Abstract
Understanding the evolution of evolvability—the evolutionary potential of populations—is key to predicting adaptation to novel environments. Despite growing evidence that evolvability structures adaptation, it remains unclear how adaptation to novel environments in turn influences evolvability. Here we address the interplay between adaptation and evolvability in the peacock fly Tephritis conura, which recently underwent an adaptive change in ovipositor length following a host shift. We compared the evolvability of morphological traits, including ovipositor length, between the ancestral and the derived host race. We found that mean evolvability was reduced in females of the derived host race compared to the ancestral host race. However, patterns of multivariate evolvability (considering trait covariances) were very similar in both host races, and populations of the derived host race had diverged from the ancestral host race in directions of greater-than-average evolvability. Exploration of phenotypic integration patterns further revealed relatively high levels of independent variation in ovipositor length compared to other measured traits, allowing some degree of independent divergence. Our findings suggest that adaptation to novel environments can reduce mean evolvability without major changes in patterns of variational constraints, and that trait autonomy helps facilitate divergence of functionally important traits.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Variation is the raw material for adaptive evolution. Natural selection acts on variation in genotypes, phenotypes and fitness, and the evolutionary potential for response to selection is determined by standing genetic variation (Barrett & Schluter, 2008). There is mounting evidence that evolutionary potential is highly variable across traits and species (Hansen & Pelabon, 2021) and that a lack of evolutionary potential may constraint adaptive divergence (Arnold et al., 2001; Bolstad et al., 2014; Bradshaw & McNeilly, 1991; Houle et al., 2017; McGlothlin et al., 2018; Opedal et al., 2023; Schluter, 2000; Voje et al., 2023). A key question is therefore how evolutionary potential evolves, and how this interacts with patterns of selection to produce specific patterns of evolutionary divergence (Berner et al., 2010; Eroukhmanoff & Svensson, 2011; Henry & Stinchcombe, 2022; Opedal et al., 2022).
Phenotypic variation plays a prominent role in the theoretical framework of evolutionary quantitative genetics, as summarized by the ‘Lande equation’ \(\Delta \overline{{\varvec{z}} }=\mathbf{G}{\varvec{\upbeta}}\) (Lande, 1979), where \(\Delta \overline{{\varvec{z}} }\) represents the change in the population trait mean in response to an episode of selection (\({\varvec{\upbeta}}\)). This framework yields the additive genetic variance–covariance matrix (\(\mathbf{G}\)) as a key measure of evolvability, representing the response per unit strength of selection (\(\mathbf{G}=\Delta \overline{{\varvec{z}} }/{\varvec{\upbeta}}\)). If G remains relatively stable over time, adaptive evolution could be well understood by combining a single G-matrix estimate with data on the dynamics of selection. However, if G itself evolves (Arnold et al., 2008; Jones et al., 2003; Milocco & Salazar-Ciudad, 2022), the predictive power associated with a contemporary estimate of G is critically dependent on our understanding of the dynamics of G (McGlothlin et al., 2018; Steppan et al., 2002; Walsh & Blows, 2009). Previous work suggests that G can evolve, and potential drivers have been identified through simulations (Jones et al., 2004, 2007, 2014; Milocco & Salazar-Ciudad, 2022; Pavlicev et al., 2011). Empirical studies assessing evolutionary divergence in G have provided somewhat conflicting results, with many studies reporting apparent stability over a broad range of time-scales (Henry & Stinchcombe, 2022; Houle et al., 2017; McGlothlin et al., 2022; Opedal et al., 2022; Puentes et al., 2016; Voje et al., 2023) while others have reported that G can change rapidly (Björklund et al., 2013; Cano et al., 2004; Doroszuk et al., 2008; Walter et al., 2018).
One challenge in studies assessing the dynamics of G is the difficulty of formulating testable hypotheses (Pélabon et al., 2010). For example, the structure of variance–covariance matrices may be altered following selection (Penna et al., 2017; Revell et al., 2010) and ancestral bottlenecks are expected to affect current evolvability by reducing genetic variation (Nei et al., 1975). A bottleneck may also increase genetic variance, however, if there are allelic combinations hidden under effects of epistasis, dominance, or the environment (Paaby & Rockman, 2014; Whitlock et al., 2002). Gene flow among diverging lineages can also alter the structure of G (Guillaume & Whitlock, 2007), either through hybridization in sympatry, or as a consequence of selection for avoiding gene flow. For instance, sympatry between two recently diverged conspecific species of woodrats have been suggested to impact the phenotypic covariance as gene flow may increase the combinations of traits available to selection (Dochtermann & Matocq, 2016), and experimental sympatry in Drosophila increase genetic variance (Blows & Higgie, 2003).
A pressing question in a changing environment is if specific selection pressures can reshape trait covariances (Melo & Marroig, 2015), hence altering the future potential for adaptation. To improve our understanding of the effect of selection on evolutionary potential, empirical studies that compare variational properties of diverging populations with known histories of selection are needed. Estimating G-matrices can be logistically challenging and is rarely achieved for more than a few populations. An alternative approach is to focus on the structure of the phenotypic variance–covariance matrix (P) reflecting the sum of genetic, environmental and developmental effects. Assuming that G and P are structurally similar, studies of P can provide a powerful approach to the study of how variational properties evolve. In this work, we apply the analytical framework of evolutionary quantitative genetics to study the evolution of phenotypic covariance structure, which allow us to include a larger set of populations. This approach has been debated (Willis et al., 1991) because developmental and plastic effects can cause P and G to differ in structure. However, there is now both theoretical (Cheverud, 1988) and substantial empirical (Kohn & Atchley, 1988; Porto et al., 2009; Roff & Mousseau, 2005; Sodini et al., 2018) evidence that genetic and environmental variances are correlated (Hansen et al., 2011). The resulting correlation between phenotypic and genetic variances is particularly strong for morphological traits, as demonstrated e.g. by recent work with fruit flies (Houle et al., 2017; Rohner & Berger, 2023; Saito et al., 2023).
Colonization of new environments provide excellent opportunities to address how variation in characters under selection evolve, as changes in selection pressures can be inferred from niche differences, and comparisons to ancestral states provide a natural control. Host shifts are especially well suited to assess how evolutionary potential evolves in a novel environment because we know a priori that ancestral and derived populations are evolving towards different phenotypic optima (Assis et al., 2016). Because changes in P following a host shift can arise from either stochastic loss of variation (genetic drift) during the colonization process or effects of selection on variation, comparing traits hypothesized to be under divergent vs. stabilizing selection in the ancestral and novel environments may yield insights into the relative importance of these processes. Here, we leverage a recent host shift in the peacock fly Tephritis conura (Diegisser et al., 2006a, 2006b, 2007, 2008; Nilsson et al., 2022; Steward et al., 2023) to empirically study the evolution of variation during population divergence.
