Abstract
To determine dietary selenium (Se) status regulates the transcriptions of selenoproteome and activities of selenoenzymes in chicken kidney, 1-day-old chickens received low Se (0.028 mg Se per kg of diet) or super-nutritional Se (3.0 or 5.0 mg Se per kg of diet) in their diets for 8 weeks. It was observed that dietary low or super-nutritional Se did not make renal appearance pathological changes in chicken. Low Se significantly reduced total antioxidant capability (T-AOC), glutathione (GSH) content, but malondialdehyde (MDA) content in the kidney increased and decreased glutathione peroxidase (Gpx) and thioredoxin reductase (TrxR) activity with changes in their mRNA levels. Super-nutritional Se (3.0 mg/kg) increased T-AOC and GSH contents then made them reduce, but it reduced MDA content significantly, elevated then reduced Gpx activity, and decreased TrxR activity with changes in their mRNA levels. Dietary low Se downregulated the mRNA expressions of Gpx1-4, Txnrd3, Sepn1, Selw, Sepx1, Selh, and SEPSECS. At super-nutritional Se, most selenoproteins were upregulated in chicken kidney, but Sepp2 and Sep15 was only upregulated in Se excess (5.0 mg/kg) bird. These results indicated that dietary Se status stabilizes normal renal physiology function via regulation of the selenoprotemic transcriptions and selenoenzyme activities in avian.
Similar content being viewed by others
Introduction
Selenium (Se) is an obligatory nutritional trace element and participates in many aspects of health such as chemoprevention, neurobiology [1, 2], immune function [3, 4], reproduction [5], aging [6], carcinogenesis [7], and muscle metabolism [8, 9]. Poultry is sensitive to Se content of diet [10]. Se deficiency manifested as deformation and necrosis [11] in sheep, cattle, horses, and chickens [12]. The kidney can be ruled out as a product of metabolism in the organism and maintain electrolyte balance. It could also be influenced by Se deficiency or excess, such as predominant pathological changes were convoluted tubule necroses. Concentration of free Se is the greatest in the renal cortex [13]. Se could protect the renal antioxidant function from oxidative damage [14], and Se deficiency or excess causes a lot of selenoprotein resultant metabolic disorders in pig kidney [15]. Evidence of cellular damage (i.e., histopathologic and physiologic, including oxidative stress) was found in kidney systems of young snowy egrets [16]. Se can protect against oxidative damage which leads to the overproduction of free radicals that exert deleterious effects on the kidney [23].
Se is regulated in the body to maintain vital selenoproteins and to avoid toxicity, and it exerts its biological effects mainly through its direct incorporation into selenoproteins as the amino acid, selenocysteine (Sec) [17] (Fig. S1A). The mRNA for a selenoprotein has two distinctive properties: UGA in the open reading frame (ORF) and a specialized stem-loop structure in the 3′ untranslated region (3’UTR). The stem-loop structure is known as a selenocysteine insertion sequence (SECIS) element [18].Three unique trans-acting factors are vital for selenoprotein synthesis. tRNA[ser]sec has the anticodon for UGA. It is initially charged with serine, which is then phosphorylated by O-phosphoseryl-tRNA[ser]sec kinase [19]. The phosphoserine attached to tRNA[ser]sec is converted to selenocysteine by selenocysteine synthase, using monoselenophosphate as the Se donor [20]. Selenocysteyl-tRNA[ser]sec binds to another vital trans-acting factor, the specific elongation factor EFsec [21]. EFsec binds to the third vital trans-acting factor, SECIS-binding protein 2 (SBP2). In addition to binding EFsec, SBP2 has binding sites for the SECIS element and the ribosome. Selenoproteins play an important role in the kidney function. There are intriguing relationships between Se physiology and several derangements associated with acute and chronic kidney disease. Selenoproteins and selenoenzyme can protect the normal function of the kidney [22]. However, effects of dietary Se on selenoproteome in chicken kidney are yet unclear.
Selenoproteome is a set of selenoproteins in eukaryotes and prokaryotes. Avian selenoproteome consists of 24 selenoproteins. Several selenoproteins have been characterized as antioxidant enzymes, serving to alleviate damage caused by reactive oxygen species (ROS) [23]. Some antioxidant enzymes contain Se called selenoenzymes; the most important metabolic roles of Se in cells are due to its function in the active site of these enzymes, for example glutathione peroxidase (Gpx) and thioredoxin reductase (TrxR) [24]. Gpx activity is essential for avoiding peroxide-generated oxidative damage and is provided by different families of enzymes, namely selenoglutathione peroxidases (Gpx; EC 1.11.1.9; glutathione: hydrogen-peroxide oxidoreductase) [25]. The thioredoxin (Trx) system catalyzes the reduction of protein disulfides such as in ribonucleotide reductase, an enzyme essential for DNA synthesis [26]; thioredoxin peroxidases (peroxiredoxins), critical enzymes in the defense against oxidative stress [27]; and protein disulfide isomerase (PDI). Most selenoproteins in the kidney are intracellular enzymes with their activities relying upon a single selenocysteine residue in their main structures [28]. Although 24–25 selenoprotein genes were identified in mammals [26], only a few studies have determined collective responses of these genes to dietary Se concentrations ranging from deficiency to moderately high levels in chicken kidney [29, 30]. The transcriptional response of Se is dependent selenoprotein genes. Thus, systematic data are needed to link selenoprotein and relationship between Se, selenoproteins, selenoenzymatic, and antioxidant function in the kidney, and associated study on poultry is still a vacancy. Here, we tested Gpx, total antioxidant capability (T-AOC), malondialdehyde (MDA) content, 24 selenoproteins, and 7 related genes (Table S2). Our results will describe changes for selenoproteome related to dietary Se at low or super-nutritional levels and the resultant metabolic disorders in chicken kidney which have yet to be determined before.
