Abstract
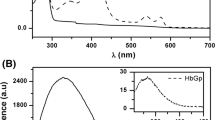
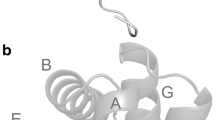
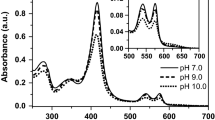
A poly-Lys tag was fused to the Lucina pectinata hemoglobin I (HbI) coding sequence and purified using an efficient and fast process. HbI is a hemeprotein that binds hydrogen sulfide (H2S) with high affinity and it has been used to understand physiologically relevant reactions of this signaling molecule. The (Lys)6-tagged rHbI construct was expressed in E. coli and purified by immobilization on a cation exchange matrix, followed by size-exclusion chromatography. The identity, structure, and function of the (Lys)6-tagged rHbI were assessed by mass spectrometry, small and wide X-ray scattering, optical spectroscopy, and kinetic analysis. The scattering and spectroscopic results showed that the (Lys)6-tagged rHbI is structurally and functionally analogous to the native protein as well as to the (His)6-tagged rHbI. Kinetics studies with H2S indicated that the association (k on) and dissociation (k off) rate constants were 1.4 × 105/M/s and 0.1 × 10−3/s, respectively. This results confirmed that the (Lys)6-tagged rHbI binds H2S with the same high affinity as its homologue.





Similar content being viewed by others
References
Young, C. L., Britton, Z. T., & Robinson, A. S. (2012). Recombinant protein expression and purification: A comprehensive review of affinity tags and microbial applications. Biotechnology Journal, 7, 620–634.
Costa, S., Almeida, A., Castro, A., & Domingues, L. (2014). Fusion tags for protein solubility, purification and immunogenicity in Escherichia coli: The novel Fh8 system. Frontiers in Microbiology, 5, 63.
Nilsson, J., Stahl, S., Lundeberg, J., Uhlen, M., & Nygren, P. A. (1997). Affinity fusion strategies for detection, purification, and immobilization of recombinant proteins. Protein Expression and Purification, 11, 1–16.
Terpe, K. (2003). Overview of tag protein fusions: From molecular and biochemical fundamentals to commercial systems. Applied Microbiology and Biotechnology, 60, 523–533.
Johnson, C. P., Jensen, I. E., Prakasam, A., Vijayendran, R., & Leckband, D. (2003). Engineered protein a for the orientational control of immobilized proteins. Bioconjugate Chemistry, 14, 974–978.
Pessela, B. C., Mateo, C., Carrascosa, A. V., Vian, A., Garcia, J. L., Rivas, G., et al. (2003). One-step purification, covalent immobilization, and additional stabilization of a thermophilic poly-His-tagged beta-galactosidase from Thermus sp. strain T2 by using novel heterofunctional chelate-epoxy Sepabeads. Biomacromolecules, 4, 107–113.
Ley, C., Holtmann, D., Mangold, K. M., & Schrader, J. (2011). Immobilization of histidine-tagged proteins on electrodes. Colloids and Surfaces B, 88, 539–551.
Allard, L., Cheynet, V., Oriol, G., Mandrand, B., Delair, T., & Mallet, F. (2002). Versatile method for production and controlled polymer-immobilization of biologically active recombinant proteins. Biotechnology Bioengineering, 80, 341–348.
Bello-Gil, D., Maestro, B., Fonseca, J., Feliu, J. M., Climent, V., & Sanz, J. M. (2014). Specific and reversible immobilization of proteins tagged to the affinity polypeptide C-LytA on functionalized graphite electrodes. PLoS One, 9, e87995.
Ladaviere, C., Delair, T., Domard, A., Novelli-Rousseau, A., Mandrand, B., & Mallet, F. (1998). Covalent immobilization of proteins onto (Maleic anhydride-alt-methyl vinyl ether) copolymers: enhanced immobilization of recombinant proteins. Bioconjugate Chemistry, 9, 655–661.
Kweon, D. H., Kim, S.-G., Han, N.-S., Lee, J.-H., Chung, K. M., & Seo, J.-H. (2005). Immobilization of Bacillus macerans cyclodextrin glycosyltransferase fused with poly-lysine using cation exchanger. Enzyme and Microbial Technology, 36, 571–578.