Adult T. conura flies oviposit into the buds of Circium thistles, and larvae and pupae develop within the buds. The ancestral host plant is Cirsium heterophyllum, and some flies have undergone a recent host shift to C. oleraceum in Northern Europe (Romstock-Volkl, 1997). Whole-genome sequencing data support the presence of two genetic clusters, one for populations infesting the ancestral host race C. heterophyllum and one for populations infesting C. oleraceum (Steward et al., 2023). The patterns of population clustering and historical population sizes are consistent with colonization of the C. heterophyllum specialized populations east of the Baltic and the northern population west of the Baltic from the east, whereas the southern population differs in genomic composition (Steward et al., 2023). The derived host race appears to have originated prior to the last glacial maximum, and the populations that have colonized from the south are genetically similar east and west of the Baltic (Fig. S1; Steward et al., 2023). The presence of populations of both host races from both east and west of the Baltic provides a possibility to test the effect of host race on variation across a set of different environments. Moreover, the host races specializing on the two different host plants coexist geographically in a broad zone where the southern C. oleraceum and the northern, ancestral, C. heterophyllum are both common (Fig. 1a), enabling us to test if regional coexistence with the other host race affects variation.
Sampling design, host plants, and traits investigated. a Parallel sampling of allopatric and sympatric populations of the two host races of T. conura flies east and west of the Baltic. CH denotes the C. heterophyllum host race and CO denotes the C. oleraceum host race. b Size measurements of T. conura. c The ancestral host plant, C. heterophyllum. d The derived host plant, C. oleraceum
There is evidence of adaptation to the specific host plants, most clearly in the length of the ovipositor. Flies infesting C. oleraceum have shorter ovipositors (mean = 1.65 ± 0.1 mm) than flies infesting C. heterophyllum (1.76 ± 0.1 mm; Nilsson et al., 2022), matching the respective bud sizes of the plants (Romstock-Volkl, 1997). This pattern is consistent with selection on the ovipositor to match the bud size of the derived host plant (Diegisser et al., 2007; Nilsson et al., 2022) and is likely genetically determined (Diegisser et al., 2007). Other traits have diverged less between the host races (Table S1-2; Nilsson et al., 2022). Given the observed divergence we expect historical and potentially current directional selection on the length of the ovipositor in the derived host race (Diegisser et al., 2007; Nilsson et al., 2022), whereas selection has likely been stabilizing in the ancestral host race. Moreover, there is empirical evidence for strongly reduced survival on the alternative host plant (Diegisser et al., 2008), suggesting strong host-plant-mediated selection. Such selection may have altered the structure of the phenotypic variance–covariance matrix (P) in the derived host race. In this study, we test hypotheses about the evolution of evolvability following colonization of a new host race by assessing variational patterns following population divergence in T. conura.
Methods
To assess variation in evolvability and how this is affected by the colonization of a new niche in T. conura, we measured morphological traits of flies (see “Quantification of Trait Values” section) from the ancestral and derived host races (see “Population Sampling” section and Fig. 1a) to estimate P-matrices, which were in turn used to quantify evolvability (see “Estimating P-Matrices and Measuring Evolvability” section). First, we test the hypothesis that the host shift has altered evolvability (see “The Effect of a Host Shift on Evolvability and P-Matrices” section). To this end, we compared mean evolvability and P-matrix structure between the host races. Second, to test the hypothesis that the evolvability in populations that coexist regionally with the alternative host race is higher than in allopatric regions due to gene flow between the host races, we compared mean evolvability between sympatric and allopatric populations (see “The Effect of Coexistence on Evolvability” section). To test the additional hypothesis that the inferred directional selection on the ovipositor in the derived host race has affected the phenotypic independence (autonomy) of the ovipositor from other traits compared to what is seen in the ancestral host race, we also quantified conditional evolvability and autonomy (see “Ovipositor Independence” section) and compared these between host races. Finally, we test the hypothesis that ancestral variation can constitute variational constraints on evolutionary divergence (Bolstad et al., 2014; Houle et al., 2017; McGlothlin et al., 2018; Opedal et al., 2023). To do so we first estimated a divergence matrix of variance across mean phenotypes across all populations and assessed its alignment with the P-matrix of the ancestral host race. Then, we specifically asked whether populations of the derived host race had diverged in directions of greater-than-average ancestral variation (see “The Effect of Ancestral Evolvability on Divergence” section).
Population Sampling
To examine the distribution of phenotypic variation within and among fly populations specialized on the derived and ancestral host plant we sampled flies from four populations of each host race (Fig. 1a). The two host races are largely reproductively isolated due to differences in mating behavior, phenology and reduced survival on the alternative host plant (Diegisser et al., 2008; Romstock-Volkl, 1997), yet there is tentative evidence of gene flow between them (Diegisser et al., 2006b; Steward et al., 2023). To examine how variational properties (P-matrix structure) vary across host plant races and between allopatric and sympatric regions, we sampled two allopatric and two sympatric populations of each host race, one on each side of the Baltic (Fig. 1a). We use the terms sympatric and allopatric to refer to the presence of one or both thistle hosts on a regional scale. We collected thistle buds infested by T. conura during the pupal stage and eclosed adults in a common laboratory environment as described in Nilsson et al. (2022). In total, we sampled 573 flies (285 females and 288 males) from eight different populations with sample sizes ranging between 16 and 47 (median = 38) for females and between 17 and 50 (median = 36.5) for males (Table S3).
Quantification of Trait Values
The data collection and data are described and illustrated in detail in Nilsson et al. (2022) and can be summarized as follows. After collecting T. conura adults, one female and one male per bud were euthanized by freezing a few days after eclosion. To quantify morphology we used a Celestron 44308 USB microscope to take magnified images. We took one lateral image of the fly body after removal of the wings and one dorsal image of the right wing on a transparent background to enable quantification of melanization. We measured body length, ovipositor length (Fig. 1b), wing length, wing width and wing area digitally from these images. Trait measurements in units of pixels were converted to units of mm using a scale that was included in all photographs. All length and width measurements were subsequently loge-transformed, while the area of the wing was square rooted, then loge-transformed to account for differences in trait dimensionality. The melanised area of the wing was measured through an automated script developed in MATLAB (Matlab, 2017) as in Nilsson et al. (2022). Because loge-transformation scale traits proportionally, they are comparable to mean-standardization and the measures of variance derived from these measurements can be interpreted as proportional variances (Houle et al., 2017; Voje et al., 2023). All subsequent statistical analyses were performed in R version 3.6.1 (R Core Team, 2019).