Materials and Methods
Animals and Treatment
Expt. 1
Chickens (Hy-Line Variety White, n = 120, 1 day old) were fed four different diets, each holding 30 chickens (cocks and hens were randomly distributed). All the groups were feed basal diet (Table S1) except Se. Four experimental diets were prepared as 0.028 mg/kg Se (low), 0.15 mg/kg Se (standard), 3.0 mg/kg Se (excess I), and 5.0 mg/kg Se (excess II). Feed (corn and soybean meal) was from Longjiang County, Heilongjiang province, which is a typically Se-deficient area in China. All other environmental conditions were maintained in accordance with existing norms for the facility. Two identical feed troughs were attached side by side to each cage front. On the seventh day, inoculate Newcastle disease virus and infectious bronchitis vaccines were given; on the 14th day, IBD vaccine; and on the 21st day, Newcastle disease vaccine. Mental state, diet condition, feathers, feces, and other ten indicators were observed with eye viewing and recorded. At the end of the second, fourth, sixth, and eighth week, experimental chickens were fasted for 24 h before weighting and sacrificing. Kidney samples were collected for analysis. Frozen chicken kidney samples were homogenized, the homogenized samples were centrifuged, and the supernatant was transferred to fresh tubes.
Expt. 2
Chickens (n = 32, 1 day old) were treated as Expt. 1. At the end of the eighth week, chickens were sacrificed. The removed kidney tissue is stored in the locker as an extracted RNA sample (CAT3060, Beijing Tiandz, Inc., P.R. China). Chicks were fasted for 24 h before weighting and sacrificing.
Analysis of Kidney Tissue
Every 2 weeks, we randomly selected ten chickens in order to record their kidneys observing the changes with eye viewing and pathological tissue slice. Kidney will be cut into small pieces on the ice packs and were fixed in 10 % formaldehyde and embedded in paraffin for microscopic examination. Sections (5 μm thick) were cut and stained with hematoxylin and eosin (H&E). The pathological changes were observed under a microscope.
Se Determination
Se was determined in feed using inductively coupled plasma mass spectrometry (ICP-MS, Agilent 7500cs-Octopole Reaction Cell, Agilent Technologies, USA). The concentrations of Se in the kidneys were determined with the calibration curve method and compared among the non-, H2, and D2 reaction modes.
Determination of Antioxidant Function
Determination of Protein Content
The protein content was measured with a protein quantitative detection kit (A045-2, Nanjing Jiancheng Bioengineering Institute, P.R. China). Bovine serum albumin (BSA) was used to construct the standard curve.
Determination of Reduced Glutathione Content in the Kidney
The glutathione (GSH) content was measured with a GSH detection kit (A006-1, Nanjing Jiancheng Bioengineering Institute, P.R. China) according to the manufacturer’s protocol. By measuring dihydronicotinamide adenine dinucleotide phosphate (NADPH) oxidation in the presence of oxidized glutathione, and we used tert-butyl hydroperoxide as the peroxide substrate.
Determination of Total Antioxidant Capability in the Kidney
The T-AOC was measured with a T-AOC detection kit (A105, Nanjing Jiancheng Bioengineering Institute, P.R. China) according to the manufacturer’s protocol by converting Fe3+ into Fe2+, whereas the latter forms complexes with phenanthroline substances, which can be measured by a spectrophotometer.
Determination of MDA Content
The content of MDA in kidney was carried out with a MDA detection kit (A003-1, Nanjing Jiancheng Bioengineering Institute, P.R. China) according to the manufacturer’s protocol. By using the thiobarbituric acid method in which MDA, the degradation product of lipid peroxidation, condenses with penthiobarbital and generates a red product with measurable absorbance by a spectrophotometer.
Selenoenzymes Activities Assays
Determination of Gpx Activities in the Kidney
The Gpx activity was measured with a Gpx detection kit (A005, Nanjing Jiancheng Bioengineering Institute, P.R. China) according to the manufacturer’s protocol. Gpx catalyzed GSH into GSSG, and glutathione reductase can use NADPH to catalyze GSSG into GSH; reduction of NADPH can be calculated through testing the vitality of glutathione peroxidase level.
Determination of TrxR Activity in the Kidney
TrxR catalyzed NADPH and made 5, 5′-dithiobis (2-nitrobenzoic acid) (DTNB) into trinitrobenzene (TNB). By measuring the increase of TNB at 412 nm, activity of TrxR can be calculated [31].
Analysis of mRNA Expressions of Selenoproteins and Factors Associated with the Regulation of the Selenoprotein Synthesis
Total RNA was isolated from the kidneys using the RNAout reagent (CAT: 3070, Beijing Tiandz Inc., P.R. China) according to the manufacturer’s protocol. The dried RNA pellets were resuspended in 40 μL water that was diethyl-pyrocarbonate treated. The content and purity of the total RNA were determined spectrophotometrically at 260/280 nm. The total RNA was immediately used to synthesize cDNA.
