaromaticum UAL26, and C. piscicola JG126, using by blast alignment, revealed 93.36% and 93.17 identity at the DNA level (Fig. 4). The gene cluster was arranged in the same direction for each of the ORFs identified. Individually, each protein of the gene cluster showed high similarity (> 85%) (Table 3).
Homology between piscicolin 126 gene cluster of A Carnobacterium maltaromaticum UAL26, B C. maltaromaticum CM22, C Carnobacterium piscicola JG126. The color scale represents the similarity between the different genes: light red and blue = 90% similarity, while dark red and blue indicate up to 100% similarity; red represents for matches in the same direction and blue for inverted matches. The entire region of the cluster shows similarity in all three strains
Discussion
Bacteriocins synthesized by LABs have gained significant attention for their potential applications as antibiotic alternatives and novel biopreservatives. Additionally, there is ongoing exploration into the potential use of these producer strains as probiotics [47]. In this work, we have isolated the C. maltaromaticum CM22 strain with a significant antimicrobial activity linked to the production capacity of piscicolin, which could suggest its potential technological use in the agri-food industry. The wide range of foods in which C. maltaromaticum has been identified renders this strain highly compelling for potential applications in the food industry. Carnobacterium strains have been mostly isolated from food, animal sources, and a wide range of environments [13]. Consistent with our results, other authors have reported the isolation of Carnobacterium strains from aquatic environments, including salmon [48] and extreme environments such as Antarctic ecosystems [49,50,51,52]. Some fish products have also been described as a source of isolation of C. maltaromaticum strains like vacuum-packed tuna [53], smoked surubim [22], and smoked salmon [54]. In addition, other authors have reported its isolation in meat products [55, 56].
Furthermore, in our investigation, we purified and characterized the inhibitory substance produced by C. maltaromaticum CM22, thereby confirming its identity as piscicolin CM22, a variant of piscicolin 126. Additionally, a genome study of the CM22 strain has been conducted. Gursky et al. [43] also analyzed the genome of the strains C. maltaromaticum JG126 and C. maltaromaticum UAL26 and observed that piscicolin 126 was encoded by an operon that included genes cataloged as ABC transporters, genes responsible for immunity, coding for histidine kinase protein, along with other accessory genes. Those findings show significant homology with our genomic analysis of C. maltaromaticum CM22, where an analogue of the piscicolin 126 cluster is maintained. This cluster comprises genes coding for bacteriocin immunity (MILBGHNJ_00131), histidine kinase (MILBGHNJ_00128), and ABC transporters (MILBGHNJ_00126). The proximity found between the pisA gene that encoded piscicolin CM22 (131,370–131558 bp) and the gene responsible for bacteriocin immunity (131,814–132,110 bp) is noteworthy.
The mass of piscicolin 126 produced by C. maltaromaticum JG126, as determined by MALDI-TOF, is 4416.6 Da [43], compared to the 4428.1 Da (theoretical Mw for the user-entered sequence 4431.93) obtained for the same bacteriocin produced by C. maltaromaticum CM22. The percentage of identity between these two proteins is around 87%, which explains this small variation (Supplementary Fig. 5S). Similar to the majority of described bacteriocins, the hydrophobicity plot of piscicolin CM22 suggests its nature as a hydrophobic peptide, with approximately 34.1% of its amino acid residues being hydrophobic. Additionally, it exhibits a notable basic character, with a predicted isoelectric point (pI) of 9.19.
Also, the genomes of strains UAL26 and JG126 had structural genes for the bacteriocin BM1. This bacteriocin contains the characteristic YGNGV motif of the family of class II bacteriocins in the N-terminal region [42, 57, 58]. A congruent result was observed compared to C. maltaromaticum CM22, wherein the bacteriocin BM1 genes are also present; however, bacteriocin BM1 production could not be detected under the assayed conditions.
