Abstract
The plant homeodomain (PHD) zinc finger is a structural motif of about 40–60 amino acid residues found in many eukaryotic proteins that are involved in chromatin-mediated gene regulation. The human chromodomain helicase DNA binding protein 4 (CHD4) is a multi-domain protein that harbours, at its N-terminal end, a pair of PHD finger motifs (dPHD) connected by a ~30 amino acid linker. This tandem PHD motif is thought to be involved in targeting CHD4 to chromatin via its interaction with histone tails. Here we report the 1H, 13C and 15N backbone and side-chain resonance assignment of the entire dPHD by heteronuclear multidimensional NMR spectroscopy. These assignments provide the starting point for the determination of the structure, dynamics and histone-binding properties of this tandem domain pair.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Biological context
Plant homeodomain (PHD) fingers are short (~40–60 amino acid) protein domains which contain two zinc ions bound by a C4HC3 ligand motif with cross-brace ligation topology. They were discovered over 20 years ago (Schindler et al. 1993) and are found in more than 400 eukaryotic proteins, many of which are involved in the recognition of histone tails and in the chromatin remodelling mechanism (Mellor 2006). In recent years, a number of PHD motifs have been shown to act as specialized histone code “reader” modules that often recognize the methylation status of histone H3 tails (Pena et al. 2006; Shi et al. 2006; Wysocka et al. 2006; Zhang 2006). The biological importance of PHD fingers in human cells is highlighted by their occurrence in many chromatin-remodelling proteins linked to a variety of human diseases including cancer, mental retardation, and immunodeficiency (Baker et al. 2008).
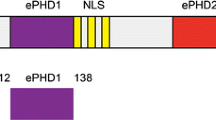
Chromodomain helicase DNA binding protein 4 (CHD4), also known as Mi2b, belongs to the SNF2 family of helicases (Eisen et al. 1995) and was first identified as a dermatomyositis-specific auto antigen (Seelig et al. 1995). CHD4 is the ATPase component of the nucleosome remodelling and deacetylases (NuRD) complex, which is involved in many repressive transcriptional regulatory processes (Ramirez and Hagman 2009). In addition to its SNF2-type ATPase motor, CHD4 harbours double PHD fingers (dPHD) and double chromodomains (dCHD) (Fig. 1) that are believed to be involved in its targeting to the chromatin (Morra et al. 2012), yet their mechanism of action is unclear. While the combination of PHD domains with other chromatin “reader” modules such as bromodomains is a feature of many chromatin-remodelling factors, the presence of tandem PHDs is characteristic of a much smaller subset of proteins including CHD4, CHD3, CHD5 and DPF3b. The structural mechanism of the acetylated histone binding by the double PHD fingers of DPF3b has been recently elucidated (Zeng et al. 2010) and represents the first solution structure of a tandem PHD finger domain pair. In the presence of an H3K14ac peptide, the PHD fingers of DPF3b act as a single functional unit with an extensive interface between the two PHD domains in which residues from both domains contribute to binding of a single peptide.
In human CHD4, the two PHD domains are separated by a 32 amino-acid linker, rich in acidic amino acids. This linker is significantly longer than that found in DPF3b. The length and sequence of the linker is well conserved in mammals, amphibians and bony fish. A linker of similar length and amino acid composition is also found in CHD3 and CHD5. Solution structures of the individual PHD1 and PHD2 of CHD4 have been reported and the mechanism of their binding to unmodified and methylated histone H3 has been described (Mansfield et al. 2011; Musselman et al. 2009). A more recent study has shown that together both PHDs of CHD4 can bind to nucleosomes in a multivalent manner, which is required for the repressive functioning of CHD4 (Musselman et al. 2012).
Here we report preliminary NMR studies of the tandem PHD fingers of human CHD4 and the complete 1H, 13C and 15N assignments. These serve as the starting point for the determination of the solution structure, dynamics and histone binding properties of dPHD which will help to elucidate the structural mechanism of histone H3 recognition by the entire tandem PHD unit of CHD4.