Estimating P-Matrices and Measuring Evolvability
Evolutionary potential—evolvability—can be measured as a mean-scaled additive genetic variance (Houle, 1992). For multivariate phenotypes, evolvability measures are typically derived from mean-scaled additive genetic variance matrices (G) (Hansen & Houle, 2008) obtained from quantitative-genetic breeding experiments. In the following analyses of mean-scaled phenotypic variance matrices (P) we will refer to patterns of variation as ‘evolvability’ but note that this rests on the assumption of strongly correlated environmental and genetic variation (Hansen et al., 2011).
We estimated P-matrices by fitting multivariate mixed models with the MCMCglmm R package (Hadfield, 2010) and subsequently postprocessed the posterior distributions with tools from the evolvability R package (Bolstad et al., 2014). For each model, we sampled the posterior distributions for 1 million MCMC iterations, with a burn-in of 500000 and a thinning interval of 500. We set uninformative priors for fixed effects (Hadfield, 2010). We estimated different P-matrices to address each research question. To test for differences in evolvability between host races, we estimated mean P-matrices for each host race while including population as a fixed factor. To test for differences in evolvability between allopatric and sympatric populations, we estimated mean P-matrices for allopatric and sympatric populations, while keeping population as a fixed factor. To assess whether the sampled populations differed in evolvability, we estimated and compared P-matrices for each population separately.
To illustrate the distribution of variation within and among populations, we fitted separate univariate linear mixed-effect models to loge-transformed trait values, with population as a random effect. We then computed the among-population variance component as the variance among populations divided by the sum of the among-population and residual (within-population) variance.
The Effect of a Host Shift on Evolvability and P-Matrices
To test for differences in evolvability between host races, we compared the estimated P-matrices in several ways. First, we tested for differences in mean evolvability by deriving posterior means and credible intervals of mean evolvability from each estimated P-matrix, and by assessing posterior support for a difference in mean evolvability (i.e. the proportion of posterior estimates for which the evolvability was greater for the more evolvable host race). Then, to assess differences in P-matrix shape (i.e. patterns of covariances) between the ancestral and derived host race, we correlated the expected responses to a set of random hypothetical selection gradients for the two P-matrices (Hansen & Houle, 2008). We generated 1000 random selection gradients drawn from the unit sphere, and used the evolvabilityBeta function of the evolvability R package (Bolstad et al., 2014) to compute the evolvability (expected response) along each selection gradient for each matrix. Highly correlated expected responses would indicate similar shape of the P-matrices.
The Effect of Coexistence on Evolvability
To test the hypothesis that phenotypic variance is higher in sympatric flies that coexist with the other host race than in allopatric flies, we compared mean evolvability between host races in the same way as above.
Ovipositor Independence
Patterns of trait divergence allowed us to formulate hypotheses about past or current patterns of selection on different traits. The ovipositor is shorter in the derived host race (Diegisser et al., 2007; Nilsson et al., 2022), and is thus likely to either be or have been under directional selection to match the bud size of the derived host plant. The directional selection on ovipositor length in the derived host race may have depleted the variation in ovipositor length, and of the available combinations of ovipositor length and other traits within the fly populations. By comparing the autonomy, i.e. the fraction of variance in a trait that is not bound up in correlations with other traits (Hansen & Houle, 2008), of ovipositor length between the ancestral host race and the derived host race, we examined if there was evidence for reduced autonomy in the derived host race. If the derived host race has a lower autonomy of ovipositor length than the ancestral host race, it may be an effect of reduced available variation. We further assessed if the autonomy of the ovipositor differed from the autonomies of other traits by comparing the autonomy of each individual trait conditioned on ovipositor length to the pairwise autonomies of the same traits conditioned on other non-ovipositor traits. We also evaluated the effect of trait covariances on evolvability using the concept of conditional evolvability, which is the response to selection in a trait given that correlated traits does not change (Hansen et al., 2003). Conditional evolvabilities are directly related to autonomy in that conditional evolvability = evolvability × autonomy (Hansen & Houle, 2008). To obtain evolvability of traits conditioned on ovipositor length, we first conditioned the entire P-matrix on ovipositor length by using the conditionalG function of the evolvability package (Bolstad et al., 2014), and then computed the ratio of the mean evolvability of this conditioned matrix and the mean evolvability of the original matrix with the ovipositor excluded.
The Effect of Ancestral Evolvability on Divergence
Our sampling of both the ancestral and derived host race also allowed us to assess whether populations of the derived host race have diverged in directions of comparatively high evolvability. We address this by asking whether patterns of host race divergence align with variation within the ancestral host race, resulting in divergence in a direction of comparatively high evolvability (Schluter, 1996). We investigated this question in two complementary ways, following Opedal et al. (2023). First, we estimated a variance–covariance matrix among loge-transformed population means (divergence matrix, D), and assessed its alignment with P estimated for the ancestral host race. Second, we considered the divergence of each population of the derived host race from the mean phenotype of the ancestral host race. We computed divergence vectors \(\Delta {\overline{{\varvec{x}}} }_{log}\) from a focal population to the mean of the ancestral host race as \(\Delta {\overline{{\varvec{x}}} }_{log}={{\text{log}}(\overline{{\varvec{x}}} }_{1})-{\text{log}}({\overline{{\varvec{x}}} }_{A}\)), where \({\overline{{\varvec{x}}} }_{A}\) is the vector of mean phenotypes for the ancestral host race. We then computed the evolvability along \(\Delta {\overline{{\varvec{x}}} }_{log}\) as e(\(\Delta {\overline{{\varvec{x}}} }_{log}\)) = \(\Delta {\overline{{\varvec{x}}} }_{log}\) TP \(\Delta {\overline{{\varvec{x}}} }_{log}\), and compared this to the minimum, mean and maximum evolvability of the ancestral P-matrix (Opedal et al., 2023; Voje et al., 2023). Divergence in a direction of greater-than-average evolvability would be consistent with some influence of ancestral variance on divergence.
To assess whether the ovipositor plays a particular role in driving patterns of divergence, we repeated the divergence-vector analyses using three different measures of evolvability, namely (1) raw evolvability, (2) evolvability conditioned only on ovipositor length, and (3) overall conditional evolvability. A different pattern of evolvability in the direction of divergence for the latter would indicate that the ovipositor plays a special role in driving patterns of population divergence.
