Abstract
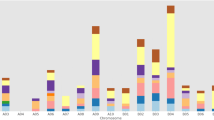
Pineapple is a major tropical fruit and the most important crop processing CAM photosynthesis. It originated in southwest Brazil and northeast Paraguay and survived the harsh, semi-arid environment. Disease resistance genes have contributed to the survival and thriving of this species. The largest class of disease resistance (R) genes in plants consists of genes encoding nucleotide-binding site (NBS) domains. The sequenced genome of pineapple (Ananas comosus (L.) Merr.) provides a resource for analyzing the NBS-encoding genes in this species. A total of 177 NBS-encoding genes were identified using automated and manual analysis criteria, and these represent about 0.6 % of the total number of predicted pineapple genes. Five genes identified here contained the N-terminal Toll/Interleukin-l receptor (TIR) domain, and 46 genes carried the N-terminal Coiled-Coil (CC) motif. A majority of these NBS-encoding genes (84 %) contained a leucine-rich repeat (LRR) domain. A total of 130 of 177 (73 %) of these NBS-encoding genes were distributed across 20 pineapple linkage groups. The identification and characterization of NBS genes in pineapple yielded a valuable genomic resource and improved understanding of R genes in pineapple, which will facilitate the development of disease resistant pineapple cultivars.



Similar content being viewed by others
Abbreviations
- NBS:
-
Nucleotide-binding site
- TIR:
-
N-terminal Toll/Interleukin-l receptor
- CC:
-
Coiled-Coil
- LRR:
-
Leucine-rich repeat
- R:
-
Resistance
References
Ameline-Torregrosa CW, O'Bleness B-B, et al. (2008) Identification and characterization of nucleotide-binding site-leucine-rich repeat genes in the model plant Medicago truncatula. Plant Physiol 146:5–21
Bai J, Pennill LA, Ning J, et al. (2002) Diversity in nucleotide binding site–leucine-rich repeat genes in cereals. Genome Res 12:1871–1884
Bailey TL, Williams N, Misleh C, et al. (2006) MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res 34:W369–W373
Bella J, Hindle K, McEwan P, et al. (2008) The leucine-rich repeat structure. Cell Mol Life Sci 65:2307–2333
Cannon SB, Zhu H, Baumgarten AM, et al. (2002) Diversity, distribution, and ancient taxonomic relationships within the TIR and non-TIR NBS-LRR resistance gene subfamilies. J Mol Evol 54:548–562
Cheng X, Jiang H, Zhao Y, et al. (2010) A genomic analysis of disease-resistance genes encoding nucleotide binding sites in Sorghum bicolor. Genet Mol Biol 33:292–297
Chisholm ST, Coaker G, Day B, et al. (2006) Host-microbe interactions: shaping the evolution of the plant immune response. Cell 124:803–814
Cooley MB, Pathirana S, Wu H-J, et al. (2000) Members of the Arabidopsis HRT/RPP8 family of resistance genes confer resistance to both viral and oomycete pathogens. Plant Cell 12:663–676
Dey KK, Borth WB, Melzer MJ, et al. (2015) Analysis of pineapple mealybug wilt associated virus-1 and-2 for potential RNA silencing suppressors and pathogenicity factors. Viruses 7:969–995
DeYoung BJ, Innes RW (2006) Plant NBS-LRR proteins in pathogen sensing and host defense. Nat Immunol 7:1243–1249
Finn RD, Bateman A, Clements J et al. (2013) Pfam: the protein families database. Nucleic Acids Res:gkt1223
Friedman AR, Baker BJ (2007) The evolution of resistance genes in multi-protein plant resistance systems. Curr Opin Genet Dev 17:493–499
Gambley C, Steele V, Geering A, et al. (2008) The genetic diversity of ampeloviruses in Australian pineapples and their association with mealybug wilt disease. Australas Plant Pathol 37:95–105
Guo Y-L, Fitz J, Schneeberger K, et al. (2011) Genome-wide comparison of nucleotide-binding site-leucine-rich repeat-encoding genes in Arabidopsis. Plant Physiol 157:757–769