LAB strains often harbor the genetic components necessary to produce multiple bacteriocins that can synergistic or additively work together, resulting in a stronger inhibitory effect against susceptible strains when expressed concurrently [59]. Cintas et al. [60] showed that LABs frequently possess genetic elements within their genomes for the production of multiple bacteriocins, since they tend to act synergistically, leading to a more pronounced inhibitory effect. However, other authors have argued that certain bacteriocins may exhibit antagonistic effect [61]. These mutually exclusive alternatives open the door to future research, exploring the production conditions of the bacteriocins piscicolin CM22 and BM1 and evaluating the effect that one exerts over the other. The genetic organization of piscicolin 126 on gene cluster depicted in Fig. 3 has been reported to be conserved between Carnobacterium and other LAB species such as Enterococcus [62]. Nevertheless, piscicolin 126 and carnobacteriocin BM1 clusters are not always present together in the same strains. Quadri et al. [42] reported that Carnobacterium piscicola LV17B strain is another producer of carnobacteriocin BM1 that does not contain the piscicolin cluster. In conjunction with the study conducted by Leisner et al. [13], both works showed that the structure of carnobacteriocin BM1 gene cluster is highly similar to that found in the present work.
In another study by Quadri et al. [41], the carnobacteriocin B2 gene cluster includes a structural gene, cbnB2, and a gene coding for bacteriocin immunity, cbiB2, providing autoimmunity not only to its own carnobacteriocin B2 but also to some other antimicrobials. These findings exhibit homology with the gene cluster of the carnobacteriocin BM1, where a structural gene (MILBGHNJ_02530) and a coding gene for immunity (MILBGHNJ_02531) are present.
In our work, we demonstrated the antimicrobial activity of C. maltaromaticum CM22 associated with its piscicolin CM22, revealing a broad spectrum of action in both in solid and liquid media against several Gram-positive bacteria relevant to human and animal health [63]. Given its antibacterial potential against both pathogenic and probiotic microorganisms, it is imperative to study and monitor the impact of these bacteriocin-producing strains on the gut microbiota. Preliminary studies, such as gut simulation trials on the target species, could be conducted [64, 65]. In addition to its antibacterial properties, this strain exhibits the capability to grow at 5 °C. This, along with its ability to grow at low temperatures, could facilitate its potential application as bioprotective agent for refrigerated food, inhibiting the growth of psychrotolerant pathogenic bacteria such as L. monocytogenes [22, 56] or other spoilage bacteria [44, 66]. Furthermore, C. maltaromaticum is recognized as a generally recognized as safe (GRAS) bacterium in the United States and it is considered a microorganism with beneficial technological use in the EU [67]. Although commercial products with this bacterium such as Micocin®, formed by a culture of the C. maltaromaticum CB1 strain, are already available for the preservation of meat products [12], C. maltaromaticum is not yet included in the European qualified presumption of safety (QPS) list due to some strains being associated with diseases in fish [68,69,70]. In contrast, some authors have highlighted the probiotic potential of Carnobacterium in different fish species, such as rainbow trout (Oncorhynchus mykiss) [52] and salmon (S. salar) [71]. For example, some authors have described the benefits of using Carnobacterium strains as probiotics in the diet of rainbow trout, exhibiting the ability to control pathogens such as Aeromonas salmonicida or to improve the immune response of fish [52, 72]. Recently, Puvanendran et al. [18] reported the ability of a strain of Carnobacterium divergens to increase growth parameters and disease resistance in cod larvae (Gadus morhua). In the present work, C. maltaromaticum CM22 demonstrated antimicrobial activity against a fish pathogen such as Streptococcus phocae subsp. salmonis [73]. Therefore, its potential use as a probiotic in the aquaculture sector is a promising avenue for research.
Conclusion
In conclusion, our work has unveiled a novel psychrotolerant C. maltaromaticum CM22 strain, isolated from S. salar, capable of producing the bacteriocin piscicolin CM22. Further studies are needed to investigate its safety properties and potential applications in the food preservation sector, either as bioprotective culture for refrigerated food or through the applications of its purified bacteriocin. Additionally, its probiotic properties could be explored for potential applications as an active ingredient in feed production.