Results
Differences among populations explained an average of 13.4% (range across traits = 2.6–21.9%) of the female trait variance in the ancestral host race, and an average of 33.7% (range = 26.9%–42.4%) in the derived host race (Fig. 2).
Proportional trait variance among and within populations for females of each host race. The lighter portion of each bar represents variance among populations and the darker portion represents variance within populations. Trait abbreviations are BL body length, OL ovipositor length, WL wing length, WW wing width, WA wing area and MA melanised area
The Effect of a Host Shift on Evolvability and P-Matrices
We found small but detectable differences in evolvability between females of the two host races. The mean evolvability for females of the derived host race was 21.9% lower than that for females of the ancestral host race (mean difference 2.44 \(\times\) 10–4 ± 3.94 \(\times\) 10–6, posterior support 99.98%; Fig. 3a and c). To illustrate host race differences in P-matrices, we graphically explored the two leading axes of the estimated P-matrices, which represented 91.1% (male) and 86.9% (female) of the total phenotypic variation (Fig. S2). Females of the ancestral host race had a slightly broader distribution along the first axis compared to the derived host race (Fig. 3a). The distribution along the second axis was similar for females of both host races (Fig. 3a). In contrast, male variances were similar between host races (Fig. 3b). More quantitatively, the ancestral and derived P-matrices were similar in structure, as indicated by a strong correlation between predicted responses to a set of random selection gradients (posterior mean R2 with 95% CI: 0.94 [0.88, 0.98]), yet the relationship differed from a one-to-one slope (posterior mean β with 95% CI 1.45 [1, 2.01]). The latter reflects the greater size of the ancestral P-matrix and indicates that the derived P carries less variation than the ancestral P in directions of high variance (Fig. 3a, c). The host race difference in female evolvability when excluding ovipositor length is a 25.9% reduction in the derived host race (mean difference = 2.92 \(\times\) 10–4 ± 4.24 \(\times\) 10–6, posterior support 99.1%). Univariate evolvability in ovipositor of the ancestral host race was 3.08% higher than in the derived host race (CH-flies = 0.115 and CO-flies = 0.111).
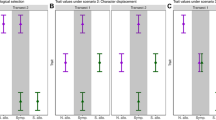
Comparisons of evolvability between host races, between sympatric and allopatric populations, and among populations. a and b Representation of phenotypic variation along the first two eigenvectors for females (a) and males (b). Purple represents flies belonging to the C. heterophyllum host race and green represents flies belonging to the C. oleraceum host race. Solid squares represent host race means, open squares represent population means. Solid ellipses show 95% confidence interval of the overall measurements per host race, whereas the dashed ellipses show 95% confidence interval of the population measurements. c Population and host race mean evolvability for all traits in females. d Population and host race mean evolvability for all traits in males. Colors indicate host races; triangles denote sympatric populations and circles allopatric populations. Solid lines represent mean evolvability and dashed lines represent 95% confidence intervals of the mean evolvability, and individual error bars represent 95% confidence intervals for each population
The male P-matrices were more similar between the host races than were the female P-matrices (Fig. 3b), and we failed to detect a difference in evolvability between males of the two host races. Even though the evolvability was 10% lower in the derived host race compared to the ancestral host race, the posterior support for a difference was moderate (79.6%; Fig. 3d; Table S4).
The Effect of Coexistence on Evolvability
We found no support for the hypothesis that gene flow increases evolvability, as there were no detectable differences in mean evolvability between allopatric and sympatric flies of either sex. Contrary to our prediction, we found a 9.7% increase in evolvability in allopatric compared to sympatric populations in females (posterior support 23.9%; Fig. 3c; Table S4). The patterns in males mirrored those found in females (posterior support 13%; Fig. 3d; Table S4).
Ovipositor Independence
The length of the ovipositor was less integrated with the other traits investigated, as indicated by greater autonomy for traits conditioned on ovipositor length than when conditioned on the average trait, consistently in both host races (Fig. 4). In females of the ancestral host race, the autonomy of an average trait conditioned on ovipositor length was 81.2%, whereas it was 89.7% for females of the derived host race. In contrast, the autonomy of traits when conditioned on the average trait (other than ovipositor length) was 45.4% for the ancestral host race and 55.4% for the derived host race.
Autonomy (proportional remaining evolvability) of any individual trait when conditioned either on ovipositor length or on any other trait. Traits are body size, wing length, wing width, wing area and melanisation area. Shaded bars represent traits conditioned on ovipositor length and filled bars traits conditioned on the average of traits other than ovipositor length. Colors represent host race or mode of coexistence. Error bars represent 95% credible intervals
The Effect of Ancestral Evolvability on Divergence
To assess if populations have diverged in directions of high evolvability, and thus how well P predicts divergence, we correlated the magnitude of evolutionary divergence among populations to the magnitude of standing variation in the ancestral host race, as given by the diagonal of P. The traits line up almost perfectly (β = 1.26 ± 0.41, R2 = 0.98; Fig. 5).
Relationship between evolutionary divergence (D-matrix) and ancestral within-population variation (P-matrix) for all traits. Circles represent traits melanisation area, wing area, wing width, wing length, ovipositor length and body length, from left to right. Posterior mean R2 = 98% [95% CI 0.96, 0.99]. Thus, evolutionary divergence was strongly correlated to the ancestral within-population variation
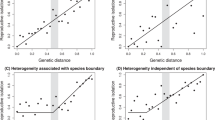
To complement the trait-focused analyses, we also asked if populations of the derived host race had diverged in directions of greater-than-average evolvability from the ancestral host race. As shown in Fig. 6, this was generally the case. The populations of the derived host race had diverged by about 13–18% from the mean phenotype of the ancestral host race, and always in directions of greater-than-average evolvability and conditional evolvability (Fig. 6, Table S5). Patterns were very similar for ovipositor-conditioned evolvability, comparative to overall conditional evolvability. We also detected a tendency for populations diverging in directions of greater evolvability to have diverged slightly more (Fig. 6), which corroborates the strong correlation between variation and divergence (Fig. 5).