First-strand cDNA was synthesized from 10 μg of total RNA using Oligo(dT) primers and Transcript Reverse Transcriptase (Beijing TransGen Biotech Co. Ltd., P.R. China) according to the manufacturer’s instructions. Synthesized cDNA was diluted ten times with sterile water and stored at −20 °C before use.
Primer Analysis Software (Oligo 7.24, Molecular Biology Insights, Inc. USA) designs specific primers (Table S1). The relative mRNA expressions of 24 selenoproteins and 7 factors associated with the regulation of the selenoprotein synthesis genes were assayed by qPCR array. To perform the qPCR array, GoTaq® qPCR Master Mix (A6001, Promega, USA), primers, template, and nuclease-free water (Promega, USA) were added to form the reaction mixture (final volumes 20 μL) in one plate according to the manufacturer’s instructions. The qPCR array was performed on a 7500 Real-Time PCR System (Applied Biosystems, USA).
Four internal reference comparisons (β-actin (Actb), glyceraldehyde 3-phosphate dehydrogenase (GADPH), 18S ribosomal RNA (18srRNA), ribosomal protein L19 (RPL19)) were designed (Table S2). All data were normalized to the four internal control genes, and the relative expression levels were calculated using the 2−ΔΔCt method [29]. When the software analyzes the data, it uses the mean value of Ct value of the selected four internal control genes to make the baseline correction. Data analysis of mRNA expression was analyzed using the GeneCopoeia-FulenGen qPCR Array data analysis system online (http://www.genecopoeia.com/product/qpcr/analyse2/index.php).
Statistical Analysis
The data were analyzed statistically using GraphPad Prism 5.1 (GraphPad Software Inc., USA). One-way analysis of variance (ANOVA) and the least significant difference (LSD) post hoc test were used to analyze the data. Differences between the means of data were analyzed by the paired T test which was utilized to determine the effects of Se supplementation. The results were expressed as mean ± S.D. of different groups. The significant differences of all data were showed by ANOVA of each experiment. P < 0.05 is considered significant unless otherwise stated.
Results
Effects of Dietary Se Status on Chicken Kidney
As can be seen in Fig. 1, the weight of chicken in the low Se group was smaller than that in the standard group which was reported in previous paper [10] and its feather was fluffy and messy. In Se excess groups (3.0 and 5.0 Se mg/kg), the size of chickens can be seen a little bigger than those in the standard group. Dietary Se content did not affect kidney performance, no significant differences observed in appearance, symptoms and pathological changes compared with the standard group with the other three groups. From the observation of pathological tissue section, kidney glomerulus, tubule, and interstitial showed no lesions (Fig. 1).
Effects of Dietary Se Status on Renal Antioxidant Function
The levels of Se greatly influence antioxidant defense. Renal GSH contents in the low Se, excess I, and excess II groups were all significantly reduced compared with that in the standard group at the eighth week (P < 0.05) (Table 1). However, lowered T-AOC activity can be seen significantly in the low Se group compared with that in the standard group (P < 0.05), and it increased then decreased significantly in the excess I and the excess II groups compared with that in the standard group (P < 0.05) (Table 1). Meanwhile, the MDA content showed the rising trend in the low Se, the excess I, and the excess II groups at the eighth week compared with that in the standard group (P < 0.05) (Table 1).
Effects of Dietary Se Status on the Selenoenzymes of Chicken Kidney
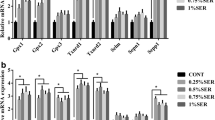
The activity of Gpx in the low-Se group decreased significantly compared with that in the standard group all the 8 weeks (P < 0.05). In the excess I group, it decreased significantly at the eighth week (P < 0.05). In the excess II group, it decreased significantly at the eighth week (P < 0.05) (Table 1). Gpx1-4 mRNA levels significantly increased in the excess I group (P < 0.05) and decreased significantly in the low-Se and excess II groups (P < 0.05) (Fig. 2).
Effects of dietary low-Se or super-nutritional Se on the Gpx1 (a), Gpx2 (b), Gpx3 (c), Gpx4 (d),Txnrd1 (e), Txnrd2 (f), Txnrd3 (g), and TNX (h) mRNA level in the kidney. Four reference comparisons in the diagram. Symbols for the significance of differences between Se standard group and another: * P < 0.05
The TrxR activity decreased significantly in response to low Se at the eighth week compared with that in the standard group (P < 0.05). In the excess I group, it went up significantly in the second week and eighth week compared with that in the standard group (P < 0.05), yet from the fourth week to sixth week it decreased significantly compared with that in the standard group (P < 0.05) (Table 1). TXN mRNA level increased significantly in all the groups compared with that in the standard group (P < 0.05) (Fig. 2). Meanwhile, Txnrd1 and Txnrd2 mRNA levels all increased significantly in the low-Se and the excess I groups (P < 0.05) compared with that in the standard group, whereas decreased significantly in the excess II group (P < 0.05); nonetheless, Txnrd3 mRNA level has no significant changes in the low-Se and excess I groups compared with that in the standard group; in Se excess II group, it decreased significantly (P < 0.05) (Fig. 2).