Data Availability
Bioproject PRJNA1004195 included the assembled genome of C. maltaromaticum CM22 deposited at NCBI under accession number JAVBVO00000000.1.
References
Esqué PC (2018) Cooking feeding future. Nutr Hosp 35:49–51. https://doi.org/10.20960/NH.2125
Clavijo V, Flórez MJV (2018) The gastrointestinal microbiome and its association with the control of pathogens in broiler chicken production: a review. Poult Sci 97:1006–1021. https://doi.org/10.3382/ps/pex359
Caprarulo V, Giromini C, Rossi L (2021) Chestnut and quebracho tannins in pig nutrition: the effects on performance and intestinal health. Animal 15:100064. https://doi.org/10.1016/j.animal.2020.100064
Maldonado Galdeano C, Cazorla SI, Lemme Dumit JM, Vélez E, Perdigón G (2019) Beneficial effects of probiotic consumption on the immune system. Annals of Nutrition Metabolism: Clinical and Experimental 74:115–124. https://doi.org/10.1159/000496426
Gradisteanu Pircalabioru G, Popa LI, Marutescu L, Gheorghe I, Popa M, Czobor Barbu I, Cristescu R, Chifiriuc M-C (2021) Bacteriocins in the era of antibiotic resistance: rising to the challenge. Pharmaceutics 13:196. https://doi.org/10.3390/pharmaceutics13020196
Darbandi A, Asadi A, Mahdizade Ari M, Ohadi E, Talebi M, Halaj Zadeh M, Darb Emamie A, Ghanavati R, Kakanj M (2022) Bacteriocins: properties and potential use as antimicrobials. J Clin Lab Anal 36:e24093. https://doi.org/10.1002/jcla.24093
Mokoena MP (2017) Lactic acid bacteria and their bacteriocins: classification, biosynthesis and applications against uropathogens: a mini-review. Mol Cells 22:1255. https://doi.org/10.3390/molecules22081255
Huang Y, Ye K, Yu K, Wang K, Zhou G (2016) The potential influence of two Enterococcus faecium on the growth of Listeria monocytogenes. Food Control 67:18–24. https://doi.org/10.1016/j.foodcont.2016.02.009
Lo R, Bansal N, Turner MS (2018) Characterisation of Lactococcus lactis isolates from herbs, fruits and vegetables for use as biopreservatives against Listeria monocytogenes in cheese. Food Control 85:472–483. https://doi.org/10.1016/j.foodcont.2017.09.036
Khorshidian N, Khanniri E, Mohammadi M, Mortazavian AM, Yousefi M (2021) Antibacterial activity of pediocin and pediocin-producing bacteria against Listeria monocytogenes in meat products. Front Microbiol 12:709959. https://doi.org/10.3389/fmicb.2021.709959
Reis J, Paula A, Casarotti S, Penna A (2012) Lactic acid bacteria antimicrobial compounds: characteristics and applications. Food Eng Rev 4:124–140. https://doi.org/10.1007/s12393-012-9051-2
Gálvez A, López RL, Abriouel H, Valdivia E, Omar NB (2008) Application of bacteriocins in the control of foodborne pathogenic and spoilage bacteria. Crit Rev Biotechnol 28:125–152. https://doi.org/10.1080/07388550802107202
Leisner JJ, Laursen BG, Prévost H, Drider D, Dalgaard P (2007) Carnobacterium: positive and negative effects in the environment and in foods. FEMS Microbiol Rev 31:592–613. https://doi.org/10.1111/j.1574-6976.2007.00080.x
Martin-Visscher LA, van Belkum MJ, Garneau-Tsodikova S, Whittal RM, Zheng J, McMullen LM, Vederas JC (2008) Isolation and characterization of carnocyclin A, a novel circular bacteriocin produced by Carnobacterium maltaromaticum UAL307. Appl Environ Microbiol 74:4756–4763. https://doi.org/10.1128/AEM.00817-08