Patterns of evolvability along divergence vectors from each population of the derived host race to the mean phenotype of the ancestral host race. Horizontal black lines indicate the minimum, mean and maximum evolvability of the ancestral P-matrix. Horizontal stippled grey line indicates mean conditional evolvability of the ancestral P-matrix. Note that all populations have diverged in directions of greater-than-average evolvability, ovipositor-conditioned evolvability and conditional evolvability. Error bars represent 95% credible intervals. Populations are, from left to right, COLI, COES, COGE and COSK
Discussion
We addressed to which extent a recent host shift, exerting directional selection on the length of the ovipositor, had altered P-matrices in the fly T. conura. Female P-matrices were smaller in the derived host race compared to the ancestral host race, yet the host races had retained very similarly shaped P-matrices. This result is consistent with the finding of reduced overall variation in mice subject to artificial selection (Penna et al., 2017). The authors also found that evolvability increased in the direction of selection, in contrast to our finding. Although we infer net selection on the ovipositor from the shorter ovipositor of the derived host race and its correspondence with host-plant bud size (Romstock-Volkl, 1997), rather than through direct estimates, this difference would be interesting to follow up on in future studies. One explanation for the reduction observed in our data is past or current directional selection acting on several traits following the host shift (Diegisser et al., 2006a, 2006b, 2007, 2008; Nilsson et al., 2022). Alternatively, reduced variation may reflect stochastic losses of variation when a subsample of the population colonized the new host plant. Interestingly, the variance reduction in the derived host race is less pronounced in males (Fig. 3), which holds true when comparing females without ovipositor and males (Fig S3). Males have similar levels of evolvability as females of the derived host race. This difference could reflect historically strong selection on ovipositor length, which may have had effects on correlated traits in males, as there is no strong evidence for sex differences in additive variation generally (Wyman & Rowe, 2014). Exploring further sex differentiated trait correlations in this system would be very intriguing.
The length of the ovipositor is functionally important because a mismatch between ovipositor length and bud size results in reduced female reproductive success (Romstock-Volkl, 1997). Therefore, the difference in ovipositor length between the host races (Diegisser et al., 2007; Nilsson et al., 2022) likely reflects adaptation to the derived host race. Historical directional selection may have caused the observed reduction in standing variation in females of T. conura, yet it is unclear whether there is current directional selection on the length of the ovipositor. If the population mean phenotype is already close to the optimum associated with the new host plant, the ovipositor may currently be under stabilizing selection. Under this scenario, the reduction in variation may result from a combination of the influence of past directional and current stabilizing selection on the ovipositor, and possibly correlated responses to stabilizing selection on other morphological traits. Future work measuring selection on the ovipositor would be necessary to shed light on current selection pressures.
The influence of selection on the ovipositor on correlated traits would be reduced if the ovipositor can evolve independently from other characters (Armbruster et al., 2014; Klingenberg, 2008; Wagner & Altenberg, 1996; Wagner et al., 2007). Using the concepts of conditional evolvability and autonomy, we demonstrated that this is at least partly true, because the covariance between ovipositor length and other traits reduced available variation less than did the covariances among other traits when conditioned among themselves (Fig. 4). This suggests that the ovipositor may be, to some extent, developmentally and functionally separated from the other study traits. Because autonomy has not changed between the host races, this “quasi-independence” might have been in place in ancestral populations and facilitated the observed divergence. Our results are consistent with previous work suggesting slow evolution of autonomy (covariance patterns) and thus a role of trait covariances as variational constraints in nature (McGlothlin et al., 2022; Pélabon et al., 2014).
One possible explanation for the observed variance reduction in the derived host race is that a stringent bottleneck event occurred during the host transition phase, as suggested by the lower variation in mitochondrial haplotypes in the derived host race (Diegisser et al., 2006b). This would substantially reduce the variation available within the ancestral gene pool (Nei et al., 1975), and potentially evolvability. Moreover, the host races overlap in phenology by only 16% (Romstock-Volkl, 1997), and larval survival on the alternative host plant is 10% (Diegisser et al., 2008) suggesting strong selection on phenological adaptation and genes involved in larval metabolism, in addition to selection on the ovipositor.
The host shift has reduced the size of P in the derived host race in females, as expected if multifarious selection to adapt to the novel host plant has contributed to the overall reduction in variation. Interestingly, our analyses suggested limited changes in the shape of P (Fig. 3a, b). Our findings add to the evidence that P (or G) may change following divergence (Björklund et al., 2013; Eroukhmanoff & Svensson, 2011), and revealed that the change can take place without changing the pattern of variational constraints. Given the small change in the structure of P-matrices across populations, it is not surprising that the contemporary P-matrices align well with the pattern of divergence among populations. Our results corroborate a growing body of evidence that the standing variation of contemporary populations are correlated with divergence in a wide range of taxa and time scales (Opedal et al., 2023; Voje et al., 2023). Our study adds a novel dimension to this line of research in that ours is one of the few that have compared populations evolving under clearly different phenotypic optima. Provided multiple lines of evidence for the adaptation associated with the host shifts in this species (Diegisser et al., 2007; Nilsson et al., 2022), it is puzzling why it did not change the covariance pattern at all. While the mechanistic basis of adaptation without alterations of genetic architecture underlying phenotypic covariances will be a question for future investigation, our study demonstrates that standing variation of contemporary populations may enable predicting and explaining patterns of evolution over long time scales even when dramatic adaptations are involved in these processes.
An additional factor that could affect evolvability is gene flow between the host races, as gene flow could increase the available genetic variation and thereby increase evolvability in sympatry (Blows & Higgie, 2003; Dochtermann & Matocq, 2016; Gompert et al., 2017). Contrary to this prediction, allopatric and sympatric populations had similar levels of evolvability. Thus, there is no indication that gene flow affects evolvability in T. conura. One interpretation of this result is that novel genetic variants introduced by gene flow does not necessarily have phenotypic consequences. This may be the case particularly because the traits in our study most likely have highly polygenic genetic architecture (Noble et al., 2017). Alternatively, the lack of effects on the evolvability of T. conura could be explained by the age of secondary sympatry, as evolvability is predicted to increase following early gene flow due to linkage disequilibrium, but may be reduced to normal levels after a period of recombination (Tufto, 2000). Alternatively, genetic drift may play a role in the lack of differences between sympatric and allopatric populations, as several of the sympatric populations were sampled from the edges of the distributions of the T. conura host races. There, effective population sizes may be smaller than in range center populations. Genetic drift affects the phenotype (P) but in stochastic ways (Roff & Mousseau, 2005), and a comparison of P and neutral sites would be needed to assess the effects of drift on T. conura evolvability.