Effects of Dietary Se Status on Partial Selenoproteins of Chicken Kidney
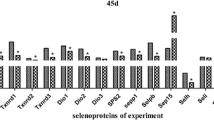
The effects of low and super-nutritional Se on selenoprotein mRNA abundance were determined by qPCR in chicken kidney. The levels of Se greatly regulate the expression of several selenoproteins. All data for mRNA expression of chicken renal selenoproteome was shown in Table S2, and the classifications of histogram were shown in Figs. 3, 4, 5, 6.
Effects of dietary low-Se or super-nutritional Se on the Dio1 (a), Dio2 (b), Dio3 (c), Sepp1 (d), Selu (e), and Selk (f) mRNA level in the kidney. Four reference comparisons in the diagram. Symbols for the significance of differences between Se standard group and another at the eighth week: * P < 0.05
Effects of dietary Se levels on selenoprotein synthesis-related transcription factor mRNAs in chicken kidney. The percentage of upregulated or downregulated multiple on relative mRNA abundance of selenoprotein synthesis factors was compared with the Se standard group at the eighth week. Four reference comparisons in the diagram: * P < 0.05
RNA generated expression values for the 24 selenoprotein transcripts and 7 related factors. Columns represent kidney expression from individual chicken fed a 0.028, 0.15, 3.0, and 5.0 Se mg/kg diet. Rows represent the probe sets. RNA gene expression is shown using the indicated pseudo color scale from green (−10) to red (+10) relative to values. Green squares represent increased significantly (P < 0.05), red squares represent decreased significantly (P < 0.05), and gray squares represent no significant difference (P > 0.05)
Two kinds of trends can be seen: one was that iodothyronine deiodinases (Dio1, Dio2, Dio3) mRNA levels all increased significantly in the low-Se and the excess I groups (P < 0.05) compared with that in the standard group, but decreased significantly in the excess II group (P < 0.05), whereas Dio2 mRNA levels in the low-Se group changed more obviously than that of Dio1 and Dio3, over ten times (Fig. 3). Selenoprotein k (Selk), selenoprotein u (Selu), and selenoprotein p1 (Sepp1) mRNA levels showed the same trend with Dios, and Selk and Selu altered more significantly than Sepp1 (P < 0.05) (Fig. 3).
The other trend was that selenoprotein n1 (Sepn1) and selenoprotein w (Selw) mRNA levels’ variation was similar with each other in chicken kidney and decreased significantly in the low-Se and the excess II groups (P < 0.05) but increased significantly in the excess I group (P < 0.05) (Fig. 4).
Effects of Dietary Se Status on the Selenoprotein Synthesis-Related Transcription Factors
The effects of low Se and super-nutritional Se on selenoprotein synthesis-related transcription factors were determined by qPCR in chicken kidney. Seven selenoprotein synthesis-related transcription factors were studied (Fig. 5). The tRNA selenocysteine-associated protein (SECp43), phosphoseryl-tRNA kinase (PSTK), and seryl-tRNA synthetase (SARS) mRNA levels were upregulated significantly under low-Se condition (P < 0.05), whereas Sep (O-phosphoserine) tRNA:Sec (selenocysteine) tRNA synthase (SEPSECS) mRNA had no expression. Eukaryotic elongation factor, selenocysteine-tRNA-specific (EFsec), SEPSECS, SECp43, PSTK, selenophosphate synthetase 1 (SEPHS1), SECIS-binding protein 2 (SECISBP2), SARS mRNA levels were upregulated significantly in the excess I group (P < 0.05) (Fig. 5). PSTK, SEPHS1, and SECISBP2 mRNA levels were downregulated significantly in the excess II group (P < 0.05), and SEPSECS mRNA had no expression (Fig. 5). We saw that SECp43 and PSTK altered more obviously, then SARS, and finally EFsec, SEPHS1, and SECISBP2.
Regulation of Low and Super-nutritional Se of Selenoproteomic Expression Spectrum
We used selenoproteomic expression spectrum to indicate all selenoproteins and selenoprotein synthesis-related transcription factors [32]. According to different colors, changes of low and super-nutritional Se regulation of the selenoproteomic mRNA expressions were demonstrated (Fig. 6). Partial avian selenoproteins including Sepx1, Selo, Selt, Sels, Sep15, SelI, Selm, and Selh mRNA levels changed differently. Sepx1 mRNA level was downregulated significantly in the low-Se and Se excess I groups (P < 0.05). However, SelI mRNA has no expression in low-Se and Se excess II groups; Selo was the same with SelI in Se excess II group (P < 0.05). Meanwhile, in Selt and Sep15, mRNA levels were upregulated with low-Se and Se excess (3.0 mg/kg). Moreover, Sels mRNA level was upregulated in Se excess (3.0 mg/kg) but downregulated in Se excess (5.0 mg/kg) (P < 0.05). In addition, Selh and Selm mRNA levels were all downregulated with Se excess (3.0 and 5.0 mg/kg), and this can also be observed on Selh in low-Se group (P < 0.05) (Fig. 6).
Discussion
In the present study, we observed that kidneys of chickens fed with low or super-nutritional Se diet evidenced no appearance changes and the renal pathological section did not show differences among the four groups. The nutritional antioxidant Se ameliorated oxidative stress and loss of cellular antioxidants and suggested that Se efficiently protected the kidneys [33]. It is reported that Se can protect the kidney antioxidant defense system [34]. Se influences selenoproteins’ and selenoenzymes’ activity which is closely related to renal function [32]. So we tested the antioxidant function of the kidney to find how Se affected the organism.