Methods and experiments
Protein expression and purification
The portion of the human CHD4 gene encoding the tandem PHD fingers (residues 367–501) was amplified by PCR using an appropriate set of primers. The amplified fragment was cloned in the IPTG-inducible pTriEx2 vector to generate a C-terminal 8xHis-tagged recombinant vector, which was used to transform Rosetta (DE3)pLysS E. coli cells. 15N and 15N/13C labelled recombinant proteins were expressed in the transformed cells grown in 15N-NH4Cl and 13C-glucose enriched M9 minimal medium supplemented with either 1 mM ZnSO4 or ZnCl2. The expressed proteins were first purified using a cobalt affinity column and eluted with a gradient of imidazole and then further purified by size exclusion chromatography in 20 mM Tris pH 7.5, 200 mM NaCl and 1–10 mM DTT.
NMR spectroscopy
NMR samples contained ~1 mM protein in 95 % H2O/5 % D2O at pH 7.5 in 20 mM Tris with 200 mM NaCl and 10 mM DTT. All NMR spectra were acquired at 298 K using either home-built 750 or 950 MHz spectrometers which are controlled with GE/Omega software and are equipped with home-built triple-resonance pulsed-field-gradient probeheads or a Bruker Avance 500 MHz spectrometer with a Cryoplatform, equipped with a TCI CryoProbe. Sequential assignments were carried out initially using 15N-labelled dPHD and 3D 15N-edited TOCSY-HSQC (52 ms mixing time), NOESY-HSQC (150 ms mixing time) and HSQC-NOESY-HSQC (75 ms mixing time) experiments acquired at 950 MHz. Further backbone and side chain assignments were obtained using 15N- and 13C-labelled dPHD and 3D HNCA, HN(CO)CA, CBCANH, CBCA(CO)NH, HBHA(CBCACO)NH, H(CCO)NH, (H)CC(CO)NH, HNCO, HN(CA)CO and HCCH-TOCSY experiments acquired at 500 or 750 MHz.
Extent of assignments and data deposition
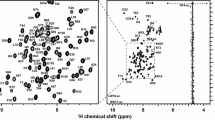
Figure 2 shows the 1H–15N HSQC spectrum of dPHD collected at 950 MHz. Assigned backbone 1HN and 15N are indicated. A total of 95 % of the backbone 1HN and 15N of non-proline residues have been assigned. The side-chain 1HN–15N peaks for the three asparagine, four glutamine and three tryptophan residues are also labelled in Fig. 2. In addition, 95 % of the 1Hα, 93 % of the 13Cα, and 85 % of the 13C′ resonances were assigned. Most of the missing assignments correspond to the N-terminal residues, M1 and E2, three residues in the linker region, E80, E81 and H84, and the C-terminal residues G136 and K137, for which peaks in the 1H–15N HSQC spectrum could not be observed.
Extensive side-chain 1H and 13C assignments have also been made including 95 % of the 1Hβ and 94 % of the 13Cβ. For aliphatic residues, 87 % of the 1Hγ, 88 % of the 13Cγ, 68 % of the 1Hδ, 76 % of the 13Cδ, 44 % of the 1Hε and 36 % of the 13Cε have been assigned. For the Phe, Tyr and Trp residues, only aromatic 1H assignments have been made.
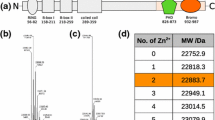
The 13Cβ and 13Cγ chemical shifts of ten of the twelve proline residues of dPHD have been analysed by the procedure of Schubert et al. (2002) to assign cis or trans peptide bond conformation (Fig. 3). We find that P114, located in the second PHD, is likely to adopt a cis conformation. This is confirmed by the 1Hα–1Hα NOE observed between N113 and P114. Interestingly, this residue is assigned a trans conformation in a previous structure determination by 1H NMR alone for the isolated 2nd PHD (Kwan et al. 2003).
Analysis of proline 13Cβ and 13Cγ chemical shifts for dPHD. Circles and triangles correspond to residues with chemical shifts characteristic of a trans and cis conformation, respectively. The open circle and open triangle show the average chemical shifts reported (Schubert et al. 2002) for trans and cis proline, respectively; the bars indicate the standard deviation of these shifts
The chemical shift assignments for dPHD have been deposited in the BioMagResBank (http://www.bmrb.wisc.edu) under the accession number 19968.
