Abstract
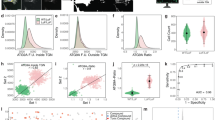
NPC1 gene encodes a transmembrane glycoprotein on the late endosome/lysosomal membrane. Its mutation leads to a rare and aggravated autosomal recessive neurovisceral condition, termed Niemann-Pick disease type C1 (NPC1), which is characterized by progressive neurodegeneration, visceral symptoms, and premature death. To investigate the influence of NPC1 gene deletion on cell morphology, adhesion, proliferation, and apoptosis, CRISPR-Cas9 technology was used to knockout the NPC1 gene in HEK 293 T cells. Sanger sequencing, western blotting, and immunofluorescence were used to confirm successful NPC1 ablation. Filipin staining results indicated that deletion of NPC1 gene led to accumulation of unesterified cholesterol in HEK 293 T cells. Phalloidin staining results revealed cell aggregation, synapse shortening, nuclear enlargement, and cytoskeleton filamentous actin thinning in HEK 293 T cells with NPC1 gene mutation. Furthermore, NPC1 gene mutated HEK 293 T cell showed enhanced cell adhesion, inhibited cell proliferation, and increased cell apoptosis. In addition, NPC1 gene mutations significantly increased the protein expression levels of E-cadherin and γ-catenin and significantly decreased the protein expression levels of Wnt 3a, c-Myc, and cyclin D1. These results suggest that NPC1 may regulate cell adhesion by affecting the cadherin–catenin complex through E-cadherin, and that the classical Wnt signaling pathway may be inhibited by restricting β-catenin from entering the nucleus to inhibit cell proliferation.






Similar content being viewed by others
References
Patterson, M. (1993). Niemann-Pick Disease Type C, in GeneReviews(R). University of Washington.
Loftus, S. K., Morris, J. A., Carstea, E. D., Gu, J. Z., Cummings, C., Brown, A., Ellison, J., Ohno, K., Rosenfeld, M. A., Tagle, D. A., Pentchev, P. G., & Pavan, W. J. (1997). Murine model of Niemann-Pick C disease: Mutation in a cholesterol homeostasis gene. Science, 277(5323), 232–235.
Blanchette-Mackie, E. J. (2000). Intracellular cholesterol trafficking: Role of the NPC1 protein. Biochimica et Biophysica Acta, 1486(1), 171–183.
Naureckiene, S., Sleat, D. E., Lackland, H., Fensom, A., Vanier, M. T., Wattiaux, R., Jadot, M., & Lobel, P. (2000). Identification of HE1 as the second gene of Niemann-Pick C disease. Science, 290(5500), 2298–2301.
Scott, C., & Ioannou, Y. A. (2004). The NPC1 protein: Structure implies function. Biochimica et Biophysica Acta, 1685(1–3), 8–13.
Vanier, M. T. (2010). Niemann-Pick disease type C. Orphanet Journal of Rare Diseases, 5, 16.
Ma, W., Xu, J., Wang, Q., Xin, Y., Zhang, L., Zheng, X., Wang, H., Sun, K., Hui, R., & Huang, X. (2010). Interaction of functional NPC1 gene polymorphism with smoking on coronary heart disease. BMC Medical Genetics, 11, 149.
Higaki, K., Ninomiya, H., Sugimoto, Y., Suzuki, T., Taniguchi, M., Niwa, H., Pentchev, P. G., Vanier, M. T., & Ohno, K. (2001). Isolation of NPC1-deficient Chinese hamster ovary cell mutants by gene trap mutagenesis. Journal of Biochemistry, 129(6), 875–880.
Rodriguez-Pascau, L., Coll, M. J., Casas, J., Vilageliu, L., & Grinberg, D. (2012). Generation of a human neuronal stable cell model for niemann-pick C disease by RNA interference. JIMD Rep, 4, 29–37.
Liu, R., Liang, L., Freed, E. F., & Gill, R. T. (2021). Directed evolution of CRISPR/Cas systems for precise gene editing. Trends in Biotechnology, 39(3), 262–273.
Wiedenheft, B., Sternberg, S. H., & Doudna, J. A. (2012). RNA-guided genetic silencing systems in bacteria and archaea. Nature, 482(7385), 331–338.
Jinek, M., Chylinski, K., Fonfara, I., Hauer, M., Doudna, J. A., & Charpentier, E. (2012). A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science, 337(6096), 816–821.
Zhang, S., Shen, J., Li, D., & Cheng, Y. (2021). Strategies in the delivery of Cas9 ribonucleoprotein for CRISPR/Cas9 genome editing. Theranostics, 11(2), 614–648.
Dalvie, N. C., Lorgeree, T., Biedermann, A. M., Love, K. R., & Love, J. C. (2022). Simplified gene knockout by CRISPR-Cas9-induced homologous recombination. ACS Synthetic Biology, 11(1), 497–501.
Guan, L., Han, Y., Yang, C., Lu, S., Du, J., Li, H., & Lin, J. (2022). CRISPR-Cas9-mediated gene therapy in neurological disorders. Molecular Neurobiology, 59(2), 968–982.
Li, Y., Glass, Z., Huang, M., Chen, Z. Y., & Xu, Q. (2020). Ex vivo cell-based CRISPR/Cas9 genome editing for therapeutic applications. Biomaterials, 234, 119711.
Li, C., Brant, E., Budak, H., & Zhang, B. (2021). CRISPR/Cas: A Nobel Prize award-winning precise genome editing technology for gene therapy and crop improvement. Journal of Zhejiang University. Science. B, 22(4), 253–284.