Kassaa IA, Rafei R, Moukhtar M, Zaylaa M, Gharsallaoui A, Asehraou A, El Omari K, Shahin A, Hamze M, Chihib N-E (2019) LABiocin database: a new database designed specifically for lactic acid bacteria bacteriocins. Int J Antimicrob Agents 54:771–779. https://doi.org/10.1016/j.ijantimicag.2019.07.012
Meade E, Slattery MA, Garvey M (2020) Bacteriocins, potent antimicrobial peptides and the fight against multi drug resistant species: resistance is futile? Antibiot Chemother 9:32. https://doi.org/10.3390/antibiotics9010032
Furlaneto-Maia L, Ramalho R, Rocha KR, Furlaneto MC (2020) Antimicrobial activity of enterocins against Listeria sp. and other food spoilage bacteria. Biotechnol Lett 42:797–806. https://doi.org/10.1007/s10529-020-02810-7
Puvanendran V, Rud I, Breiland MSW, Arnesen J-A, Axelsson L (2021) Probiotic Carnobacterium divergens increase growth parameters and disease resistance in farmed Atlantic cod (Gadus morhua) larvae without influencing the microbiota. Aquaculture 532:736072. https://doi.org/10.1016/j.aquaculture.2020.736072
Peralta-Sanchez JM, Martin-Platero AM, Ariza-Romero JJ, Rabelo-Ruiz M, Zurita-Gonzalez MJ, Baños A, Rodriguez-Ruano SM, Maqueda M, Valdivia E, Martinez-Bueno M (2019) Egg production in poultry farming is improved by probiotic bacteria. Front Microbiol 10:1042. https://doi.org/10.3389/fmicb.2019.01042
Baños A, Ariza JJ, Nuñez C, Gil-Martínez L, García-López JD, Martínez-Bueno M, Valdivia E (2019) Effects of Enterococcus faecalis UGRA10 and the enterocin AS-48 against the fish pathogen Lactococcus garvieae. Studies in vitro and in vivo. Food Microbiol 77:69–77. https://doi.org/10.1016/j.fm.2018.08.002
Józefiak D, Sip A, Rutkowski A, Rawski M, Kaczmarek S, Wołuń-Cholewa M, Engberg R, Højberg O (2012) Lyophilized Carnobacterium divergens AS7 bacteriocin preparation improves performance of broiler chickens challenged with Clostridium perfringens. Poult Sci 91:1899–1907. https://doi.org/10.3382/ps.2012-02151
Alves VF, De Martinis EC, Destro MT, Vogel BF, Gram L (2005) Antilisterial activity of a Carnobacterium piscicola isolated from Brazilian smoked fish (surubim [Pseudoplatystoma sp.]) and its activity against a persistent strain of Listeria monocytogenes isolated from surubim. J Food Prot 68:2068–2077. https://doi.org/10.4315/0362-028x-68.10.2068
Dos Reis FB, De Souza VM, Thomaz MR, Fernandes LP, De Oliveira WP, De Martinis EC (2011) Use of Carnobacterium maltaromaticum cultures and hydroalcoholic extract of Lippia sidoides Cham. against Listeria monocytogenes in fish model systems. Int J Food Microbiol 146:228–234. https://doi.org/10.1016/j.ijfoodmicro.2011.02.012
Yagi Y, Clewell D (1980) Recombination-deficient mutant of Streptococcus faecalis. J Bacteriol Parasitol 143:966–970. https://doi.org/10.1128/jb.143.2.966-970.1980
Alonso S, Carmen Castro M, Berdasco M, de la Banda IG, Moreno-Ventas X, de Rojas AH (2019) Isolation and partial characterization of lactic acid bacteria from the gut microbiota of marine fishes for potential application as probiotics in aquaculture. Probiotics Antimicrobial Proteins 11:569–579. https://doi.org/10.1007/s12602-018-9439-