We have employed phenotypic covariance structure (P) rather than genetic covariance structure (G) in this study. The advantage of focusing on P rather than G is that P can more readily be estimated for a large set of populations. The potential downside is that P may differ structurally from G, so that P yields biased estimates of expected response to selection. Although we left the flies to hatch in a common environment, larval development took place inside the buds of two different host plants and in many different locations. Hence, it is possible that plastic responses to environmental effects from the early larval life stage could have contributed to our estimates of variational properties examined in this study. We partly addressed this by studying two replicates for each of the four host-locality combination so that each type of population experienced at least two different environments. The patterns found were largely consistent between replicates, suggesting that our main results are not driven by plastic responses to local weather or microclimates. Moreover, previous studies have shown that P and G are often aligned with the environmental variance–covariance (E, including plasticity) in a variety of traits (Noble et al., 2019) including size and shape traits in Diptera (Rohner & Berger, 2023; Saito et al., 2023). Furthermore, the strong correlation between P and divergence between the host races (Figs. 5 and 6) suggest that P indeed captures the general pattern of variational constraints, because we should not have been able to find such a strong correlation (R2 = 0.98) unless P is correlated with G and E. Taken together, the data suggest that our results are not likely to be strongly influenced by our focus on P, yet further work estimating G and E would be an important step towards uncovering how (cryptic) genetic variation, plasticity, recombination, or adaptation may have generated the patterns we demonstrated in this research.
Conclusions
We find evidence for reduced evolvability in response to past or current directional selection and potentially bottlenecks in population size resulting from the colonization of a new niche in females, but to a much lesser degree in males, of T. conura. The differences between sexes could suggest that selection on the ecologically important ovipositor is responsible for the observed reduction in evolvability. The host races also retained very similar structures of phenotypic variation despite the reduction in overall variation in females. Our study adds to growing evidence that evolvability is predictive of divergence between populations (Bolstad et al., 2014; Houle et al., 2017; McGlothlin et al., 2018; Opedal et al., 2023), and illustrates that evolvability is a dynamic entity that evolves when populations are exposed to novel environments.
Data availability
All data and code used in this study will be published on github (directory https://github.com/kalnil/tephritis) with a permanent DOI through Zenodo upon acceptance.
References
Armbruster, W. S., Pélabon, C., Bolstad, G. H., & Hansen, T. F. (2014). Integrated phenotypes: Understanding trait covariation in plants and animals. Philosophical Transactions of the Royal Society B-Biological Sciences, 369(1649), 20130245. https://doi.org/10.1098/rstb.2013.0245
Arnold, S. J., Bürger, R., Hohenlohe, P. A., Ajie, B. C., & Jones, A. G. (2008). Understanding the evolution and stability of the G-matrix. Evolution, 62(10), 2451–2461. https://doi.org/10.1111/j.1558-5646.2008.00472.x
Arnold, S. J., Pfrender, M. E., & Jones, A. G. (2001). The adaptive landscape as a conceptual bridge between micro- and macroevolution. Genetica, 112(1), 9–32. https://doi.org/10.1023/A:1013373907708
Assis, A. P. A., Patton, J. L., Hubbe, A., & Marroig, G. (2016). Directional selection effects on patterns of phenotypic (co)variation in wild populations. Proceedings of the Royal Society B-Biological Sciences, 283(1843), 20161615. https://doi.org/10.1098/rspb.2016.1615
Barrett, R. D. H., & Schluter, D. (2008). Adaptation from standing genetic variation. Trends in Ecology & Evolution, 23(1), 38–44. https://doi.org/10.1016/j.tree.2007.09.008
Berner, D., Stutz, W. E., & Bolnick, D. I. (2010). Foraging trait (co)variances in stickleback evolve deterministically and do not predict trajectories of adaptive diversification. Evolution, 64(8), 2265–2277. https://doi.org/10.1111/j.1558-5646.2010.00982.x
Björklund, M., Husby, A., & Gustafsson, L. (2013). Rapid and unpredictable changes of the G-matrix in a natural bird population over 25 years. Journal of Evolutionary Biology, 26(1), 1–13. https://doi.org/10.1111/jeb.12044
Blows, M., & Higgie, M. (2003). Genetic constraints on the evolution of mate recognition under natural selection. The American Naturalist, 161(2), 240–253. https://doi.org/10.1086/345783
Bolstad, G. H., Hansen, T. F., Pelabon, C., Falahati-Anbaran, M., Perez-Barrales, R., & Armbruster, W. S. (2014). Genetic constraints predict evolutionary divergence in Dalechampia blossoms. Philosophical Transactions of the Royal Society B-Biological Sciences, 369(1649), 20130255. https://doi.org/10.1098/rstb.2013.0255
Bradshaw, A. D., & McNeilly, T. (1991). Evolutionary response to global climatic change. Annals of Botany, 67(supp1), 5–14. https://doi.org/10.1093/oxfordjournals.aob.a088209
Cano, J. M., Laurila, A., Palo, J., & Merilä, J. (2004). Population differentiation in G matrix structure due to natural selection in Rana temporaria. Evolution, 58(9), 2013–2020. https://doi.org/10.1111/j.0014-3820.2004.tb00486.x
Cheverud, J. M. (1988). A comparison of genetic and phenotypic correlations. Evolution, 42(5), 958–968. https://doi.org/10.2307/2408911
Diegisser, T., Johannesen, J., & Seitz, A. (2006a). The role of geographic setting on the diversification process among Tephritis conura (Tephritidae) host races. Heredity, 96(5), 410–418. https://doi.org/10.1038/sj.hdy.6800821
Diegisser, T., Johannesen, J., & Seitz, A. (2008). Performance of host-races of the fruit fly, Tephritis conura on a derived host plant, the cabbage thistle Cirsium oleraceum: Implications for the original host shift. Journal of Insect Science, 8(1), 66. https://doi.org/10.1673/031.008.6601
