Abstract
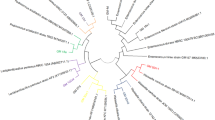
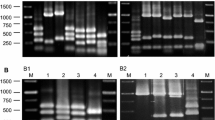
Milk kefir fermentation has been used in households for generations. Consumption of milk kefir has been associated with various health benefits, presumably from the probiotics of yeast and bacteria that make up the kefir grains. In addition, many of the microbes are known to produce novel antimicrobial compounds that can be used for other applications. The microbes living inside kefir grains differ significantly depending on geographical location and production methods. In this study, we aimed to use metagenomic analysis of fermented milk by using three different kefir grains (kefir 1, kefir 2, and kefir 3) from different US sources. We analyzed the microbial compositions of the three milk fermentation samples. This study revealed that each sample contains unique and distinct groups of microbes, kefir 1 showed the least diversity, and kefir 3 showed the highest diversity. Kefir 3 is rich in Proteobacteria while kefir 2 is dominated by the Firmicutes. Using bacterial indicator growth analyses carried out by continuous readings from microplate-based bioreactor assays suggested that kefir 2 fermentation filtrate has higher antibacterial property. We have screened 30 purified cultures of kefir 2 sample and isolated two lactic acid bacteria strains with higher antibacterial activities; the two strains were identified as Leuconostoc mesenteroides 28–1 and Lentilactobacillus kefiri 25–2 by 16S genomic PCR with confirmed antibacterial activities of fermentation filtrate after growing under both aerobic and anaerobic conditions.








Similar content being viewed by others
Data Availability
All data generated and analyzed during this study are included in this published article.
References
Rosa DD et al (2017) Milk kefir: nutritional, microbiological and health benefits. Nutr Res Rev 30(1):82–96
Nielsen B, Gürakan GC, Ünlü G (2014) Kefir: a multifaceted fermented dairy product. Probiotics and antimicrobial proteins 6(3):123–135
Bengoa AA et al (2019) Kefir micro-organisms: their role in grain assembly and health properties of fermented milk. J Appl Microbiol 126(3):686–700
Marshall VM, Cole WM, Brooker B (1984) Observations on the structure of kefir grains and the distribution of the microflora. J Appl Bacteriol 57(3):491–497
Garrote GL, Abraham AG, De Antoni GL (2001) Chemical and microbiological characterisation of kefir grains. J Dairy Res 68:639–652
Garofalo C et al (2015) Bacteria and yeast microbiota in milk kefir grains from different Italian regions. Food Microbiol 49:123–133
Chen HC, Wang SY, Chen MJ (2008) Microbiological study of lactic acid bacteria in kefir grains by culture-dependent and culture-independent methods. Food Microbiol 25(3):492–501
Azizi NF et al (2021) Kefir and its biological activities. Foods 10
Cevikbas A et al (1994) Antitumoural antibacterial and antifungal activities of kefir and kefir grain. Phytother Res 8(2):78–82
Guzel-Seydim ZB et al (2011) Review: functional properties of kefir. Crit Rev Food Sci Nutr 51(3):261–268
Bourrie BCT, Willing BP, Cotter PD (2016) The microbiota and health promoting characteristics of the fermented beverage kefir. Front Microbiol 7
Marshall VM, Cole WM (1985) Methods for making kefir and fermented milks based on kefir. J Dairy Res 52(3):451–456
Figler M et al (2006) Effect of special Hungarian probiotic kefir on faecal microflora. World J Gastroenterol: WJG 12(7):1129
Chen Y et al (2013) Effects of Lactobacillus kefiranofaciens M1 isolated from kefir grains on enterohemorrhagic Escherichia coli infection using mouse and intestinal cell models. J Dairy Sci 96(12):7467–7477
Santos A et al (2003) The antimicrobial properties of different strains of Lactobacillus spp. isolated from kefir. Syst Appl Microbiol 26(3):434–437
Leite AM et al (2012) Assessment of the microbial diversity of Brazilian kefir grains by PCR-DGGE and pyrosequencing analysis. Food Microbiol 31(2):215–221
Fu L et al (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28(23):3150–3152
Ondov BD, Bergman NH, Phillippy AM (2011) Interactive metagenomic visualization in a web browser. BMC Bioinformatics 12:385
