Abstract
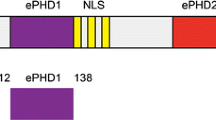
Krüppel-like factor 4 (KLF4) is a transcription factor involved in diverse biological processes, including development, cellular differentiation and proliferation, and maintenance of tissue homeostasis. KLF4 has also been associated with disease states, such as cardiovascular disease and several cancers. KLF4 contains an activation domain, repression domain, and a structurally characterized C-terminal zinc finger domain that mediates its binding to DNA. The structurally uncharacterized KLF4 activation domain is critical for transactivation by KLF4 and mediates its binding to the transcriptional coactivator CBP/p300. Here, we report the 1H, 15N, 13CO, 13Cα and 13Cβ NMR chemical shift assignments of KLF41–130, which contains the KLF4 activation domain. Narrow chemical shift dispersion in the 1H dimension of the 1H-15N HSQC spectrum suggests that the KLF41–130 fragment is intrinsically disordered.


Similar content being viewed by others
References
Bieker JJ (2001) Kruppel-like Factors: Three Fingers in Many Pies. J Biol Chem 276:34355–34358. doi:10.1074/jbc.R100043200
Clubb RT, Thanabal V, Wagner G (1992) A constant-time three-dimensional Triple-resonance pulse scheme to correlate intraresidue 1HN, 15 N, and 13C’ chemical shifts in 15 N-13C-labeled proteins. J Magn Reson 97:213–217
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Evans PM, Liu C, 2008. Roles of Krüpel-like factor 4 in normal homeostasis, cancer and stem cells. Acta Biochim. Biophys. Sin (Shanghai) 40:554–564
Feinberg MW, Cao Z, Wara AK, Lebedeva MA, Senbanerjee S, Jain MK (2005) Kruppel-like factor 4 is a mediator of proinflammatory signaling in macrophages. J Biol Chem 280:38247–38258. doi:10.1074/jbc.M509378200
Foster KW, Frost AR, McKie-Bell P, Lin CY, Engler JA, Grizzle WE, Ruppert JM (2000) Increase of GKLF messenger RNA and protein expression during progression of breast cancer. Cancer Res 60:6488–6495
Garrett-Sinha LA, Eberspaecher H, Seldin MF, de Crombrugghe B (1996) A gene for a novel Zinc-finger protein expressed in differentiated epithelial cells and transiently in certain mesenchymal cells. J Biol Chem 271:31384–31390. doi:10.1074/jbc.271.49.31384
Geiman DE, Ton-That H, Johnson JM, Yang VW (2000) Transactivation and growth suppression by the gut-enriched Krüppel-like factor (Krüppel-like factor 4) are dependent on acidic amino acid residues and protein–protein interaction. Nucleic Acids Res 28:1106–1113. doi:10.1093/nar/28.5.1106
Grzesiek S, Bax A (1992a) Improved 3D triple-resonance NMR techniques applied to a 31 kDa protein. J Magn Reson. doi:10.1016/0022-2364(92)90099-S
Grzesiek S, Bax A (1992b) An efficient experiment for sequential backbone assignment of medium-sized isotopically enriched proteins. J Magn Reson 99(1):201–207. doi:10.1016/0022-2364(92)90169-8
Grzesiek S, Bax A (1992c) Correlating backbone amide and side chain resonances in larger proteins by multiple relayed triple resonance NMR. J Am Chem Soc 114:6291–6293. doi:10.1021/ja00042a003
Grzesiek S, Anglister J, Bax A (1993) Correlation of backbone amide and aliphatic side-chain resonances in 13C/15N-enriched proteins by isotropic mixing of 13C magnetization. J Magn Reson 101:114–119
Kay LE, Ikura M, Tschudin R, Bax A (1990) Three-dimensional triple-resonance NMR Spectroscopy of isotopically enriched proteins. J Magn Reson 89:496–514. doi:10.1016/j.jmr.2011.09.004
Liu Y, Olanrewaju YO, Zheng Y, Hashimoto H, Blumenthal RM, Zhang X, Cheng X (2014) Structural basis for Klf4 recognition of methylated DNA. Nucleic Acids Res 42:4859–4867. doi:10.1093/nar/gku134
Maherali N, Sridharan R, Xie W, Utikal J, Eminli S, Arnold K, Stadtfeld M, Yachechko R, Tchieu J, Jaenisch R, Plath K, Hochedlinger K (2007) Directly reprogrammed fibroblasts show global epigenetic remodeling and widespread tissue contribution. Cell Stem Cell 1:55–70. doi:10.1016/j.stem.2007.05.014