GSH and associated antioxidant enzymes are major combatants of oxidative stress that influence redox status [35]. GSH also works as a specific substrate for the enzyme Gpx [36]. Se supplementation was protective of kidney by providing substantial elevations of GSH levels [37]. Super-nutritional Se enhanced the endogenous antioxidant capacity, increasing GSH content. However, the lack of thiol response in the kidney might indicate that the redox cycling between the Se compounds and the reduced intracellular GSH in tench did not occur or other cellular defenses against oxidative stress were induced to prevent oxidative damage [38], and GSH act both as a scavenger of various undesired compounds and their toxic metabolites [39]. Suitable content of Se greatly improved T-AOC and decreased MDA concentration in the kidney [40]. Our result is consistent with previous research. At low or super-nutritional Se levels, chickens increased kidney content of MDA, whereas decreased T-AOC. GSH level increased at first and then decreased. Our results indicated that low or super-nutritional Se increased MDA content, which caused an organism’s GSH to protect itself from oxidative stress and in turn prevents it from being damaged in detoxification reactions.
Measurement of Gpx activity has been used as a marker of adequacy of intake because activity of the enzyme is proportional to dietary intake [41]. Some studies previously indicated that low Se can result in inactivation of Gpx and development of oxidative stress in the kidneys [41], and induced Gpx activity in the kidney of juvenile carp fed 1 mg Se/kg diet for 60 days was the result of the enhanced detoxification ability [42]. In this study, compared with those fed with a normal Se diet, chicken fed with a low or super-nutritional Se diet display significant decreases in renal Gpx activities. It is interesting that Gpx1-4 mRNA levels increased in excess I group, whereas Gpx activity decreased. We suppose that super-nutritional Se (3.0 mg/kg) could enhance mRNA levels, but regulation of mRNA translation can be affected by many factors, such as microRNAs, or the precursor protein translation requires some modification and activation, and this process requires effects of other selenoproteins.
TrxR is a selenocysteine-containing enzyme which catalyzes the NADPH-dependent reduction of Trx and therefore plays a regulatory role in its metabolic activity [43]. Our results indicated that low or super-nutritional Se made chicken renal TrxR activity decrease. It may be because dietary insufficient Se influences TrxR directly or 3.0 and 5.0 mg/kg Se excess indirectly as a result of damaging ER. TrxR is much more vulnerable to oxidative damage than Gpx [44]; hence, its changes were different from Gpx. Txnrd1 and Txnrd2 had different trends maybe because Txnrd1 and 2 are mostly cytoplasmic and mostly mitochondrial, respectively. Txnrd 3 is involved in sperm maturation [43]. It is intriguing that Txnrd1, Txnrd2, and TXN mRNAs increased, whereas TrxR activity decreased. It may be caused by the same reason as those of Gpx we mentioned above.
Up to now, 24 selenoproteins were identified in chicken. In this study, to obtain whether low or super-nutritional Se would affect the renal selenoprotein transcription, mRNA expressions of all avian selenoproteins were analyzed in chicken kidney. It reported that 20 selenoprotein mRNAs in mice kidney were identified, including Gpx1, Selh, Sepw1, Selm, Txnrd1, Dio1, Sep15, and Sels that were significantly downregulated in Se-deficient kidney. In this study, Dio1, Dio2, Dio3, Selk, Selu, and Sepp1 had the same trend that increased significantly in low Se and 3.0 mg/kg Se excess but decreased significantly in 5.0 mg/kg Se excess. These results suggested that dietary Se status influenced the expressions of selenoprotein mRNAs in chicken kidney. In low Se diet, these selenoprotein mRNAs’ levels increased significantly but selenoenzyme activity and antioxidant capacity decreased. One possible explanation could be the mobilization of the Se from the liver to the kidney [45]. A selenoprotein hierarchy is when Se supply is insufficient to support full expressions of all the selenoproteins, the synthesis of some of them is downregulated so that others can be expressed more fully. Moreover, we suggest although Se deficiency made selenoprotein mRNAs’ expression higher, these selenoproteins did not exert function or too many selenoproteins accumulating in the kidney competed with each other leading to selenoproteins not used rationally in the kidney. A 3.0 mg/kg Se excess can increase kidney function and promoted selenoprotein mRNAs’ expression; however, 5.0 mg/kg Se excess might become a harmful accumulation, decrease selenoprotein mRNA level, and then damage antioxidative function. We also suggest that too much Se could cause organism disorder and lead to cell autophagy. Even more noteworthy is Sepp1 is a biomarker of Se status, the kidney is dependent on hepatically derived Sepp1 protein as a Se source, and Selu could potentially be a molecular biomarker of Se status too. Both of them may be near the top of the abundance hierarchy, so their mRNA levels increased in low Se diet. However, it needs to be further validated.
We found the effects of dietary Se status on avian Sepn1 and Selw expression were similar in chicken kidney, decreased significantly in low Se and 5.0 mg/kg Se excess, and increased significantly in 3.0 mg/kg Se excess. Gpx3 and Selw regulation was in agreement with a previous study in rat’s kidney that these two selenoproteins were highly regulated by Se [22, 31]; Selw downregulated by dietary low Se was also associated with our research [22, 31]. Insufficient Se influenced organism antioxidant capacity that decreased these selenoprotein mRNA levels. Se status on mRNA regulation occurs at post-transcription; organism can reduce some selenoprotein mRNAs to save more important selenoproteins in chicken kidney because of hierarchy. We noticed that 3.0 mg/kg Se excess could improve kidney function and increase enzyme activity, as selenoprotein mRNA levels increased. However, 5.0 mg/kg Se excess might burden the kidneys, which caused decreased levels of selenoprotein mRNA in chicken.
