Abstract
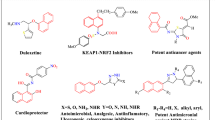
In this study, firstly, 22 thiosemicarbazone derivatives (3a-y) were synthesized. Then, ADME parameters, pharmacokinetic properties, drug-like structures, and suitability for medicinal chemistry of these molecules were studied theoretically by using SwissADME and admetSAR programs. According to the results of these theoretical studies, it can be said that the bioavailability and bioactivity of these compounds may be high. In silico molecular docking between ligands (thiosemicarbazone derivatives) and targeted proteins (protein-78 (GRP78) for C6 and quinone reductase-2 (4ZVM for MCF 7) was analyzed using Hex 8.0.0 docking software. According to the docking data, almost all molecules had higher negative E values than Imatinib (already used as a drug). For this, in vitro anticancer studies of these molecules were done. The cytotoxic activities of thiosemicarbazone derivatives (3a-y) were evaluated on C6 glioma and MCF7 breast cancer cell lines at 24 h, and Imatinib was used as the positive control. According to the results of the cytotoxicity assay, it can be said that the five compounds (3b, c, f, g, and m with IC50 = 10.59–9.08 μg/mL; Imatinib IC50 = 11.68 μg/mL) showed more potent cytotoxic activity than Imatinib on C6 cell line. Together with to these results ten compounds (3b, d, f, g, I, k, l, m, n, and r with IC50 = 7.02–9.08 μg/mL; Imatinib IC50 = 9.24 μg/mL) had a more effective cytotoxic activity against MCF7 cell line than Imatinib. Compound 3 m showed the highest antiproliferative effect against C6 and MCF7 cell lines.


Similar content being viewed by others
References
Hu K, Yang Z, Pan S-S, et al. Synthesis and antitumor activity of liquiritigenin thiosemicarbazone derivatives. Eur J Med Chem. 2010;45:3453–8.
Gürdere MB, Kamo E, Budak Y, et al. Synthesis and anticancer and cytotoxic effects of novel 1,4-phenylene-bis-N-thiocarbamoylpyrazole and 1,4-phenylene-bis-pyrazolylthiazole derivatives. Turk J Chem. 2017;41:179–89.
Joseph M, Kuriakose M, Kurup MRP, et al. Structural, antimicrobial, and spectral studies of copper (II) complexes of 2-benzoylpyridine N (4)-phenyl thiosemicarbazone. Polyhedron. 2006;25:61–70.
Gupta RP, Narayana NL. Synthesis of some Mannich bases of 1-cyclohexylidene-N(1,2-dihydro-2-oxo-3H-indol-3- ylidene) thiosemicarbazones and their antibacterial activity. Pharm Acta Helv. 1997;72:43–5.
Budak Y, Koçyiğit UM, Gürdere MB, et al. Synthesis and investigation of antibacterial activities and carbonic anhydrase and acetyl cholinesterase inhibition profiles of novel 4,5-dihydropyrazol and pyrazolyl-thiazole derivatives containing methanoisoindol-1,3-dion unit. Synth Commun. 2017;47:2313–23.
Khan SA, Asiri AMAA, Khan KA, et al. Synthesis of novel schiff bases by microwave irradiation and their in vitro antibacterial activity. Asian J Chem. 2013;25:8643–6.
Hashmi S, Khan S, Shafiq Z, et al. Probing 4-(diethylamino)-salicylaldehyde-based thiosemicarbazones as multi-target directed ligands against cholinesterases, carbonic anhydrases and α-glycosidase enzymes. Bioorg Chem. 2021;107:104554.
Zambre AP, Kulkarni VM, Padhye S, et al. Novel curcumin analogs targeting TNF-induced NF-jB activation and proliferation in human leukemic KBM-5 cells. Bioorg Med Chem. 2006;14:7196–204.
Pavan FR, da Maia SP, Leite SRA, et al. Thiosemicarbazones, semicarbazones, dithiocarbazates and hydrazide/hydrazones: anti—mycobacterium tuberculosis activity and cytotoxicity. Eur J Med Chem. 2010;45:1898–905.
Denny WA. Prodrug strategies in cancer therapy. Eur J Med Chem. 2001;36:577–95.
Huseynova M, Taslimi P, Medjidov A, et al. Synthesis, characterization, crystal structure, electrochemical studies and biological evaluation of metal complexes with thiosemicarbazone of glyoxylic acid. Polyhedron. 2018;2018(155):25–33.
