Abstract
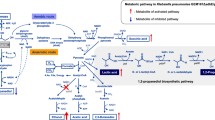
Plasmid-dependent overexpression of enzyme(s) aims to divert carbon flux toward a desired compound. One drawback of this strategy is compromise of growth due to massive consumption of host resources. Here we show that replenishment of sigma factor rpoE improves the growth of Klebsiella pneumoniae. The gene rpoE was expressed alone or coexpressed with Ald4 (an aldehyde dehydrogenase from Saccharomyces cerevisiae) in K. pneumoniae. We found that the Ald4 activity was higher in the strain coexpressing Ald4 and rpoE (32.3 U/mg) than that expressing Ald4 alone (29.9 U/mg). Additionally, under shake-flask conditions, the strain coexpressing Ald4 and rpoE produced 0.5 g 3-hydroxypropionic acid (3-HP) and 9.8 g 1,3-propanediol (1,3-PD) per liter in 24 h, which were 1.6- and 0.85-fold enhancement, respectively, compared to those expressing Ald4 alone. Notably, under non-optimized bioreactor conditions, the strain coexpressing Ald4 and rpoE produced 13.5 g 3-HP and 37.8 g 1,3-PD per liter with glycerol conversion ratio of 0.45 mol/mol. These results indicate that replenishment of rpoE enhanced promoter activity and stimulated glycerol consumption.




Similar content being viewed by others
References
Kumar V, Ashok S, Park S (2013) Recent advances in biological production of 3-hydroxypropionic acid. Biotechnol Adv 31:945–961. doi:10.1016/j.biotechadv.2013.02.008
Ji XJ, Huang H, Ouyang PK (2011) Microbial 2,3-butanediol production: a state-of-the-art review. Biotechnol Adv 29:351–364. doi:10.1016/j.biotechadv.2011.01.007
Li Y, Su MY, Ge XZ, Tian PF (2013) Enhanced aldehyde dehydrogenase activity by regenerating NAD+ in Klebsiella pneumoniae and implications for the glycerol dissimilation pathways. Biotechnol Lett 35:1609–1615. doi:10.1007/s10529-013-1243-1
Forage RG, Lin EC (1982) DHA system mediating aerobic and anaerobic dissimilation of glycerol in Klebsiella pneumoniae NCIB 418. J Bacteriol 151:591–599
Forage RG, Foster MA (1982) Glycerol fermentation in Klebsiella pneumoniae: functions of the coenzyme B12-dependent glycerol and diol dehydratases. J Bacteriol 149:413–419
Ashok S, Raj SM, Rathnasingh C, Park S (2011) Development of recombinant Klebsiella pneumoniae∆dhaT strain for the co-production of 3-hydroxypropionic acid and 1, 3-propanediol from glycerol. Appl Microbiol Biotechnol 90:1253–1265. doi:10.1007/s00253-011-3148-z
Ma Z, Rao Z, Zhuge B, Fang H, Liao X, Zhuge J (2010) Construction of a novel expression system in Klebsiella pneumoniae and its application for 1,3-propanediol production. Appl Biochem Biotechnol 162:399–407. doi:10.1007/s12010-009-8743-4
Wang X, Sa N, Wang FH, Tian PF (2013) Engineered constitutive pathway in Klebsiella pneumoniae for 3-hydroxypropionic acid production and implications for decoupling glycerol dissimilation pathways. Curr Microbiol 66:293–299. doi:10.1007/s00284-012-0271-8
Loewen PC, Hu B, Strutinsky J, Sparling R (1998) Regulation in the rpoS regulon of Escherichia coli. Can J Microbiol 44:707–717
Hengge-Aronis R (2002) Signal transduction and regulatory mechanisms involved in control of the σS (RpoS) subunit of RNA polymerase. Microbiol Mol Biol Rev 66:373–395. doi:10.1128/MMBR.66.3.373-395.2002
Heydorn A, Ersboll B, Kato J, Hentzer M, Parsek MR, Tolker-Nielsen T, Givskov M, Molin S (2002) Statistical analysis of Pseudomonas aeruginosa biofilm development: impact of mutations in genes involved in twitching motility, cell-to-cell signaling, and stationary-phase sigma factor expression. Appl Environ Microbiol 68:2008–2017. doi:10.1128/AEM.68.4.2008-2017.2002
Alper H, Stephanopoulos G (2007) Global transcription machinery engineering: a new approach for improving cellular phenotype. Metab Eng 9:258–267. doi:10.1016/j.ymben.2006.12.002
Weber H, Polen T, Heuveling J, Wendisch VF, Hengge R (2005) Genome-wide analysis of the general stress response network in Escherichia coli: σS-dependent genes, promoters, and sigma factor selectivity. J Bacteriol 187:1591–1603. doi:10.1128/JB.187.5.1591-1603.2005
Merrikh H, Ferrazzoli AE, Bougdour A, Olivier-Mason A, Lovett ST (2009) A DNA damage response in Escherichia coli involving the alternative sigma factor, RpoS. Proc Natl Acad Sci USA 106:611–616. doi:10.1073/pnas.0803665106
Riordan JT, Tietjen JA, Walsh CW, Gustafson JE, Whittam TS (2010) Inactivation of alternative sigma factor 54 (RpoN) leads to increased acid resistance, and alters locus of enterocyte effacement (LEE) expression in Escherichia coli O157: H7. Microbiology 156:719–730. doi:10.1099/mic.0.032631-0
Huang Y, Li Z, Shimizu K, Ye Q (2013) Co-production of 3-hydroxypropionic acid and 1,3-propanediol by Klebseilla pneumoniae expressing aldH under microaerobic conditions. Bioresour Technol 128:505–512. doi:10.1016/j.biortech.2012.10.143
Seoane J, Sin G, Lardon L, Gernaey KV, Smets BF (2010) A new extant respirometric assay to estimate intrinsic growth parameters applied to study plasmid metabolic burden. Biotechnol Bioeng 105:141–149. doi:10.1002/bit.22518
Segall-Shapiro TH, Meyer AJ, Ellington AD, Sontag ED, Voigt CA (2014) A ‘resource allocator’ for transcription based on a highly fragmented T7 RNA polymerase. Mol Syst Biol 10:742. doi:10.15252/msb.20145299
Alper H, Moxley J, Nevoigt E, Fink GR, Stephanopoulos G (2006) Engineering yeast transcription machinery for improved ethanol tolerance and production. Science 314:1565–1568. doi:10.1126/science.1131969
Acknowledgments
This work was supported by grants from National Basic Research Program of China (973 Program) (2012CB725200), National High Technology Research and Development Program (863 Program) (No. 2015AA021003), National Natural Science Foundation of China (Nos. 21276014, 21476011), and the Fundamental Research Funds for the Central Universities (YS1407).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, L., Li, Y. & Tian, P. Enhanced Promoter Activity by Replenishment of Sigma Factor rpoE in Klebsiella pneumoniae . Indian J Microbiol 56, 190–197 (2016). https://doi.org/10.1007/s12088-016-0576-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12088-016-0576-6