Some studies found that most selenoprotein mRNA expressions were not significantly regulated by Se status in mice. In rats’ kidney, Dio1, Dio2, Dio3, Gpx2, Gpx4, Seli, Selm, Selo, Sepp1, Sels, Selt, Sep15, Txnrd2, and Txnrd3 were expressed but not regulated by dietary Se levels. However, the authors disagree with these findings. In chicken kidney, Sepx1, Txnrd3, Sels, and Selt mRNA expressions differed from one another. It was discovered that low or 5.0 mg/kg Se excess levels did not affect Selo or Seli. Sepp2 mRNA levels kept on increasing in low-Se and 5.0 mg/kg Se excess groups. Selh decreased all the time in low or super-nutritional Se, and Selm decreased in Se excess. These results may be due to different selenoproteins in different animal disaffinity. Insufficient Se resulted in selenoenzyme activity decrease, but mRNAs were not reduced in the meantime; 5.0 mg/kg Se excess causes selenoenzyme activity decrease with mRNAs decreasing. Se deficiency may not decrease some mRNA transcription, but prevent mRNAs’ translation into some of selenoproteins, and Se excess directly affects the expression of mRNAs. Ultimately, it results in the decrease of selenoenzyme activity. These discrepancies from observation let us begin to further analyze on synthesis factors related to selenoprotein.
Selenoprotein synthesis requires translational recoding of the UGA codon from a stop signal to a SECIS. Any of these gene mRNAs’ change may influence selenoprotein expression. What we can see in our research is that dietary low Se made PSTK, Secp43, SPS2, and SAR mRNA levels increase significantly. All these gene mRNA levels increased significantly in 3.0 mg/kg Se excess. They, however, decreased in the 5.0 mg/kg Se excess samples. Some selenoproteins seemed to change as the same trend of these genes. The amounts of selenoprotein expression changes were not fully consistent with these factors. Some selenoproteins could be expressed in other ways and this conjecture should be further discussed. It reported that SBP2 is potentially reduced by the thioredoxin and glutaredoxin systems, facilitating its relocation to the ribosomes and reinitiation of selenoprotein translation [46]. SECp43 plays a role in the formation or stabilization of the EFSec–SBP2–Sec tRNA[Ser]Sec complex and promotes the formation and subcellular localization of the SPS1/SLA/SECp43 complex. SECp43 may also assist in the decoding of multiple UGA-Sec codons in selenoprotein and in preventing degradation of selenoprotein mRNAs by the nonsense-mediated-decay pathway [47]. Some genes can directly affect the function of selenoenzyme, resulting in kidney function to reduce, and these genes might influence each other, all linked with one another. One of these factors damaged cause of selenoprotein synthesis interruption, other related factors accumulated. We can also suggest that in low-Se diet, description of selenoprotein synthesis regulation occurs at or before the beginning of the translation for polyclonal antibody protein or UGA encoding selenocysteine terminal polypeptide; description of selenoprotein synthesis regulation occurs at or before the beginning of translation.
Conclusion
Numerous researches suggested that variational Se intake regulated the expression of some selenoproteins more than others. From this observation, the concept of a selenoprotein hierarchy developed. In the present study, it was observed that dietary low or super-nutritional Se did not make renal appearance pathological changes in chicken. However, dietary low Se downregulated the mRNA expressions of Gpx1-4, Txnrd3, Sepn1, Selw, Sepx1, Selh, and SEPSECS. At super-nutritional Se, most selenoproteins were upregulated in chicken kidney, but Selm and Selh were downregulated. These results indicated that dietary Se status stabilizes normal physiology function via regulation of the selenoprotemic transcriptions. We hypothesize that hierarchy of regulation of the transcriptions of selenoproteome makes an important role for renal Se metabolism and transport in avian.