Kweon, D. H., Lee, D. H., Han, N. S., Rha, C. S., & Seo, J. H. (2002). Characterization of polycationic amino acids fusion systems for ion-exchange purification of cyclodextrin glycosyltransferase from recombinant Escherichia coli. Biotechnology Progress, 18, 303–308.
Li, J., Zhang, Y., Shen, F., & Yang, Y. (2012). Comparison of magnetic carboxymethyl chitosan nanoparticles and cation exchange resin for the efficient purification of lysine-tagged small ubiquitin-like modifier protease. Journal of Chromatography B, 907, 159–162.
Nock, S., Spudich, J. A., & Wagner, P. (1997). Reversible, site-specific immobilization of polyarginine-tagged fusion proteins on mica surfaces. FEBS Letters, 414, 233–238.
Brewer, S. J., & Sassenfeld, J. M. (1985). The purification of recombinant proteins using C-terminal polyarginine fusions. Trends in Biotechnology, 3, 119–122.
Stempfer, G., Holl-Neugebauer, B., & Rudolph, R. (1996). Improved refolding of an immobilized fusion protein. Nature Biotechnology, 14, 329–334.
Kraus, D. W., & Wittenberg, J. B. (1990). Hemoglobins of the Lucina pectinata/bacteria symbiosis. I. Reactions with ligands. Journal of Biological Chemistry, 265, 16043–16053.
Pietri, R., Roman-Morales, E., & Lopez-Garriga, J. (2011). Hydrogen sulfide and hemeproteins: Knowledge and mysteries. Antioxidants & Redox Signaling, 15, 393–404.
Pietri, R., Lewis, A., Leon, R. G., Casabona, G., Kiger, L., Yeh, S. R., et al. (2009). Factors controlling the reactivity of hydrogen sulfide with hemeproteins. Biochemistry, 48, 4881–4894.
Rizzi, M., Wittenberg, J. B., Coda, A., Fasano, M., Ascenzi, P., & Bolognesi, M. (1994). Structure of the sulfide-reactive hemoglobin from the clam Lucina pectinata. Crystallographic analysis at 1.5 A resolution. Journal of Molecular Biology, 244, 86–99.
Leon, R. G., Munier-Lehmann, H., Barzu, O., Baudin-Creuza, V., Pietri, R., Lopez-Garriga, J., & Cadilla, C. L. (2004). High-level production of recombinant sulfide-reactive hemoglobin I from Lucina pectinata in Escherichia coli. High yields of fully functional holoprotein synthesis in the BLi5 E. coli strain. Protein Expression and Purification, 38, 184–195.
Steen Redeker, E., Ta, D. T., Cortens, D., Billen, B., Guedens, W., & Adriaensens, P. (2013). Protein engineering for directed immobilization. Bioconjugate Chemistry, 24, 1761–1777.
Banerjee, R., Chiku, T., Kabil, O., Libiad, M., Motl, N., & Yadav, P. K. (2015). Assay methods for H2S biogenesis and catabolism enzymes. Methods in Enzymology, 554, 189–200.
Luo, H., Zhao, H., Chang, Y., Wang, Q., Yu, H., & Shen, Z. (2015). Oriented immobilization and characterization of a poly-lysine-tagged cephalosporin C acylase on glyoxyl agarose support. Applied Biochemistry and Biotechnology, 175, 2114–2123.
Chen, Y. H., Chi, M. C., Wang, T. F., Chen, J. C., & Lin, L. L. (2012). Preparation of magnetic nanoparticles and their use for immobilization of C-terminally lysine-tagged Bacillus sp. TS-23 alpha-amylase. Applied Biochemistry and Biotechnology, 166, 1711–1722.
Collazo, E., Pietri, R., De Jesus, W., Ramos, C., Del Toro, A., Leon, R. G., et al. (2004). Functional characterization of the purified holo form of hemoglobin I from Lucina pectinata overexpressed in Escherichia coli. The Protein Journal, 23, 239–245.
Antommattei-Perez, F. M., Rosado-Ruiz, T., Cadilla, C. L., & Lopez-Garriga, J. (1999). The cDNA-derived amino acid sequence of hemoglobin I from Lucina pectinata. Journal of Protein Chemistry, 18, 831–836.
