Abstract
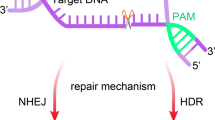
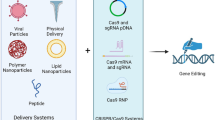
The clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated protein 9 (Cas9) system has been used for the insertion of large transgenes into Chinese hamster ovary cells via co-transfection of a Cas9/guide RNA expression vector and donor plasmid. The Cas9 protein includes nuclear localization sequences that are used as peptide tags for the import of Cas9 into the nucleus. However, the import of a donor plasmid into the nucleus is passive because of the absence of such localization signals; thus, the delivery of Cas9 and the donor plasmid is not synchronized, resulting in low knock-in (KI) efficiency. Here, we modified the Cas9 expression vector expressing a Cas9 protein fused to a zinc finger (ZF) domain, Cas9-ZF, to expedite the translocation of the donor plasmid into the nucleus and the co-localization of the donor plasmid with a CRISPR/Cas9-mediated DNA double-strand break site by tethering Cas9-ZF and the donor plasmid. Compared to the typical donor plasmid and wild-type Cas9, the donor plasmid harboring the ZF-binding motif showed increased homology-mediated KI efficiency, while the engineered Cas9 protein showed decreased expression and gene-editing efficiency. Moreover, the pair of Cas9-ZF and the donor plasmid with the ZF motif did not improve KI efficiency, but rather negated the positive effect of the donor plasmid with the ZF motif. This study demonstrates the importance of the transport of donor plasmids and the limitations of using the plasmid-based Cas9-ZF fusion system to improve KI efficiency.
Similar content being viewed by others
References
Lee, J. S., H. F. Kildegaard, N. E. Lewis, and G. M. Lee (2019) Mitigating clonal variation in recombinant mammalian cell lines. Trends Biotechnol. 37: 931–942.
Bandyopadhyay, A. A., S. A. O’Brien, L. Zhao, H. Y. Fu, N. Vishwanathan, and W. S. Hu (2019) Recurring genomic structural variation leads to clonal instability and loss of productivity. Biotechnol. Bioeng. 116: 41–53.
Lee, J. S., T. B. Kallehauge, L. E. Pedersen, and H. F. Kildegaard (2015) Site-specific integration in CHO cells mediated by CRISPR/Cas9 and homology-directed DNA repair pathway. Sci. Rep. 5: 8572.
Shin, S. W. and J. S. Lee (2020) CHO cell line development and engineering via site-specific integration: challenges and opportunities. Biotechnol. Bioprocess Eng. 25: 633–645.
Carlson-Stevermer, J., A. A. Abdeen, L. Kohlenberg, M. Goedland, K. Molugu, M. Lou, and K. Saha (2017) Assembly of CRISPR ribonucleoproteins with biotinylated oligonucleotides via an RNA aptamer for precise gene editing. Nat. Commun. 8: 1711.
Roche, P. J. R., H. Gytz, F. Hussain, C. J. F. Cameron, D. Paquette, M. Blanchette, J. Dostie, B. Nagar, and U. D. Akavia (2018) Double-stranded biotinylated donor enhances homology-directed repair in combination with Cas9 monoavidin in mammalian cells. CRISPR J. 1: 414–430.
Gu, B., E. Posfai, and J. Rossant (2018) Efficient generation of targeted large insertions by microinjection into two-cell-stage mouse embryos. Nat. Biotechnol. 36: 632–637.
Aird, E. J., K. N. Lovendahl, A. St Martin, R. S. Harris, and W. R. Gordon (2018) Increasing Cas9-mediated homology-directed repair efficiency through covalent tethering of DNA repair template. Commun. Biol. 1: 54.
Savic, N., F. C. Ringnalda, H. Lindsay, C. Berk, K. Bargsten, Y. Li, D. Neri, M. D. Robinson, C. Ciaudo, J. Hall, M. Jinek, and G. Schwank (2018) Covalent linkage of the DNA repair template to the CRISPR-Cas9 nuclease enhances homology-directed repair. Elife 7: e33761.
Ling, X., B. Xie, X. Gao, L. Chang, W. Zheng, H. Chen, Y. Huang, L. Tan, M. Li, and T. Liu (2020) Improving the efficiency of precise genome editing with site-specific Cas9-oligonucleotide conjugates. Sci. Adv. 6: eaaz0051.
Strecker, J., A. Ladha, Z. Gardner, J. L. Schmid-Burgk, K. S. Makarova, E. V. Koonin, and F. Zhang (2019) RNA-guided DNA insertion with CRISPR-associated transposases. Science 365: 48–53.
Klompe, S. E., P. L. H. Vo, T. S. Halpin-Healy, and S. H. Sternberg (2019) Transposon-encoded CRISPR-Cas systems direct RNA-guided DNA integration. Nature 571: 219–225.
Anzalone, A. V., L. W. Koblan, and D. R. Liu (2020) Genome editing with CRISPR-Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 38: 824–844.
Wu, S. C., Y. J. Meir, C. J. Coates, A. M. Handler, P. Pelczar, S. Moisyadi, and J. M. Kaminski (2006) piggyBac is a flexible and highly active transposon as compared to sleeping beauty, Tol2, and Mos1 in mammalian cells. Proc. Natl. Acad. Sci. U. S. A. 103: 15008–15013.
Hew, B. E., R. Sato, D. Mauro, I. Stoytchev, and J. B. Owens (2019) RNA-guided piggyBac transposition in human cells. Synth. Biol. (Oxf.) 4: ysz018.
Luo, W., D. L. Galvan, L. E. Woodard, D. Dorset, S. Levy, and M. H. Wilson (2017) Comparative analysis of chimeric ZFP-, TALE- and Cas9-piggyBac transposases for integration into a single locus in human cells. Nucleic Acids Res. 45: 8411–8422.