References
Combs Jr GF, Clark LC, Turnbull BW (2001) An analysis of cancer prevention by selenium. Biofactors 14:153–159
Li JL, Gao R, Li S, Wang JT, Tang ZX, Xu SW (2010) Testicular toxicity induced by dietary cadmium in cocks and ameliorative effect by selenium. Biometals 23:695–705
Rayman MP (2000) The importance of selenium to human health. Lancet 356:233–241
Hoffmann PR, Berry MJ (2008) The influence of selenium on immune responses. Mol Nutr Food Res 52:1273–1280
Kaur P, Bansal MP (2005) Effect of selenium-induced oxidative stress on the cell kinetics in testis and reproductive ability of male mice. Nutrition 21:351–357
Martin-Romero FJ, Kryukov GV, Lobanov AV, Carlson BA, Lee BJ, Gladyshev VN, Hatfield DL (2001) Selenium metabolism in Drosophila: selenoproteins, selenoprotein mRNA expression, fertility, and mortality. J Biol Chem 276:29798–29804
Jackson-Rosario SE, Self WT (2010) Targeting selenium metabolism and selenoproteins: novel avenues for drug discovery. Metallomics 2:112–116
Chariot P, Bignani O (2003) Skeletal muscle disorders associated with selenium deficiency in humans. Muscle Nerve 27:662–668
Mahmoud KZ, Edens FW (2005) Influence of organic selenium on hsp70 response of heat-stressed and enteropathogenic Escherichia coli-challenged broiler chickens (Gallus gallus). Comp Biochem Physiol C Toxicol Pharmacol 141:69–75
Xu JX, Cao CY, Sun YC, Wang LL, Li N, Xu SW, Li JL (2014) Effects on liver hydrogen peroxide metabolism induced by dietary selenium deficiency or excess in chickens. Biol Trace Elem Res 159:174–182
Zhang ZW, Zhang JL, Zhang YH, Wang QH, Li S, Wang XL, Xu SW (2013) Effect of oxygen free radicals and nitric oxide on apoptosis of immune organ induced by selenium deficiency in chickens. Biometals 26:355–365
Papp LV, Holmgren A, Khanna KK (2010) Selenium and selenoproteins in health and disease. Antioxid Redox Signal 12:793–795
Drasch G, Mail Der S, Schlosser C, Roider G (2000) Content of non-mercury-associated selenium in human tissues. Biol Trace Elem Res 77:219–230
Hoffman DJ, Eagles-Smith CA, Ackerman JT, Adelsbach TL, Stebbins KR (2011) Oxidative stress response of Forster’s terns (Sterna forsteri) and Caspian terns (Hydroprogne caspia) to mercury and selenium bioaccumulation in liver, kidney, and brain. Environ Toxicol Chem 30:920–929
Liu Y, Zhao H, Zhang Q, Tang J, Li K, Xia XJ, Wang KN, Li K, Lei XG (2012) Prolonged dietary selenium deficiency or excess does not globally affect selenoprotein gene expression and/or protein production in various tissues of pigs. J Nutr 142:1410–1416
Hoffman DJ, Henny CJ, Hill EF, Grove RA, Kaiser JL, Stebbins KR (2009) Mercury and drought along the lower Carson river, Nevada: III. Effects on blood and organ biochemistry and histopathology of snowy egrets and black-crowned night-herons on Lahontan reservoir, 2002-2006. J Toxicol Environ Health A 72:1223–1241
Hoffmann FW, Hashimoto AS, Lee BC, Rose AH, Shohet RV, Hoffmann PR (2011) Specific antioxidant selenoproteins are induced in the heart during hypertrophy. Arch Biochem Biophys 512:38–44
Touat-Hamici Z, Legrain Y, Bulteau AL, Chavatte L (2014) Selective up-regulation of human selenoproteins in response to oxidative stress. J Biol Chem 289:14750–14761
Carlson BA, Xu XM, Kryukov GV, Rao M, Berry MJ, Gladyshev VN, Hatfield DL (2004) Identification and characterization of phosphoseryl-tRNA[ser]sec kinase. Proc Natl Acad Sci U S A 101:12848–12853
Xu XM, Carlson BA, Mix H, Zhang Y, Saira K, Glass RS, Berry MJ, Gladyshev VN, Hatfield DL. (2007) Biosynthesis of selenocysteine on its tRNA in eukaryotes. PLoS Biol 5:e4
Tujebajeva RM, Copeland PR, Xu XM, Carlson BA, Harney JW, Driscoll DM, Hatfield DL, Berry MJ (2000) Decoding apparatus for eukaryotic selenocysteine insertion. EMBO Rep 1:158–163
Benner MJ, Settles ML, Murdoch GK, Hardy RW, Robison BD (2013) Sex-specific transcriptional responses of the zebrafish (Danio rerio) brain selenoproteome to acute sodium selenite supplementation. Physiol Genomics 45:653–666
Sunde RA, Raines AM, Barnes KM, Evenson JK (2009) Selenium status highly regulates selenoprotein mRNA levels for only a subset of the selenoproteins in the selenoproteome. Biosci Rep 29:329–338
Pacini N, Abete MC, Dörr AJ, Prearo M, Natali M, Elia AC (2012) Detoxifying response in juvenile tench fed by selenium diet. Environ Toxicol Pharmacol 33:46–52
Wang Y, Zhan X, Zhang X, Wu R, Yuan D (2011) Comparison of different forms of dietary selenium supplementation on growth performance, meat quality, selenium deposition, and antioxidant property in broilers. Biol Trace Elem Res 143:261–273
Brown KM, Arthur JR (2001) Selenium, selenoproteins and human health: a review. Public Health Nutr 4:593–599
Shanu A, Groebler L, Kim HB, Wood S, Weekley CM, Aitken JB, Harris HH, Witting PK (2013) Selenium inhibits renal oxidation and inflammation but not acute kidney injury in an animal model of rhabdomyolysis. Antioxid Redox Signal 18:756–769
Olson GE, Winfrey VP, Hill KE, Burk RF (2008) Megalin mediates selenoprotein P uptake by kidney proximal tubule epithelial cells. J Biol Chem 283:6854–6860
