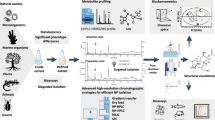
Abstract
Stable isotopes are increasingly used to trace microbial activities and metabolic pathways and are widely applied in microbiology, biochemistry, and risk assessment. This development is tightly linked to advance of mass spectrometric instrumentation as the key analysis for stable isotopes. Also the concepts for stable isotope tracing are progressing fast. We briefly review such concepts and indicate some of our own studies.
Similar content being viewed by others
Literatur
Wang S, Seiwert B, Kästner M et al. (2016) (Bio)degradation of glyphosate in water-sediment microcosms – a stable isotope co-labeling approach. Water Res 99:91–100
Adrian L, Marco-Urrea E (2016) Isotopes in geobiochemistry: tracing metabolic pathways in microorganisms of environmental relevance with stable isotopes. Curr Opin Biotechnol 41:19–25
North KP, Mackay DM, Annable MD et al. (2012) An ex situ evaluation of TBA-and MTBE-baited bio-traps. Water Res 46:3879–3888
Nowak KM, Miltner A, Gehre M et al. (2011) Formation and fate of bound residues from microbial biomass during 2,4-D degradation in soil. Environ Sci Technol 45:999–1006
Marco-Urrea E, Seifert J, von Bergen M et al. (2012) Stable isotope peptide mass spectrometry to decipher amino acid metabolism in Dehalococcoides strain CBDB1. J Bacteriol 194:4169–4177
Zhuang W-Q, Yi S, Feng X et al. (2011) Selective utilization of exogenous amino acids by Dehalococcoides ethenogenes strain 195 and the effects on growth and dechlorination activity. Appl Environ Microbiol 77: 7797–7803
Zamboni N, Saghatelian A, Patti Gary J (2015) Defining the metabolome: size, flux, and regulation. Mol Cell 58:699–706
Jang C, Chen L, Rabinowitz JD (2018) Metabolomics and isotope tracing. Cell 173:822–837
Tang YJ, Yi S, Zhuang W-Q et al. (2009) Investigation of carbon metabolism in „Dehalococcoides ethenogenes“ strain 195 by use of isotopomer and transcriptomic analyses. J Bacteriol 191:5224–5231
Jehmlich N, Schmidt F, von Bergen M et al. (2008) Protein-based stable isotope probing (Protein-SIP) reveals active species within anoxic mixed cultures. ISME J 2:1122–1133
Author information
Authors and Affiliations
Corresponding author
Additional information
Karolina Małgorzata Nowak Jahrgang 1980. Studium der Umweltwissenschaften an der Universität Warmia und Masuren, Olsztyn, Polen. 2011 Promotion in der Umweltchemie an der RWTH Aachen. 2011–2012 Postdoc am Helmholtz-Zentrum für Umweltforschung (UFZ), Leipzig. 2012–2017 Postdoc an der RWTH Aachen. Seit 2018 wissenschaftliche Mitarbeiterin in einem eigenem DFG-Projekt am Fachgebiet Geobiotechnologie, Institut für Biotechnologie der TU Berlin.
Ernest Marco-Urrea Jahrgang 1977. 1998 Studium Technical Industrial Engineering an der Universität Politècnica de Catalunya, Barcelona, Spanien und 2001 der Umweltwissenschaften an der Universität Autònoma de Barcelona (UAB). 2007 Promotion an der UAB. 2009 Postdoc als Marie-Curie-Fellow am Helmholtz-Zentrum für Umweltforschung (UFZ), Leipzig. Seit 2011 Associate Professor an der Fakultät für Chemische, Biologische und Umwelttechnik, UAB, Barcelona, Spanien.
Lorenz Adrian Jahrgang 1966. Biologiestudium an der Universität Bonn; Biochemiestudium an der Universität Manchester, UK. 1999 Promotion an der TU Berlin. 2003 und 2007 Postdoc an der Cornell University und am Rensselaer Polytechnic Institute, Troy, NY, USA. 2006 Habilitation an der TU Berlin. 2008–2013 ERC-Gruppe und seit 2013 Arbeitsgruppen - leiter am Helmholtz-Zentrum für Umweltforschung (UFZ), Leipzig. Seit 2017 Professur für Geobiotechnologie am Institut für Biotechnologie der TU Berlin.
Rights and permissions
About this article
Cite this article
Nowak, K.M., Marco-Urrea, E. & Adrian, L. Markierung im mikrobiellen Stoffwechsel. Biospektrum 24, 491–494 (2018). https://doi.org/10.1007/s12268-018-0951-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12268-018-0951-4