Rasul, M. F., Hussen, B. M., Salihi, A., Ismael, B. S., Jalal, P. J., Zanichelli, A., Jamali, E., Baniahmad, A., Ghafouri-Fard, S., Basiri, A., & Taheri, M. (2022). Strategies to overcome the main challenges of the use of CRISPR/Cas9 as a replacement for cancer therapy. Molecular Cancer, 21(1), 64.
Du, X., Lukmantara, I., & Yang, H. (2017). CRISPR/Cas9-mediated generation of Niemann-Pick C1 knockout cell line. Methods in Molecular Biology, 1583, 73–83.
Giuliano, C. J., Lin, A., Girish, V., & Sheltzer, J. M. (2019). Generating single cell-derived knockout clones in mammalian cells with CRISPR/Cas9. Current Protocols in Molecular Biology, 128(1), e100.
Livak, K. J., & Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods, 25(4), 402–408.
Guan, L., Chi, W., Xiao, W., Chen, L., & He, S. (2014). Analysis of hypoxia-inducible factor alpha polyploidization reveals adaptation to Tibetan Plateau in the evolution of schizothoracine fish. BMC Evolutionary Biology, 14, 192.
Wang, J., Liu, X., Qiu, Y., Shi, Y., Cai, J., Wang, B., Wei, X., Ke, Q., Sui, X., Wang, Y., Huang, Y., Li, H., Wang, T., Lin, R., Liu, Q., & Xiang, A. P. (2018). Cell adhesion-mediated mitochondria transfer contributes to mesenchymal stem cell-induced chemoresistance on T cell acute lymphoblastic leukemia cells. Journal of Hematology & Oncology, 11(1), 11.
Zhang, C., Wang, W., Lin, J., Xiao, J., & Tian, Y. (2019). lncRNA CCAT1 promotes bladder cancer cell proliferation, migration and invasion. International Brazilian Journal of Urology, 45(3), 549–559.
Bazzoni, G., & Dejana, E. (2004). Endothelial cell-to-cell junctions: Molecular organization and role in vascular homeostasis. Physiological Reviews, 84(3), 869–901.
Garcia, M. A., Nelson, W. J., & Chavez, N. (2018). Cell–cell junctions organize structural and signaling networks. Cold Spring Harbor Perspectives in Biology, 10(4), a029181.
Campbell, H. K., Maiers, J. L., & DeMali, K. A. (2017). Interplay between tight junctions & adherens junctions. Experimental Cell Research, 358(1), 39–44.
Marie, P. J., Hay, E., Modrowski, D., Revollo, L., Mbalaviele, G., & Civitelli, R. (2014). Cadherin-mediated cell–cell adhesion and signaling in the skeleton. Calcified Tissue International, 94(1), 46–54.
van Roy, F., & Berx, G. (2008). The cell-cell adhesion molecule E-cadherin. Cellular and Molecular Life Sciences, 65(23), 3756–3788.
Kaszak, I., Witkowska-Pilaszewicz, O., Niewiadomska, Z., Dworecka-Kaszak, B., Ngosa Toka, F., & Jurka, P. (2020). Role of cadherins in cancer—A review. International Journal of Molecular Sciences, 21(20), 7624.
Hiroki, O. (2012). Evolution of the cadherin–catenin complex. SubCellular Biochemistry, 60, 9–35.
MacDonald, B. T., Tamai, K., & He, X. (2009). Wnt/beta-catenin signaling: Components, mechanisms, and diseases. Developmental Cell, 17(1), 9–26.
Zhang, Y., & Wang, X. (2020). Targeting the Wnt/beta-catenin signaling pathway in cancer. Journal of Hematology & Oncology, 13(1), 165.
Efthymiou, A. G., Steiner, J., Pavan, W. J., Wincovitch, S., Larson, D. M., Porter, F. D., Rao, M. S., & Malik, N. (2015). Rescue of an in vitro neuron phenotype identified in Niemann-Pick disease, type C1 induced pluripotent stem cell-derived neurons by modulating the WNT pathway and calcium signaling. Stem Cells Translational Medicine, 4(3), 230–238.
Funding
This work was supported by grants from the National Natural Science Foundation of China (Grant Nos. 81801127, 81771226, 81800792, and U1804186), Science and Technology Innovative Research Team in Higher Educational Institutions of Henan Province (Grant No. 19IRTSTHN003), Henan Province Science and Technology Project (Grant No. 212102310215), Natural Science Foundation of Henan Province (Grant No. 212300410225), Foundation for the University Key Teacher by the Henan Province (Grant No. 2020GGJS148), Major Cultivation Plan of Scientific and Technological Achievements from Natural science class of Xinxiang Medical University (Grant No. 20172DCG-03), Doctoral Scientific Research Program Foundation of Xinxiang Medical University (Grant No. XYBSKYZZ201523), and Open Program of Henan Key Lab of Biological Psychiatry (Grant No. ZDSYS2015004), Preferential Scientific Research Foundation for Overseas Scholars of Henan Province (Grant No. 40), Research Innovation Support Program for Postgraduates of Xinxiang Medical University (Grant No. YJSCX202135Y).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by ZJ, MY and YZ. The first draft of the manuscript was written by ZJ and LG, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare no financial or commercial conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jia, Z., Yang, M., Zhao, Y. et al. CRISPR-Cas9-Mediated NPC1 Gene Deletion Enhances HEK 293 T Cell Adhesion by Regulating E-Cadherin. Mol Biotechnol 65, 252–262 (2023). https://doi.org/10.1007/s12033-022-00503-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-022-00503-2