Diegisser, T., Seitz, A., & Johannesen, J. (2006b). Phylogeographic patterns of host-race evolution in Tephritis conura (Diptera: Tephritidae). Molecular Ecology, 15(3), 681–694. https://doi.org/10.1111/j.1365-294X.2006.02792.x
Diegisser, T., Seitz, A., & Johannesen, J. (2007). Morphological adaptation in host races of Tephritis conura. Entomologia Experimentalis Et Applicata, 122(2), 155–164. https://doi.org/10.1111/j.1570-7458.2006.00501.x
Dochtermann, N. A., & Matocq, M. D. (2016). Speciation along a shared evolutionary trajectory. Current Zoology, 62(6), 507–511. https://doi.org/10.1093/cz/zow059
Doroszuk, A., Wojewodzic, M. W., Gort, G., & Kammenga, J. E. (2008). Rapid divergence of genetic variance-covariance matrix within a natural population. The American Naturalist, 171(3), 291–304. https://doi.org/10.1086/527478
Eroukhmanoff, F., & Svensson, E. I. (2011). Evolution and stability of the G-matrix during the colonization of a novel environment. Journal of Evolutionary Biology, 24(6), 1363–1373. https://doi.org/10.1111/j.1420-9101.2011.02270.x
Gompert, Z., Mandeville, E. G., & Buerkle, C. A. (2017). Analysis of population genomic data from hybrid zones. Annual Review of Ecology, Evolution, and Systematics, 48(1), 207–229. https://doi.org/10.1146/annurev-ecolsys-110316-022652
Guillaume, F., & Whitlock, M. C. (2007). Effects of migration on the genetic covariance matric. Evolution, 61(10), 2398–2409. https://doi.org/10.1111/j.1558-5646.2007.00193.x
Hadfield, J. D. (2010). MCMC methods for multi-response generalized linear mixed models: The MCMCglmm R package. Journal of Statistical Software, 33(2), 1–22. https://doi.org/10.18637/jss.v033.i02
Hansen, T. F., Armbruster, W. S., Carlson, M. L., & Pelabon, C. (2003). Evolvability and genetic constraint in Dalechampia blossoms: Genetic correlations and conditional evolvability. Journal of Experimental Zoology Part B-Molecular and Developmental Evolution, 296B(1), 23–39. https://doi.org/10.1002/jez.b.00014
Hansen, T. F., & Houle, D. (2008). Measuring and comparing evolvability and constraint in multivariate characters. Journal of Evolutionary Biology, 21(5), 1201–1219. https://doi.org/10.1111/j.1420-9101.2008.01573.x
Hansen, T. F., & Pelabon, C. (2021). Evolvability: A quantitative-genetics perspective. Annual Review of Ecology, Evolution, and Systematics, 52, 153–175. https://doi.org/10.1146/annurev-ecolsys-011121-021241
Hansen, T. F., Pélabon, C., & Houle, D. (2011). Heritability is not evolvability. Evolutionary Biology, 38(3), 258–277. https://doi.org/10.1007/s11692-011-9127-6
Henry, G. A., & Stinchcombe, J. R. (2022). G-matrix stability in clinally diverging populations of an annual weed. Evolution, 77(1), 49–62. https://doi.org/10.1093/evolut/qpac005
Houle, D. (1992). Comparing evolvability and variability of quantitative traits. Genetics, 130(1), 195–204. https://doi.org/10.1093/genetics/130.1.195
Houle, D., Bolstad, G. H., van der Linde, K., & Hansen, T. F. (2017). Mutation predicts 40 million years of fly wing evolution. Nature, 548(7668), 447–450. https://doi.org/10.1038/nature23473
Jones, A. G., Arnold, S. J., & Bürger, R. (2003). Stability of the G-matrix in a population experiencing pleiotropic mutation, stabilizing selection, and genetic drift. Evolution, 57(8), 1747–1760. https://doi.org/10.1111/j.0014-3820.2003.tb00583.x
Jones, A. G., Arnold, S. J., & Burger, R. (2004). Evolution and stability of the G-matrix on a landscape with a moving optimum. Evolution, 58(8), 1639–1654. https://doi.org/10.1111/j.0014-3820.2004.tb00450.x
Jones, A. G., Arnold, S. J., & Bürger, R. (2007). The mutation matrix and the evolution of evolvability. Evolution, 61(4), 727–745. https://doi.org/10.1111/j.1558-5646.2007.00071.x
Jones, A. G., Bürger, R., & Arnold, S. J. (2014). Epistasis and natural selection shape the mutational architecture of complex traits. Nature Communications, 5(1), 3709. https://doi.org/10.1038/ncomms4709
Klingenberg, C. P. (2008). Morphological integration and developmental modularity. Annual Review of Ecology, Evolution, and Systematics, 39(1), 115–132. https://doi.org/10.1146/annurev.ecolsys.37.091305.110054
Kohn, L. P., & Atchley, W. R. (1988). How similar are genetic correlations structures? Data from Mice and Rats. Evolution, 42(3), 467–481. https://doi.org/10.1111/j.1558-5646.1988.tb04153.x
Lande, R. (1979). Quantitative genetic analysis of multivariate evolution, applied to brain: Body size allometry. Evolution, 33(1), 402–416. https://doi.org/10.2307/2407630
Matlab (2017). 9.3.0.713579 (R2017b). Natick, Massachusetts: The MathWorks Inc.
McGlothlin, J. W., Kobiela, M. E., Wright, H. V., Kolbe, J. J., Losos, J. B., & III, E. D. B. (2022). Conservation and convergence of genetic architecture in the adaptive radiation of Anolis lizards. The American Naturalist, 200(5), E207–E220. https://doi.org/10.1086/721091
McGlothlin, J. W., Kobiela, M. E., Wright, H. V., Mahler, D. L., Kolbe, J. J., Losos, J. B., & Brodie, E. D., III. (2018). Adaptive radiation along a deeply conserved genetic line of least resistance in Anolis lizards. Evolution Letters, 2(4), 310–322. https://doi.org/10.1002/evl3.72
Melo, D., & Marroig, G. (2015). Directional selection can drive the evolution of modularity in complex traits. Proceedings of the National Academy of Sciences, 112(2), 470–475. https://doi.org/10.1073/pnas.1322632112
Milocco, L., & Salazar-Ciudad, I. (2022). Evolution of the G matrix under nonlinear genotype-phenotype maps. The American Naturalist, 199(3), 420–435. https://doi.org/10.1086/717814
Nei, M., Maruyama, T., & Chakraborty, R. (1975). The bottleneck effect and genetic variability in populations. Evolution, 29(1), 1–10. https://doi.org/10.1111/j.1558-5646.1975.tb00807.x
Nilsson, K. J., Ortega, J., Friberg, M., & Runemark, A. (2022). Non-parallel morphological divergence following colonization of a new host plant. Evolutionary Ecology, 36, 859–877. https://doi.org/10.1007/s10682-022-10189-2