Hulbert SH, Webb CA, Smith SM, et al. (2001) Resistance gene complexes: evolution and utilization. Annu Rev Phytopathol 39:285–312
Jia RZ, Ming R, Zhu YJ (2013) Genome-wide analysis of nucleotide-binding site (NBS) disease resistance (R) genes in Sacred Lotus (Nelumbo nucifera Gaertn.) reveals their transition role during early evolution of land plants. Trop Plant Biol 6:98–116
Jiao Y, Li J, Tang H, et al. (2014) Integrated syntenic and phylogenomic analyses reveal an ancient genome duplication in monocots. Plant Cell 26:2792–2802
Jones JD, Dangl JL (2006) The plant immune system. Nature 444:323–329
Jupe F, Pritchard L, Etherington GJ, et al. (2012) Identification and localisation of the NB-LRR gene family within the potato genome. BMC Genomics 13:75
Kobe B, Kajava AV (2001) The leucine-rich repeat as a protein recognition motif. Curr Opin Struct Biol 11:725–732
Kohler A, Rinaldi C, Duplessis S, et al. (2008) Genome-wide identification of NBS resistance genes in Populus trichocarpa. Plant Mol Biol 66:619–636
Konin E, Aravind L (2000) The NACHT family–a new group of predicted NTPases implicated in apoptosis and MHC transcription activation. Trends Biochem Sci 25:223–224
Korf I (2004) Gene finding in novel genomes. BMC Bioinf 5:59
Kunkel BN, Bent AF, Dahlbeck D, et al. (1993) RPS2, an Arabidopsis disease resistance locus specifying recognition of pseudomonas syringae strains expressing the avirulence gene avrRpt2. Plant Cell 5:865–875
Larkin MA, Blackshields G, Brown N, et al. (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Leister D (2004) Tandem and segmental gene duplication and recombination in the evolution of plant disease resistance genes. Trends Genet 20:116–122
Lomsadze A, Ter-Hovhannisyan V, Chernoff YO, et al. (2005) Gene identification in novel eukaryotic genomes by self-training algorithm. Nucleic Acids Res 33:6494–6506
Lozano R, Ponce O, Ramirez M, et al. (2012) Genome-wide identification and mapping of NBS-encoding resistance genes in Solanum tuberosum group phureja. PLoS One 7:e34775
Lozano R, Hamblin MT, Prochnik S, et al. (2015) Identification and distribution of the NBS-LRR gene family in the cassava genome. BMC Genomics 16:360
Malacarne G, Perazzolli M, Cestaro A, et al. (2012) Deconstruction of the (paleo) polyploid grapevine genome based on the analysis of transposition events involving NBS resistance genes. PLoS One 7:e29762
Marchler-Bauer A, Derbyshire MK, Gonzales NR, et al. (2014) CDD: NCBI's conserved domain database. Nucleic Acids Res. doi:10.1093/nar/gku1221
Martin GB, Bogdanove AJ, Sessa G (2003) Understanding the functions of plant disease resistance proteins. Annu Rev Plant Biol 54:23–61
McDonnell AV, Jiang T, Keating AE, et al. (2006) Paircoil2: improved prediction of coiled coils from sequence. Bioinformatics 22:356–358
Mcdowell JM, Simon SA (2006) Recent insights into R gene evolution. Mol Plant Pathol 7:437–448
McDowell JM, Woffenden BJ (2003) Plant disease resistance genes: recent insights and potential applications. Trends Biotechnol 21:178–183
McDowell JM, Dhandaydham M, Long TA, et al. (1998) Intragenic recombination and diversifying selection contribute to the evolution of downy mildew resistance at the RPP8 locus of Arabidopsis. Plant Cell 10:1861–1874
McHale L, Tan X, Koehl P, et al. (2006) Plant NBS-LRR proteins: adaptable guards. Genome Biol 7:212
Meyers BC, Morgante M, Michelmore RW (2002) TIR-X and TIR-NBS proteins: two new families related to disease resistance TIR-NBS-LRR proteins encoded in Arabidopsis and other plant genomes. Plant J 32:77–92
Meyers BC, Kozik A, Griego A, et al. (2003) Genome-wide analysis of NBS-LRR–encoding genes in Arabidopsis. Plant Cell 15:809–834