Kryukov GV, Castellano S, Novoselov SV, Lobanov AV, Zehtab O, Guigo R, Gladyshev VN (2003) Characterization of mammalian selenoproteomes. Science 300:1439–1443
Huang JQ, Li DL, Zhao H, Sun LH, Xia XJ, Wang KN, Luo X, Lei XG (2011) The selenium deficiency disease exudative diathesis in chicks is associated with downregulation of seven common selenoprotein genes in liver and muscle. J Nutr 141:1605–1610
Barnes KM, Evenson JK, Raines AM, Sunde RA (2009) Transcript analysis of the selenoproteome indicates that dietary selenium requirements of rats based on selenium-regulated selenoprotein mRNA levels are uniformly less than those based on glutathione peroxidase activity. J Nutr 139:199–206
Sunde RA, Raines AM (2011) Selenium regulation of the selenoprotein and nonselenoprotein transcriptomes in rodents. Adv Nutr 2:138–150
Kipp AP, Banning A, van Schothorst EM, Méplan C, Coort SL, Evelo CT, Keijer J, Hesketh J, Brigelius-Flohé R (2012) Marginal selenium deficiency down-regulates inflammation-related genes in splenic leukocytes of the mouse. J Nutr Biochem 23:1170–1177
Ognjanović BI, Djordjević NZ, Matić MM, Obradović JM, Mladenović JM, Stajn AŠ, Saičić ZS (2012) Lipid peroxidative damage on cisplatin exposure and alterations in antioxidant defense system in rat kidneys: a possible protective effect of selenium. Int J Mol Sci 13:1790–1803
Marković SD, Djačić DS, Cvetković DM, Obradović AD, Žižić JB, Ognjanović BI, Štajn AŠ (2011) Effects of acute in vivo cisplatin and selenium treatment on hematological and oxidative stress parameters in red blood cells of rats. Biol Trace Elem Res 142:660–670
Burk RF, Hill KE (2015) Regulation of selenium metabolism and transport. Annu Rev Nutr DOI:. doi:10.1146/annurev-nutr-071714-034250
Franson JC, Hoffman DJ, Flint PL (2011) Selenium concentrations and enzyme activities of glutathione metabolism in wild long-tailed ducks and common eiders. Environ Toxicol Chem 30:1479–1481
Sk UH, Bhattacharya S (2006) Prevention of cadmium induced lipid peroxidation, depletion of some antioxidative enzymes and glutathione by a series of novel organoselenocyanates. Environ Toxicol Pharmacol 22:298–308
Erkekoglu P, Giray BK, Kızilgün M, Rachidi W, Hininger-Favier I, Roussel AM, Favier A, Hincal F (2012) Di(2-ethylhexyl)phthalate-induced renal oxidative stress in rats and protective effect of selenium. Toxicol Mech Methods 22:415–423
Flora SJ, Kannan GM, Pant BP, Jaiswal DK (2002) Combined administration of oxalic acid, succimer and its analogue for the reversal of gallium arsenide-induced oxidative stress in rats. Arch Toxicol 76:269–276
Combs Jr GF (2015) Biomarkers of selenium status. Nutrients 7:2209–2236
Elia AC, Prearo M, Pacini N, Dörr AJ, Abete MC (2011) Effects of selenium diets on growth, accumulation and antioxidant response in juvenile carp. Ecotoxicol Environ Saf 74:166–173
Diamond AM (2015) The subcellular location of selenoproteins and the impact on their function. Nutrients 7:3938–3948
Branco V, Canário J, Lu J, Holmgren A, Carvalho C (2012) Mercury and selenium interaction in vivo: effects on thioredoxin reductase and glutathione peroxidase. Free Radic Biol Med 52:781–793
Habibian M, Sadeghi G, Ghazi S, Moeini MM (2015) Selenium as a feed supplement for heat-stressed poultry: a review. Biol Trace Elem Res 165:183–193
Rand JD, Grant CM (2006) The thioredoxin system protects ribosomes against stress-induced aggregation. Mol Biol Cell 17:387–401
Small-Howard A, Morozova N, Stoytcheva Z, Forry EP, Mansell JB, Harney JW, Carlson BA, Xu XM, Hatfield DL, Berry MJ (2006) Supramolecular complexes mediate selenocysteine incorporation in vivo. Mol Cell Biol 26:2337–2346
Acknowledgments
This study was supported by the Program for New Century Excellent Talents in University (grant no. NECT-1207-02), the Program for New Century Excellent Talents In Heilongjiang Provincial University (grant no. 1252-NCET-009), and the Doctor Initial Funding of Northeast Agricultural University (grant no. 2012RBC52). We also acknowledge the valuable help provided by Prof. Shi-Wen Xu in Northeast Agricultural University and all involved workers.
Author information
Authors and Affiliations
Corresponding author
Additional information
Jing-Xiu Xu, Cong Zhang, and Chang-Yu Cao contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
Primers used in qRT-PCR array (DOC 52.5 kb)
Table S2
Results of kidney genes expression in chicken fed the different Se level of diets using qRT-PCR array (XLS 28.5 kb)
Figure S1
The process of selenoprotein synthesis. A: The selenocysteine biosynthesis pathway in eukaryotes. B: Mechanism of selenoproteins biosynthesis in eukaryotes (PPT 210 kb)
Rights and permissions
About this article
Cite this article
Xu, JX., Zhang, C., Cao, CY. et al. Dietary Selenium Status Regulates the Transcriptions of Selenoproteome and Activities of Selenoenzymes in Chicken Kidney at Low or Super-nutritional Levels. Biol Trace Elem Res 170, 438–448 (2016). https://doi.org/10.1007/s12011-015-0470-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12011-015-0470-9