Tagg J, McGiven A (1971) Assay system for bacteriocins. Appl Microbiol Biotechnol 21:943–943. https://doi.org/10.1128/am.21.5.943-943.1971
Martín-Platero AM, Valdivia E, Maqueda M, Martínez-Bueno M (2007) Fast, convenient, and economical method for isolating genomic DNA from lactic acid bacteria using a modification of the protein “salting-out” procedure. Anal Biochem 366:102–104. https://doi.org/10.1016/j.ab.2007.03.010
Ogier JC, Son O, Gruss A, Tailliez P, Delacroix-Buchet A (2002) Identification of the bacterial microflora in dairy products by temporal temperature gradient gel electrophoresis. Appl Environ Microbiol 68:3691–3701. https://doi.org/10.1128/aem.68.8.3691-3701.2002
Nurk S, Bankevich A, Antipov D, Gurevich A, Korobeynikov A, Lapidus A, Prjibelsky A, Pyshkin A, Sirotkin A, Sirotkin Y (2013) Assembling genomes and mini metagenomes from highly chimeric reads. Annual International Conference on Research in Computational Molecular Biology. RECOMB 2013. Lecture Notes in Computer Science, vol 7821. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-37195-0_13
Bosi E, Donati B, Galardini M, Brunetti S, Sagot M-F, Lió P, Crescenzi P, Fani R, Fondi M (2015) MeDuSa: a multi-draft based scaffolder. Bioinformatics 31:2443–2451. https://doi.org/10.1093/bioinformatics/btv171
Boetzer M, Pirovano W (2012) Toward almost closed genomes with GapFiller. Genome Biol 13:R56. https://doi.org/10.1186/gb-2012-13-6-r56
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Carver T, Harris SR, Berriman M, Parkhill J, McQuillan JA (2012) Artemis: an integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 28:464–469. https://doi.org/10.1093/bioinformatics/btr703
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinformatics 10:1–9. https://doi.org/10.1186/1471-2105-10-421
Hammami R, Zouhir A, Ben Hamida J, Fliss I (2007) BACTIBASE: a new web-accessible database for bacteriocin characterization. BMC Microbiol 7:1–6. https://doi.org/10.1186/1471-2180-7-8
Abriouel H, Valdivia E, Martınez-Bueno M, Maqueda M, Gálvez A (2003) A simple method for semi-preparative-scale production and recovery of enterocin AS-48 derived from Enterococcus faecalis subsp. liquefaciens A-48-32. J Microbiol Methods 55:599–605. https://doi.org/10.1016/s0167-7012(03)00202-1
Slootweg JC, Liskamp RM, Rijkers DT (2013) Scalable purification of the lantibiotic nisin and isolation of chemical/enzymatic cleavage fragments suitable for semi-synthesis. J Pept Sci 19:692–699. https://doi.org/10.1002/psc.2551
Bhunia AK, Johnson M, Ray B (1987) Direct detection of an antimicrobial peptide of Pediococcus acidilactici in sodium dodecyl sulfate-polyacrylamide gel electrophoresis. J Ind Microbiol Biotechnol Adv 2:319–322. https://doi.org/10.1002/(SICI)1097-0061
Herbin S, Mathieu F, Brulé F, Branlant C, Lefebvre G, Lebrihi A (1997) Characteristics and genetic determinants of bacteriocin activities produced by Carnobacterium piscicola CP5 isolated from cheese. Curr Microbiol 35:319–326. https://doi.org/10.1007/s002849900262
Quadri LE, Yan LZ, Stiles ME, Vederas JC (1997) Effect of amino acid substitutions on the activity of carnobacteriocin B2: overproduction of the antimicrobial peptide, its engineered variants, and its precursor in Escherichia coli. J Biol Chem 272:3384–3388. https://doi.org/10.1074/jbc.272.6.3384