Sunagawa S et al (2015) Ocean plankton. Structure and function of the global ocean microbiome. Science 348(6237):1261359
Mende DR et al (2012) Assessment of metagenomic assembly using simulated next generation sequencing data. PloS One 7(2):e31386
Liu S et al (2019) Increased ethanol tolerance associated with the pntAB locus of Oenococcus oeni and Lactobacillus buchneri. J Ind Microbiol Biotechnol 46(11):1547–1556
Nielsen HB et al (2014) Identification and assembly of genomes and genetic elements in complex metagenomic samples without using reference genomes. Nat Biotechnol 32(8):822–828
Zhu W, Lomsadze A, Borodovsky M (2010) Ab initio gene identification in metagenomic sequences. Nucleic Acids Res 38(12):e132
Li J et al (2014) An integrated catalog of reference genes in the human gut microbiome. Nat Biotechnol 32(8):834–841
Buchfink B, Xie C, Huson DH (2015) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12(1):59–60
Huson DH et al (2011) Integrative analysis of environmental sequences using MEGAN4. Genome Res 21(9):1552–1560
Huson DH et al (2007) MEGAN analysis of metagenomic data. Genome Res 17(3):377–386
Minot SS, Willis AD (2019) Clustering co-abundant genes identifies components of the gut microbiome that are reproducibly associated with colorectal cancer and inflammatory bowel disease. Microbiome 7(1):110
Kanehisa M et al (2006) From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res 34(Database issue): p. D354–7
Kanehisa M et al (2014) Data, information, knowledge and principle: back to metabolism in KEGG. Nucleic Acids Res 42(Database issue): p. D199–205
Cantarel BL et al (2009) The Carbohydrate-Active EnZymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res 37(Database issue): p. D233–8
Garrote GL, Abraham AG, De Antoni GL (2000) Inhibitory power of kefir: the role of organic acids. J Food Prot 63(3):364–369
García JA et al (2000) Pulsed-field gel electrophoresis analysis of Aeromonas salmonicida ssp. salmonicida. FEMS Microbiol Lett 190(1): p. 163–166
Tong Z et al (2012) An in vitro investigation of Lactococcus lactis antagonizing cariogenic bacterium Streptococcus mutans. Arch Oral Biol 57(4):376–382
Reddy S et al (2021) Comparative evaluation of efficacy of kefir milk probiotic curd and probiotic drink on Streptococcus mutans in 8–12-year-old children: an in vivo study. Int J Clin Pediatr Dent 14(1):120–127
Cooney S et al (2014) Bacteria: other pathogenic Enterobacteriaceae – Enterobacter and other genera. In: Motarjemi Y (ed) Encyclopedia of food safety. Academic Press, Waltham, pp 433–441
Zeng X et al (2022) Metagenomic analysis of microflora structure and functional capacity in probiotic Tibetan kefir grains. Food Res Int 151:110849
Siedler S et al (2020) Competitive exclusion is a major bioprotective mechanism of lactobacilli against fungal spoilage in fermented milk products. Appl Environ Microbiol 86(7)
Ioannou A, Knol J, Belzer C (2021) Microbial glycoside hydrolases in the first year of life: an analysis review on their presence and importance in infant gut. Front Microbiol 12:631282–631282
Biely P (2012) Microbial carbohydrate esterases deacetylating plant polysaccharides. Biotechnol Adv 30(6):1575–1588
Shoseyov O, Shani Z, Levy I (2006) Carbohydrate binding modules: biochemical properties and novel applications. Microbiol Mol Biol Rev 70(2):283–295
Chakraborty S et al (2017) 23 - Polysaccharide lyases, in Current developments in biotechnology and bioengineering, A. Pandey, S. Negi, and C.R. Soccol, Editors Elsevier. p. 527–539
Acknowledgements
We thank Eric Hoecker and Amber Anderson for their excellent technical assistance.
Funding
USDA is an equal opportunity provider and employer. This work was supported in part by the US Department of Agriculture, Agricultural Research Service. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics Approval
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, S., Lu, SY., Qureshi, N. et al. Antibacterial Property and Metagenomic Analysis of Milk Kefir. Probiotics & Antimicro. Prot. 14, 1170–1183 (2022). https://doi.org/10.1007/s12602-022-09976-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-022-09976-8