References
Baker LA, Allis CD, Wang GG (2008) PHD fingers in human diseases: disorders arising from misinterpreting epigenetic marks. Mutat Res 647:3–12
Eisen JA, Sweder KS, Hanawalt PC (1995) Evolution of the SNF2 family of proteins: subfamilies with distinct sequences and functions. Nucleic Acids Res 23:2715–2723
Kwan AHY, Gell DA, Verger A, Crossley M, Matthews JM, Mackay JP (2003) Engineering a protein scaffold from a PHD finger. Structure 11:803–813
Mansfield RE, Musselman CA, Kwan AH, Oliver SS, Garske AL, Davrazou F, Denu JM, Kutateladze TG, Mackay JP (2011) Plant homeodomain (PHD) fingers of CHD4 are histone H3-binding modules with preference for unmodified H3K4 and methylated H3K9. J Biol Chem 286:11779–11791
Mellor J (2006) It takes a PHD to read the histone code. Cell 126:22–24
Morra R, Lee BM, Shaw H, Tuma R, Mancini EJ (2012) Concerted action of the PHD, chromo and motor domains regulates the human chromatin remodelling ATPase CHD4. FEBS Lett 586:2513–2521
Musselman CA, Mansfield RE, Garske AL, Davrazou F, Kwan AH, Oliver SS, O’Leary H, Denu JM, Mackay JP, Kutateladze TG (2009) Binding of the CHD4 PHD2 finger to histone H3 is modulated by covalent modifications. Biochem J 423:179–187
Musselman CA, Ramirez J, Sims JK, Mansfield RE, Oliver SS, Denu JM, Mackay JP, Wade PA, Hagman J, Kutateladze TG (2012) Bivalent recognition of nucleosomes by the tandem PHD fingers of the CHD4 ATPase is required for CHD4-mediated repression. Proc Natl Acad Sci USA 109:787–792
Pena PV, Davrazou F, Shi XB, Walter KL, Verkhusha VV, Gozani O, Zhao R, Kutateladze TG (2006) Molecular mechanism of histone H3K4me3 recognition by plant homeodomain of ING2. Nature 442:100–103
Ramirez J, Hagman J (2009) The Mi-2/NuRD complex: a critical epigenetic regulator of hematopoietic development, differentiation and cancer. Epigenetics 4:532–536
Schindler U, Beckmann H, Cashmore AR (1993) HAT3.1, a novel Arabidopsis homeodomain protein containing a conserved cysteine-rich region. Plant J 4:137–150
Schubert M, Labudde D, Oschkinat H, Schmeider P (2002) A software tool for the prediction of Xaa-Pro peptide bond conformations in proteins based on 13C chemical shift statistics. J Biomol NMR 24:149–154
Seelig HP, Moosbrugger I, Ehrfeld H, Fink T, Renz M, Genth E (1995) The major dermatomyositis-specific Mi-2 autoantigen is a presumed helicase involved in transcriptional activation. Arthritis Rheum 38:1389–1399
Shi X, Hong T, Walter KL, Ewalt M, Michishita E, Hung T, Carney D, Pena P, Lan F, Kaadige MR, Lacoste N, Cayrou C, Davrazou F, Saha A, Cairns BR, Ayer DE, Kutateladze TG, Shi Y, Cote J, Chua KF, Gozani O (2006) ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. Nature 442:96–99
Wysocka J, Swigut T, Xiao H, Milne TA, Kwon SY, Landry J, Kauer M, Tackett AJ, Chait BT, Badenhorst P, Wu C, Allis CD (2006) A PHD finger of NURF couples histone H3 lysine 4 trimethylation with chromatin remodelling. Nature 442:86–90
Zeng L, Zhang Q, Li S, Plotnikov AN, Walsh MJ, Zhou MM (2010) Mechanism and regulation of acetylated histone binding by the tandem PHD finger of DPF3b. Nature 466:258–262
Zhang Y (2006) It takes a PHD to interpret histone methylation. Nat Struct Mol Biol 13:572–574
Acknowledgments
L.J.W. was supported by a BBSRC studentship. E.J.M. is a Royal Society University Research Fellow. This work was funded by the Medical Research Council UK [New Investigator grant number G0700762/1 to E.J.M.] and supported by the Wellcome Trust core award [090532/Z/09/Z] to the Wellcome Trust Centre for Human Genetics, Oxford. The 500 MHz spectrometer was funded in part by the Wellcome Trust (Grant Number 075330). The 950 MHz spectrometer was funded by awards from the Wellcome Trust Joint Infrastructure Fund and the E.P. Abraham Fund.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Walport, L.J., Morra, R., Mancini, E.J. et al. 1H, 13C, and 15N resonance assignments for the tandem PHD finger motifs of human CHD4. Biomol NMR Assign 9, 239–242 (2015). https://doi.org/10.1007/s12104-014-9582-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-014-9582-y