Quadri L, Sailer M, Roy KL, Vederas JC, Stiles ME (1994) Chemical and genetic characterization of bacteriocins produced by Carnobacterium piscicola LV17B. J Biol Chem 269:12204–12211. https://doi.org/10.1016/S0021-9258(17)32702-3
Gursky LJ, Martin NI, Derksen DJ, Van Belkum MJ, Kaur K, Vederas JC, Stiles ME, McMullen LM (2006) Production of piscicolin 126 by Carnobacterium maltaromaticum UAL26 is controlled by temperature and induction peptide concentration. Arch Microbiol 186:317–325. https://doi.org/10.1007/s00203-006-0147-z
Iskandar CF, Borges F, Taminiau B, Daube G, Zagorec M, Remenant B, Leisner JJ, Hansen MA, Sørensen SJ, Mangavel C (2017) Comparative genomic analysis reveals ecological differentiation in the genus Carnobacterium. Front Microbiol 8:357. https://doi.org/10.3389/fmicb.2017.00357
Leisner J, Hansen M, Larsen M, Hansen L, Ingmer H, Sørensen S (2012) The genome sequence of the lactic acid bacterium, Carnobacterium maltaromaticum ATCC 35586 encodes potential virulence factors. Int J Food Microbiol 152:107–115. https://doi.org/10.1016/j.ijfoodmicro.2011.05.012
Cailliez-Grimal C, Chaillou S, Anba-Mondoloni J, Loux V, Afzal MI, Rahman A, Kergourlay G, Champomier-Vergès M-C, Zagorec M, Dalgaard P (2013) Complete chromosome sequence of Carnobacterium maltaromaticum LMA 28. Genome Announc 1:1. https://doi.org/10.1128/genomea.00115-12
Todorov SD, Popov I, Weeks R, Chikindas ML (2022) Use of bacteriocins and bacteriocinogenic beneficial organisms in food products: benefits, challenges, concerns. Foods 11:3145. https://doi.org/10.3390/foods11193145
Pastorino P, Colussi S, Pizzul E, Varello K, Menconi V, Mugetti D, Tomasoni M, Esposito G, Bertoli M, Bozzetta E, Dondo A, Acutis P, Prearo M (2021) The unusual isolation of carnobacteria in eyes of healthy salmonids in high-mountain lakes. Sci Rep 11:2314. https://doi.org/10.1038/s41598-021-82133-3
Franzmann P, Höpfl P, Weiss N, Tindall B (1991) Psychrotrophic lactic acid-producing bacteria from anoxic waters in Ace Lake Antarctica Carnobacterium funditum sp nov and Carnobacterium alterfunditum sp nov. Arch Microbiol 156:255–262. https://doi.org/10.1007/BF00262994
Bratina BJ, Stevenson BS, Green WJ, Schmidt TM (1998) Manganese reduction by microbes from oxic regions of the Lake Vanda (Antarctica) water column. Appl Environ Microbiol Rep 64:3791–3797. https://doi.org/10.1128/AEM.64.10.3791-3797.1998
Zhu S, Lin D, Xiong S, Wang X, Xue Z, Dong B, Shen X, Ma X, Chen J, Yang J (2018) Carnobacterium antarcticum sp. nov., a psychrotolerant, alkaliphilic bacterium isolated from sandy soil in Antarctica. Int J Syst Evol Microbiol 68:1672–1677. https://doi.org/10.1099/ijsem.0.002727
Kim DH, Brunt J, Austin B (2007) Microbial diversity of intestinal contents and mucus in rainbow trout (Oncorhynchus mykiss). J Appl Microbiol 102:1654–1664. https://doi.org/10.1111/j.1365-2672.2006.03185.x
Emborg J, Laursen BG, Dalgaard P (2005) Significant histamine formation in tuna (Thunnus albacares) at 2 C—effect of vacuum-and modified atmosphere-packaging on psychrotolerant bacteria. Int J Food Microbiol 101:263–279. https://doi.org/10.1016/j.ijfoodmicro.2004.12.001