Kovač, A., C. Miskey, M. Menzel, E. Grueso, A. Gogol-Döring, and Z. Ivics (2020) RNA-guided retargeting of Sleeping Beauty transposition in human cells. Elife 9: e53868.
Shin, S. W., D. Kim, and J. S. Lee (2021) Controlling ratios of plasmid-based double cut donor and CRISPR/Cas9 components to enhance targeted integration of transgenes in Chinese hamster ovary cells. Int. J. Mol. Sci. 22: 2407.
Dean, D. A., D. D. Strong, and W. E. Zimmer (2005) Nuclear entry of nonviral vectors. Gene Ther. 12: 881–890.
Shin, S. W. and J. S. Lee (2020) Optimized CRISPR/Cas9 strategy for homology-directed multiple targeted integration of transgenes in CHO cells. Biotechnol. Bioeng. 117: 1895–1903.
Zelphati, O., X. Liang, P. Hobart, and P. L. Felgner (1999) Gene chemistry: functionally and conformationally intact fluorescent plasmid DNA. Hum. Gene Ther. 10: 15–24.
Ludtke, J. J., M. G. Sebestyén, and J. A. Wolff (2002) The effect of cell division on the cellular dynamics of microinjected DNA and dextran. Mol. Ther. 5: 579–588. (Erratum published 2002, Mol. Ther. 6: 134)
Chu, V. T., T. Weber, B. Wefers, W. Wurst, S. Sander, K. Rajewsky, and R. Kühn (2015) Increasing the efficiency of homology-directed repair for CRISPR-Cas9-induced precise gene editing in mammalian cells. Nat. Biotechnol. 33: 543–548. (Erratum published 2018, Nat. Biotechnol. 36: 196)
Lam, A. P. and D. A. Dean (2010) Progress and prospects: nuclear import of nonviral vectors. Gene Ther. 17: 439–447.
Xiong, T., G. E. Meister, R. E. Workman, N. C. Kato, M. J. Spellberg, F. Turker, W. Timp, M. Ostermeier, and C. D. Novina (2017) Targeted DNA methylation in human cells using engineered dCas9-methyltransferases. Sci. Rep. 7: 6732.
Bolukbasi, M. F., A. Gupta, S. Oikemus, A. G. Derr, M. Garber, M. H. Brodsky, L. J. Zhu, and S. A. Wolfe (2015) DNA-binding-domain fusions enhance the targeting range and precision of Cas9. Nat. Methods 12: 1150–1156.
Drummond, I. A., P. Rohwer-Nutter, and V. P. Sukhatme (1994) The zebrafish egr1 gene encodes a highly conserved, zinc-finger transcriptional regulator. DNA Cell Biol. 13: 1047–1055.
Sera, T. and C. Uranga (2002) Rational design of artificial zinc-finger proteins using a nondegenerate recognition code table. Biochemistry 41: 7074–7081.
Kim, J. S. and C. O. Pabo (1998) Getting a handhold on DNA: design of poly-zinc finger proteins with femtomolar dissociation constants. Proc. Natl. Acad. Sci. U. S. A. 95: 2812–2817.
Gersbach, C. A., T. Gaj, and C. F. Barbas3rd (2014) Synthetic zinc finger proteins: the advent of targeted gene regulation and genome modification technologies. Acc. Chem. Res. 47: 2309–2318.
Mesika, A., I. Grigoreva, M. Zohar, and Z. Reich (2001) A regulated, NFkappaB-assisted import of plasmid DNA into mammalian cell nuclei. Mol. Ther. 3: 653–657.
Dean, D. A., B. S. Dean, S. Muller, and L. C. Smith (1999) Sequence requirements for plasmid nuclear import. Exp. Cell Res. 253: 713–722.
Shin, S., S. H. Kim, J. S. Lee, and G. M. Lee (2021) Streamlined human cell-based recombinase-mediated cassette exchange platform enables multigene expression for the production of therapeutic proteins. ACS Synth. Biol. 10: 1715–1727.
Standage-Beier, K., N. Brookhouser, P. Balachandran, Q. Zhang, D. A. Brafman, and X. Wang (2019) RNA-guided recombinase-Cas9 fusion targets genomic DNA deletion and integration. CRISPR J. 2: 209–222.
Maeder, M. L., S. Thibodeau-Beganny, J. D. Sander, D. F. Voytas, and J. K. Joung (2009) Oligomerized pool engineering (OPEN): an ‘open-source’ protocol for making customized zinc-finger arrays. Nat. Protoc. 4: 1471–1501.
Konermann, S., M. D. Brigham, A. E. Trevino, J. Joung, O. O. Abudayyeh, C. Barcena, P. D. Hsu, N. Habib, J. S. Gootenberg, H. Nishimasu, O. Nureki, and F. Zhang (2015) Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 517: 583–588.
Acknowledgements
This work was supported by the NRF funded by the Korean government (2021R1A2C4002733 and 2019R1A6A1A11 051471).
Author information
Authors and Affiliations
Contributions
Dongwoo Kim: Conceptualization, Investigation, Methodology, Writing - original draft, Writing - review & editing. Jae Seong Lee: Conceptualization, Funding acquisition, Methodology, Supervision, Writing - original draft, Writing - review & editing.
Corresponding author
Ethics declarations
The authors declare no financial or commercial conflict of interest.
Neither ethical approval nor informed consent was required for this study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Kim, D., Lee, J.S. Limitations of the Plasmid-Based Cas9-Zinc Finger Fusion System for Homology-Directed Knock-In in Chinese Hamster Ovary Cells. Biotechnol Bioproc E 28, 289–299 (2023). https://doi.org/10.1007/s12257-022-0348-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-022-0348-6