Ming R, VanBuren R, Wai CM, et al. (2015) The pineapple genome and the evolution of CAM photosynthesis. Nat Genet 47:1435–1442
Monosi B, Wisser R, Pennill L, et al. (2004) Full-genome analysis of resistance gene homologues in rice. Theor Appl Genet 109:1434–1447
Paterson A, Bowers J, Chapman B (2004) Ancient polyploidization predating divergence of the cereals, and its consequences for comparative genomics. Proc Natl Acad Sci USA 101:9903–9908
Porter BW, Paidi M, Ming R, et al. (2009) Genome-wide analysis of Carica papaya reveals a small NBS resistance gene family. Mol Gen Genomics 281:609–626
Ronald PC, Beutler B (2010) Plant and animal sensors of conserved microbial signatures. Science 330:1061–1064
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sether D, Hu J (2002) Yield impact and spread of Pineapple mealybug wilt associated virus-2 and mealybug wilt of pineapple in Hawaii. Plant Dis 86:867–874
Sether D, Melzer M, Busto J, et al. (2005) Diversity and mealybug transmissibility of ampeloviruses in pineapple. Plant Dis 89:450–456
Sievers F, Wilm A, Dineen D, et al. (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal omega. Mol Syst Biol 7:539
Stanke M, Schöffmann O, Morgenstern B, et al. (2006) Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources. BMC Bioinf 7:62
Tamura K, Stecher G, Peterson D, et al. (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tan X, Meyers BC, Kozik A, et al. (2007) Global expression analysis of nucleotide binding site-leucine rich repeat-encoding and related genes in Arabidopsis. BMC Plant Biol 7:56
Tang H, Bowers JE, Wang X, et al. (2010) Angiosperm genome comparisons reveal early polyploidy in the monocot lineage. Proc Natl Acad Sci USA 107:472–477
Tarr DEK, Alexander HM (2009) TIR-NBS-LRR genes are rare in monocots: evidence from diverse monocot orders. BMC research notes 2:197
Ting JP, Davis BK (2005) CATERPILLER: a novel gene family important in immunity, cell death, and diseases. Annu Rev Immunol 23:387–414
Xiao S, Calis O, Patrick E, et al. (2005) The atypical resistance gene, RPW8, recruits components of basal defence for powdery mildew resistance in Arabidopsis. Plant J 42:95–110
Yang S, Feng Z, Zhang X, et al. (2006) Genome-wide investigation on the genetic variations of rice disease resistance genes. Plant Mol Biol 62:181–193
Yang S, Gu T, Pan C, et al. (2008a) Genetic variation of NBS-LRR class resistance genes in rice lines. Theor Appl Genet 116:165–177
Yang S, Zhang X, Yue J-X, et al. (2008b) Recent duplications dominate NBS-encoding gene expansion in two woody species. Mol Gen Genomics 280:187–198
Yue JX, Meyers BC, Chen JQ, et al. (2012) Tracing the origin and evolutionary history of plant nucleotide-binding site-leucine-rich repeat (NBS-LRR) genes. New Phytol 193:1049–1063
Zhou T, Wang Y, Chen J-Q, et al. (2004) Genome-wide identification of NBS genes in japonica rice reveals significant expansion of divergent non-TIR NBS-LRR genes. Mol Gen Genomics 271:402–415
Acknowledgments
We thank Zhenyang Liao and Jingping Fang for providing bioinformatics technical assistance. This work was supported by Fujian Agriculture and Forestry University startup fund to RM.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by: Paulo Arruda
Electronic supplementary material
ESM 1
(XLSX 50 kb)
Rights and permissions
About this article
Cite this article
Zhang, X., Liang, P. & Ming, R. Genome-Wide Identification and Characterization of Nucleotide-Binding Site (NBS) Resistance Genes in Pineapple. Tropical Plant Biol. 9, 187–199 (2016). https://doi.org/10.1007/s12042-016-9178-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12042-016-9178-z