Lakshmanan R, Dalgaard P (2004) Effects of high-pressure processing on Listeria monocytogenes, spoilage microflora and multiple compound quality indices in chilled cold-smoked salmon. J Appl Microbiol 96:398–408. https://doi.org/10.1046/j.1365-2672.2004.02164.x
Larrouture-Thiveyrat C, Pepin M, Leroy-Sétrin S, Montel M-C (2003) Effect of Carnobacterium piscicola on aroma formation in sausage mince. Meat Sci 63:423–426. https://doi.org/10.1016/s0309-1740(02)00083-9
Danielski GM, Imazaki PH, de Andrade Cavalari CM, Daube G, Clinquart A, de Macedo REF (2020) Carnobacterium maltaromaticum as bioprotective culture in vitro and in cooked ham. Meat Sci 162:108035. https://doi.org/10.1016/j.meatsci.2019.108035
Fimland G, Eijsink VG, Nissen-Meyer J (2002) Mutational analysis of the role of tryptophan residues in an antimicrobial peptide. Biochem 41:9508–9515. https://doi.org/10.1021/bi025856
Fimland G, Johnsen L, Dalhus B, Nissen-Meyer J (2005) Pediocin-like antimicrobial peptides (class IIa bacteriocins) and their immunity proteins: biosynthesis, structure, and mode of action. J Pept Sci 11:688–696. https://doi.org/10.1002/psc.699
Martin NI, Sprules T, Carpenter MR, Cotter PD, Hill C, Ross RP, Vederas JC (2004) Structural characterization of lacticin 3147, a two-peptide lantibiotic with synergistic activity. Biochem 43:3049–3056. https://doi.org/10.1021/bi0362065
Cintas LM, Casaus P, Herranz C, Håvarstein LS, Holo H, Hernández PE, Nes IF (2000) Biochemical and genetic evidence that Enterococcus faecium L50 produces enterocins L50A and L50B, the sec-dependent enterocin P, and a novel bacteriocin secreted without an N-terminal extension termed enterocin Q. J Bacteriol Parasitol 182:6806–6814. https://doi.org/10.1128/JB.182.23.6806-6814.2000
Mulet-Powell N, Lacoste-Armynot A, Vinas M, De Buochberg MS (1998) Interactions between pairs of bacteriocins from lactic bacteria. J Food Prot 61:1210–1212. https://doi.org/10.4315/0362-028x-61.9.1210
Ishibashi N, Himeno K, Masuda Y, Perez RH, Iwatani S, Zendo T, Wilaipun P, Leelawatcharamas V, Nakayama J, Sonomoto K (2014) Gene cluster responsible for secretion of and immunity to multiple bacteriocins, the NKR-5-3 enterocins. Appl Environ Microbiol Rep 80:6647–6655. https://doi.org/10.1128/AEM.02312-14
Hammi I, Delalande F, Belkhou R, Marchioni E, Cianferani S, Ennahar S (2016) Maltaricin CPN, a new class IIa bacteriocin produced by Carnobacterium maltaromaticum CPN isolated from mould-ripened cheese. J Appl Microbiol 121:1268–1274. https://doi.org/10.1111/jam.13248
Rychen G, Aquilina G, Azimonti G, Bampidis V, de Lourdes Bastos M, Bories G, Chesson A, Cocconcelli PS, Flachowsky G (2018) Guidance on the characterisation of microorganisms used as feed additives or as production organisms. EFSA J 16(3):5206. https://doi.org/10.2903/j.efsa.2018.5206
Lone A, Mottawea W, Mehdi Y, Hammami R (2022) Bacteriocinogenic probiotics as an integrated alternative to antibiotics in chicken production-why and how? Crit Rev Food Sci Nutr 62:8744–8760. https://doi.org/10.1080/10408398.2021.1932722
Orihuel A, Bonacina J, Vildoza MJ, Bru E, Vignolo G, Saavedra L, Fadda S (2018) Biocontrol of Listeria monocytogenes in a meat model using a combination of a bacteriocinogenic strain with curing additives. Food Res Int 107:289–296. https://doi.org/10.1016/j.foodres.2018.02.043