Ramos, C., Pietri, R., Lorenzo, W., Roman, E., Granell, L. B., Cadilla, C. L., & Lopez-Garriga, J. (2010). Recombinant hemoglobin II from Lucina pectinata: A large-scale method for hemeprotein expression in E. coli. The Protein Journal, 29, 143–151.
Blanchet, C. E., & Svergun, D. I. (2013). Small-angle X-ray scattering on biological macromolecules and nanocomposites in solution. Annual Review of Physical Chemistry, 64, 37–54.
Mertens, H. D., & Svergun, D. I. (2010). Structural characterization of proteins and complexes using small-angle X-ray solution scattering. Journal of Structural Biology, 172, 128–141.
Svergun, D. I., Petoukhov, M. V., & Koch, M. H. (2001). Determination of domain structure of proteins from X-ray solution scattering. Biophysical Journal, 80, 2946–2953.
Svergun, D. I. (1992). Determination of the regularization parameter in indirect-transform methods using perceptual criteria. Journal of Applied Crystallography, 25, 9.
Zhang, Y. (2008). I-TASSER server for protein 3D structure prediction. BMC Bioinformatics, 9, 40–48.
Roy, A., Kucukural, A., & Zhang, Y. (2010). I-TASSER: A unified platform for automated protein structure and function prediction. Nature Protocols, 5, 725–738.
Yang, J., Yan, R., Roy, A., Xu, D., Poisson, J., & Zhang, Y. (2015). The I-TASSER Suite: Protein structure and function prediction. Nature Methods, 12, 7–8.
McNicholas, S., Potterton, E., Wilson, K. S., & Noble, M. E. (2011). Presenting your structures: the CCP4 mg molecular-graphics software. Acta Crystallographica Section D, 67, 386–394.
Laspiur, J. P., Anderson, E. R., Ciborowski, P., Wojna, V., Rozek, W., Duan, F., et al. (2007). CSF proteomic fingerprints for HIV-associated cognitive impairment. Journal of Neuroimmunology, 192, 157–170.
Omenn, G. S., States, D. J., Adamski, M., Blackwell, T. W., Menon, R., Hermjakob, H., et al. (2005). Overview of the HUPO Plasma Proteome Project: results from the pilot phase with 35 collaborating laboratories and multiple analytical groups, generating a core dataset of 3020 proteins and a publicly-available database. Proteomics, 5, 3226–3245.
Fernandez-Alberti, S., Bacelo, D. E., Binning, R. C, Jr, Echave, J., Chergui, M., & Lopez-Garriga, J. (2006). Sulfide-binding hemoglobins: Effects of mutations on active-site flexibility. Biophysical Journal, 91, 1698–1709.
Nallamsetty, S., & Waugh, D. S. (2007). A generic protocol for the expression and purification of recombinant proteins in Escherichia coli using a combinatorial His6-maltose binding protein fusion tag. Nature Protocols, 2, 383–391.
Acknowledgments
We are most grateful to Prof. Rafael A. Estremera-Andújar for the helpful discussion in the protein purification process, Ms. Ingrid M. Montes-Rodríguez, Mrs. Jessicca Renta, and Dr. José Casanovas-Nieves for their assistance in the molecular cloning analysis as well as undergraduate students Jay J. Rivera Romero and Ruth Martell. The investigators are most grateful to the assistance of Dr. Ariel Lewis-Ballester and Dr. Syun-Ru Yeh for the use of Stopped Flow equipment for kinetic analysis. The support of the RISE 2BEST Program (NIH-RISE NIGMS grant number R25GM088023, RDA), MARC USTAR Program (5T34GM008419-23), the IGERT Nanomedicine Doctoral Fellowship and the Alfred P. Sloan program is deeply appreciated. This project was supported in part by grants from the National Science Foundation (Grant 0843608, JLG), the National Institutes of Health: National Center for Research Resources (G12-RR003051), the National Institute on Minority Health and Health Disparities (G12-MD007600) and Advancing Competitive Biomedical Research in Puerto Rico (P20GM103475).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Díaz-Ayala, R., Moya-Rodríguez, A., Pietri, R. et al. Molecular Cloning and Characterization of a (Lys)6-Tagged Sulfide-Reactive Hemoglobin I from Lucina pectinata . Mol Biotechnol 57, 1050–1062 (2015). https://doi.org/10.1007/s12033-015-9896-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-015-9896-8