Abstract
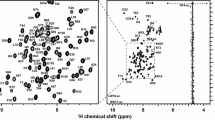
Plant homeodomains (PHD) and Bromo domains are both chromatin reader domains that recognise histone methylation degree and acetylation state, respectively. The tripartite motif protein TRIM24 is a multidomain protein carrying a PHD-Bromo motif at its C-terminus, through which it is able to bind to histone 3 (H3) N-terminal tails with a specific modification pattern, namely unmethylated at K4 and acetylated at K23 (H3-K4me0K23ac). Here we report the \({}^{1}\hbox {H}, {}^{13}\hbox {C}\) and \({}^{15}\hbox {N}\) backbone resonance assignment of this 23 kDa motif, which we have obtained by heteronuclear multidimensional NMR spectroscopy. Furthermore we show that the secondary \(\hbox {C}_\alpha\) and \(\hbox {C}_\beta\) chemical shifts are in good agreement with a previously published crystal structure.




Similar content being viewed by others
References
Allton K, Jain AK, Herz HM, Tsai WW, Jung SY, Qin J, Bergmann A, Johnson RL, Barton MC (2009) TRIM24 targets endogenous p53 for degradation. Proc Natl Acad Sci USA 106(28):11,612–11,616
Chen C, Seth AK, Aplin AE (2006) Genetic and expression aberrations of E3 ubiquitin ligases in human breast cancer. Mol Cancer Res 4(10):695–707
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6(3):277–293
Grzesiek S, Bax A (1992a) An efficient experiment for sequential backbone assignment of medium-sized isotopically enriched proteins. J Magn Reson 99:201–207
Grzesiek S, Bax A (1992b) Correlating backbone amide and side chain resonances in larger proteins by multiple relayed triple resonance NMR. J Am Chem Soc 114:6291–6293
Kay LE, Ikura M, Tschudin R, Bax A (1990) Three-dimensional triple-resonance NMR spectroscopy of isotopically enriched proteins. J Magn Reson 89:496–514
Keller RLJ (2004) The computer aided resonance tutorial. Cantina, Goldau
Le Douarin B, Zechel C, Garnier JM, Lutz Y, Tora L, Pierrat P, Heery D, Gronemeyer H, Chambon P, Losson R (1995) The N-terminal part of TIF1, a putative mediator of the ligand-dependent activation function (AF-2) of nuclear receptors, is fused to B-raf in the oncogenic protein T18. EMBO J 14(9):2020–2033
Le Douarin B, Nielsen AL, Garnier JM, Ichinose H, Jeanmougin F, Losson R, Chambon P (1996) A possible involvement of TIF1 alpha and TIF1 beta in the epigenetic control of transcription by nuclear receptors. EMBO J 15(23):6701–6715
Live DH, Davis DG, Agosta WC, Cowburn D (1984) Observation of 1000-fold enhancement of nitrogen-15 NMR via proton-detected multiquantum coherences: studies of large peptides. J Am Chem Soc 106(20):6104–6105
Reddy BA, Etkin LD, Freemont PS (1992) A novel zinc finger coiled-coil domain in a family of nuclear proteins. Trends Biochem Sci 17(9):344–345
Reymond A, Meroni G, Fantozzi A, Merla G, Cairo S, Luzi L, Riganelli D, Zanaria E, Messali S, Cainarca S, Guffanti A, Minucci S, Pelicci PG, Ballabio A (2001) The tripartite motif family identifies cell compartments. EMBO J 20(9):2140–2151
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44(4):213–223
Tsai WW, Wang Z, Yiu TT, Akdemir KC, Xia W, Winter S, Tsai CY, Shi X, Schwarzer D, Plunkett W, Aronow B, Gozani O, Fischle W, Hung MC, Patel DJ, Barton MC (2010) TRIM24 links a non-canonical histone signature to breast cancer. Nature 468(7326):927–932
Wishart DS, Sykes BD (1994) The 13C chemical-shift index: a simple method for the identification of protein secondary structure using 13C chemical-shift data. J Biomol NMR 4(2):171–180
Acknowledgments
We would like to thank Liz Flavell for establishing the protein expression and purification protocols.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Walser, R., Renshaw, J. & Milbradt, A.G. Backbone resonance assignments for the PHD-Bromo dual-domain of the human chromatin reader TRIM24. Biomol NMR Assign 10, 207–211 (2016). https://doi.org/10.1007/s12104-016-9668-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-016-9668-9