Marsh J, Singh V, Jia Z, Forman-Kay J (2006) Sensitivity of secondary structure propensities to sequence differences between α- and γ-synuclein: implications for fibrillation. Protein Sci 15:2795–2804. doi:10.1110/ps.062465306
McConnell BB, Yang VW (2010) Mammalian kruppel-like factors in health and diseases. Physiol Rev 90:1337–1381. doi:10.1152/physrev.00058.2009
Muhandiram DR, Kay LE (1994) Gradient-enhanced triple-resonance three-dimensional NMR experiments with improved sensitivity. J Magn Reson Ser B 103:203–216. doi:10.1006/jmrb.1994.1032
Schuetz A, Nana D, Rose C, Zocher G, Milanovic M, Koenigsmann J, Blasig R, Heinemann U, Carstanjen D (2011) The structure of the Klf4 DNA-binding domain links to self-renewal and macrophage differentiation. Cell Mol Life Sci 68:3121–3131. doi:10.1007/s00018-010-0618-x
Segre JA, Bauer C, Fuchs E (1999) Klf4 is a transcription factor required for establishing the barrier function of the skin. Nat Genet 22:356–360. doi:10.1038/11926
Shields JM, Christy RJ, Yang VW (1996) Identification and characterization of a gene encoding a gut-enriched Krüppel-like factor expressed during growth arrest. J Biol Chem 271:20009–20017
Suske G, Bruford E, Philipsen S (2005) Mammalian SP/KLF transcription factors: bring in the family. Genomics 85:551–556. doi:10.1016/j.ygeno.2005.01.005
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, Yamanaka S (2007) Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131:861–872. doi:10.1016/j.cell.2007.11.019
Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas M, Ulrich EL, Markley JL, Ionides J, Laue ED (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins Struct Funct Bioinform 59:687–696. doi:10.1002/prot.20449
Wernig M, Meissner A, Foreman R, Brambrink T, Ku M, Hochedlinger K, Bernstein BE, Jaenisch R (2007) In vitro reprogramming of fibroblasts into a pluripotent ES-cell-like state. Nature 448:318–324. doi:10.1038/nature05944
Yet SF, McA’Nulty MM, Folta SC, Yen HW, Yoshizumi M, Hsieh CM, Layne MD, Chin MT, Wang H, Perrella MA, Jain MK, Lee ME (1998) Human EZF, a Krüppel-like zinc finger protein, is expressed in vascular endothelial cells and contains transcriptional activation and repression domains. J Biol Chem 273:1026–1031
Yoshida T, Gan Q, Owens GK (2008) Kruppel-like factor 4, Elk-1, and histone deacetylases cooperatively suppress smooth muscle cell differentiation markers in response to oxidized phospholipids. Am J Physiol Cell Physiol 295:C1175–C1182. doi:10.1152/ajpcell.00288.2008
Yoshida T, Yamashita M, Hayashi M (2014) Protective effects of Krüppel- like factor 4 against cardiovascular disease. Ann Vasc Med Res 1:1002
Yu T, Chen X, Zhang W, Li J, Xu R, Wang TC, Ai W, Liu C, Hotchin NA (2012) Krü ppel-like factor 4 regulates intestinal epithelial cell morphology and polarity. PLoS ONE 7. doi:10.1371/journal.pone.0032492
Zhang W, Geiman DE, Shields JM, Dang DT, Mahatan CS, Kaestner KH, Biggs JR, Kraft AS, Yang VW (2000) The gut-enriched Kruppel-like factor (Kruppel-like factor 4) mediates the transactivating effect of p53 on the p21WAF1/Cip1 promoter. J Biol Chem 275:18391–18398. doi:10.1074/jbc.C000062200
Zhao W, Hisamuddin IM, Nandan MO, Babbin BA, Lamb NE, Yang VW (2004) Identification of Krüppel-like factor 4 as a potential tumor suppressor gene in colorectal cancer. Oncogene 23:395–402. doi:10.1038/sj.onc.1207067
Acknowledgements
This work was funded by a Natural Science and Engineering Council of Canada discovery grant to SPS. DNL is a recipient of a CIHR Postdoctoral Fellowship and BSC was supported by a Queen’s University Chancellor’s Scholarship.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animal research information
This work does not involve any studies involving human or animal subjects.
Rights and permissions
About this article
Cite this article
Conroy, B.S., Weiss, E.R., Smith, S.P. et al. Backbone 1H, 13C, and 15N NMR resonance assignments of the Krüppel-like factor 4 activation domain. Biomol NMR Assign 11, 95–98 (2017). https://doi.org/10.1007/s12104-017-9727-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-017-9727-x