Bourdichon F, Casaregola S, Farrokh C, Frisvad JC, Gerds ML, Hammes WP, Harnett J, Huys G, Laulund S, Ouwehand A (2012) Food fermentations: microorganisms with technological beneficial use. Int J Food Microbiol 154:87–97. https://doi.org/10.1016/j.ijfoodmicro.2011.12.030
Mohamed MH, Abogabal II, Einas H (2017) Isolation and molecular characterization of β-hemolytic Carnobacterium maltaromaticum from naturally infected longnose parrot fish (hipposcarusharid) in hurghada red sea, Egypt. Int J Innov Stud Aquat Biol Fish 3:1–8. https://doi.org/10.20431/2454-7670.0301001
Steele LM, Lowe CG, Okihiro MS, Dillon JG, Berlemont R (2019) Pseudogenization, genome streamlining and specific gene repertoire landmark the genomes of Carnobacterium maltaromaticum isolated from diseased sharks. bioRxiv 600684. https://doi.org/10.1101/600684
Roh H, Kim BS, Lee MK, Park CI, Kim DH (2020) Genome-wide comparison of Carnobacterium maltaromaticum derived from diseased fish harbouring important virulence-related genes. J Fish Dis 43:1029–1037. https://doi.org/10.1111/jfd.13208
Robertson P, O’Dowd C, Burrells C, Williams P, Austin B (2000) Use of Carnobacterium sp. as a probiotic for Atlantic salmon (Salmo salar L.) and rainbow trout (Oncorhynchus mykiss, Walbaum). Aquaculture 185:235–243. https://doi.org/10.1016/S0044-8486(99)00349-X
Irianto A, Austin B (2002) Probiotics in aquaculture. J Fish Dis 25:633–642. https://doi.org/10.1046/j.1365-2761.2002.00422.x
Romalde JL, Ravelo C, Valdés I, Magariños B, de La Fuente E, San Martín C, Avendaño-Herrera R, Toranzo AE (2008) Streptococcus phocae, an emerging pathogen for salmonid culture. Vet Microbiol 130:198–207. https://doi.org/10.1016/j.vetmic.2007.12.021
Acknowledgements
We would like to acknowledge the work of Jose Manuel García-Madero and Alicia González Gragera in editing the text.
Funding
Funding for open access publishing: Universidad de Granada/CBUA. This study was financially supported by FEDER-INNTERCONECTA-CDTI (Centro para el Desarrollo Tecnológico Industrial), Spanish Ministry of Economy and Competitiveness; (ALQUABIOTIC project, ITC-20181099). We thank the funding through Universidad de Granada-Junta de Andalucía (Programa Operativo FEDER Andalucía 2014–2020), grant number B-BIO-604-UGR20.
Author information
Authors and Affiliations
Contributions
Conceptualization E.G.-G., A.B and M.M.-B.; methodology, E.G.-G., I.G.-H., C.T.-P., A.B. and J.D.G.-L.; software, C.T.-P., A.M.M.-P. and J.M.P.-S.; validation, J.F.-J., A.M.M-P., M.M.-L. and J.M.P.-S.; investigation, E.G.-G., I.G.-H., A.B., C.T.-P. and J.D.G-L.; resources, A.B. and D.G.-L.; writing original draft preparation, E.G.-G., J.M.P.-S., J. D.G.-L. and A.B.; writing review and editing, E.G.-G., E.V., J.D.G.-L., J.M.P-S., A.M.M-P., M.M.-B. and A.B.; supervision, A.M.M-P, E.V., A.B. and M.M.-B. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
González-Gragera, E., García-López, J.D., Teso-Pérez, C. et al. Genomic Characterization of Piscicolin CM22 Produced by Carnobacterium maltaromaticum CM22 Strain Isolated from Salmon (Salmo salar). Probiotics & Antimicro. Prot. (2024). https://doi.org/10.1007/s12602-024-10316-1
Accepted:
Published:
DOI: https://doi.org/10.1007/s12602-024-10316-1