Ozbek O, Isildak O, Gürdere MB, et al. Cadmium (II)-selective potentiometric sensor based on synthesised (E)-2-benzylidenehydrazinecarbothioamide for the determination of Cd2+ in different environmental samples. Int J Environ Anal Chem. 2020. https://doi.org/10.1080/03067319.2020.1817427.
Isildak Ö, Özbek O, Gürdere MB. Development of chromium(III)-selective potentiometric sensor by using synthesized pyrazole derivative as an ionophore in PVC matrix and its applications. J Anal Test. 2020;4:273–80.
Özbek O. A novel potentiometric sensor for the determination of Pb(II) Ions based on a carbothioamide derivative in PVC matrix. J Turk Chem Soc Sect A. 2022;9(3):651–62.
DeSantis CE, Lin CC, Mariotto AB, et al. Cancer treatment and survivorship statistics. CA Cancer J Clin. 2014;64:252–71.
Fitzmaurice C, Dicker D, Pain A, et al. The global burden of cancer 2013. JAMA Oncol. 2015;1(4):505–27.
Pavlopoulou A, Spandidos DA, Michalopoulos I. Human cancer databases (review). Oncol Rep. 2015;33(1):3–18.
Ritchie DW. Evaluation of protein docking predictions using Hex 3.1 in CAPRI Rounds 1 and 2. Proteins Struct Funct Genet. 2003;52(1):98–106.
Ghoorah AW, Smail-Tabbone M, Devignes MD, et al. Protein docking using case-based reasoning. Proteins Struct Funct Genet. 2013;81:2150–8.
Ritchie DW. Recent progress and future directions in protein-protein docking. Curr Prot Pep Sci. 2008;9(1):1–15.
Macindoe G, Mavridis L, Venkatraman V, et al. HexServer: an FFT-based protein docking server powered by graphics processors. Nucleic Acids Res. 2010;38:W445–9.
Daina A, Michielin O, Zoete V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep. 2017;7:42717.
Yang H, Lou C, Sun L, et al. AdmetSAR 2.0: web-service for prediction and optimization of chemical ADMET properties. Bioinformatics. 2018;35(6):1067–9.
Yang H, Lou C, Sun L, et al. AdmetSAR 2.0: web-service for prediction and optimization of chemical ADMET properties. Bioinformatics. 2019;35(6):1067–9.
Cheng F, Li W, Zhou Y, et al. AdmetSAR: a comprehensive source and free tool for assessment of chemical ADMET properties. J Chem Inf Model. 2012;52(11):3099–105.
Lipinski CA, Lombardo F, Dominy BW, et al. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev. 1997;23(1–3):3–25.
Wolf NB, Kuchler S, Radowski MR, et al. Influences of opioids and nanoparticles on in vitro wound healing models. Eur J Pharm Biopharm. 2009;73:34–42.
da Silva SJ, de Melos LRJ, Lima GS, et al. Synthesis, anti-Trypanosoma cruzi activity and quantitative structure relationships of some fluorinated thiosemicarbazones. J Fluor Chem. 2017;195:31–6.
Matsaa R, Makamb P, Kaushikc M, et al. Thiosemicarbazone derivatives: Design, synthesis and in vitro antimalarial activity studies. Eur J Pharm Sci. 2019;137:104986.
Sardari S, Feizi S, Rezayan AH, et al. Synthesis and biological evaluation of thiosemicarbazide derivatives endowed with high activity toward Mycobacterium Bovis. Iran J Pharm Sci. 2017;16(3):1128–40.
Yakan H, Koçyiğit ÜM, Muğlu H, et al. Potential thiosemicarbazone-based enzyme inhibitors: assessment of antiproliferative activity, metabolic enzyme inhibition properties, and molecular docking calculations. J Biochem Mol Toxicol. 2022;36(5):e23018.
Koçyiğit ÜM, Doğan M, Muğlu H, et al. Determination of biological studies and molecular docking calculations of isatin-thiosemicarbazone hybrid compounds. J Mol Struct. 2022;1264:133249.
Acknowledgements
This work is supported by the Scientific Research Project Fund of Sivas Cumhuriyet University under the project number SHMYO-013 and Scientific Research Projects Commission of Tokat Gaziosmanpasa University (Project Number: 2019/54).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Doğan, M., Koçyiğit, Ü.M., Gürdere, M.B. et al. Synthesis and biological evaluation of thiosemicarbazone derivatives. Med Oncol 39, 157 (2022). https://doi.org/10.1007/s12032-022-01784-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-022-01784-y