Noble, D. W. A., Radersma, R., & Uller, T. (2019). Plastic responses to novel environments are biased towards phenotype dimensions with high additive genetic variation. Proceedings of the National Academy of Sciences, 116(27), 13452–13461. https://doi.org/10.1073/pnas.1821066116
Noble, L. M., Chelo, I., Guzella, T., Afonso, B., Riccardi, D. D., Ammerman, P., Dayarian, A., Carvalho, S., Crist, A., Pino-Querido, A., Shraiman, B., Rockman, M. V., & Teotónio, H. (2017). Polygenicity and epistasis underlie fitness-proximal traits in the Caenorhabditis elegans multiparental experimental evolution (CeMEE) panel. Genetics, 207(4), 1663–1685. https://doi.org/10.1534/genetics.117.300406
Opedal, Ø. H., Armbruster, W. S., Hansen, T. F., Holstad, A., Pélabon, C., Andersson, S., Campbell, D. R., Caruso, C. M., Delph, L. F., Eckert, C. G., Lankinen, Å., Walter, G. M., Ågren, J., & Bolstad, G. H. (2023). Evolvability and trait function predict phenotypic divergence of plant populations. Proceedings of the National Academy of Sciences, 120(1), e2203228120. https://doi.org/10.1073/pnas.2203228120
Opedal, Ø. H., Hildesheim, L. S., & Armbruster, W. S. (2022). Evolvability and constraint in the evolution of three-dimensional flower morphology. American Journal of Botany, 109(11), 1906–1917. https://doi.org/10.1002/ajb2.16092
Paaby, A. B., & Rockman, M. V. (2014). Cryptic genetic variation: Evolution’s hidden substrate. Nature Reviews Genetics, 15(4), 247–258. https://doi.org/10.1038/nrg3688
Pavlicev, M., Cheverud, J. M., & Wagner, G. P. (2011). Evolution of adaptive phenotypic variation patterns by direct selection for evolvability. Proceedings of the Royal Society B-Biological Sciences, 278(1713), 1903–1912. https://doi.org/10.1098/rspb.2010.2113
Pélabon, C., Firmat, C., Bolstad, G. H., Voje, K. L., Houle, D., Cassara, J., Rouzic, A. L., & Hansen, T. F. (2014). Evolution of morphological allometry. Annals of the New York Academy of Sciences, 1320(1), 58–75. https://doi.org/10.1111/nyas.12470
Pélabon, C., Hansen, T. F., Carter, A. J. R., & Houle, D. (2010). Evolution of variation and variability under fluctuating, stabilizing, and disruptive selection. Evolution, 64(7), 1912–1925. https://doi.org/10.1111/j.1558-5646.2010.00979.x
Penna, A., Melo, D., Bernardi, S., Oyarzabal, M. I., & Marroig, G. (2017). The evolution of phenotypic integration: How directional selection reshapes covariation in mice. Evolution, 71(10), 2370–2380. https://doi.org/10.1111/evo.13304
Porto, A., de Oliveira, F. B., Shirai, L. T., De Conto, V., & Marroig, G. (2009). The evolution of modularity in the mammalian skull I: Morphological integration patterns and magnitudes. Evolutionary Biology, 36(1), 118–135. https://doi.org/10.1007/s11692-008-9038-3
Puentes, A., Granath, G., & Ågren, J. (2016). Similarity in G matrix structure among natural populations of Arabidopsis lyrata. Evolution, 70(10), 2370–2386. https://doi.org/10.1111/evo.13034
R Core Team (2019). R: A language and environment for statistical computing. Vienna, Austria. : R Foundation for Statistical Computing.
Revell, L. J., Mahler, D. L., Sweeney, J. R., Sobotka, M., Fancher, V. E., & Losos, J. B. (2010). Nonlinear selection and the evolution of variances and covariances for continuous characters in an anole. Journal of Evolutionary Biology, 23(2), 407–421. https://doi.org/10.1111/j.1420-9101.2009.01911.x
Roff, D. A., & Mousseau, T. (2005). The evolution of the phenotypic covariance matrix: Evidence for selection and drift in Melanoplus. Journal of Evolutionary Biology, 18(4), 1104–1114. https://doi.org/10.1111/j.1420-9101.2005.00862.x
Rohner, P. T., & Berger, D. (2023). Developmental bias predicts 60 million years of wing shape evolution. Proceedings of the National Academy of Sciences, 120(19), e2211210120. https://doi.org/10.1073/pnas.2211210120
Romstock-Volkl, M. (1997). Host race formation in Tephritis conura: determinants from three trophic levels. In K. Dettner, G. Bauer, & W. Völkl (Eds.), Vertical Food Web Interactions (pp. 21–38). Springer.
Saito, K., Tsuboi, M., & Takahashi, Y. (2023). Developmental noise and phenotypic plasticity are correlated in Drosophila simulans. bioRxiv, 2023.2007.2028.550919. https://doi.org/10.1101/2023.07.28.550919
Schluter, D. (1996). Adaptive radiation along genetic lines of least resistance. Evolution, 50(5), 1766–1774. https://doi.org/10.1111/j.1558-5646.1996.tb03563.x
Schluter, D. (2000). The ecology of adaptive radiation. Oxford University Press.
Sodini, S. M., Kemper, K. E., Wray, N. R., & Trzaskowski, M. (2018). Comparison of genotypic and phenotypic correlations: Cheverud’s conjecture in humans. Genetics, 209(3), 941–948. https://doi.org/10.1534/genetics.117.300630
Steppan, S. J., Phillips, P. C., & Houle, D. (2002). Comparative quantitative genetics: Evolution of the G matrix. Trends in Ecology & Evolution, 17(7), 320–327. https://doi.org/10.1016/s0169-5347(02)02505-3
Steward, R. A., Nilsson, K. J., Giménez, J. O., Nolen, Z. J., Yan, C., Huang, Y., López, J. A., & Runemark, A. (2023). The genomic landscape of adaptation to a new host plant. bioRxiv, 2023.2004.2017.537225. https://doi.org/10.1101/2023.04.17.537225
Tufto, J. (2000). The evolution of plasticity and nonplastic spatial and temporal adaptations in the presence of imperfect environmental cues. The American Naturalist, 156(2), 121–130. https://doi.org/10.1086/303381
Voje, K. L., Grabowski, M., Holstad, A., Porto, A., Tsuboi, M., & Bolstad, G. H. (2023). Does lack of evolvability constrain adaptation? If so, on what timescales? In T. F. Hansen, D. Houle, M. Pavličev, & C. Pélabon (Eds.), Evolvability: A unifying concept in evolutionary biology? The MIT Press.
Wagner, G. P., & Altenberg, L. (1996). Perspective: Complex adaptations and the evolution of evolvability. Evolution, 50(3), 967–976. https://doi.org/10.1111/j.1558-5646.1996.tb02339.x
Wagner, G. P., Pavlicev, M., & Cheverud, J. M. (2007). The road to modularity. Nature Reviews Genetics, 8(12), 921–931. https://doi.org/10.1038/nrg2267
Walsh, B., & Blows, M. W. (2009). Abundant genetic variation plus strong selection = multivariate genetic constraints: A geometric view of adaptation. Annual Review of Ecology, Evolution, and Systematics, 40, 41–59. https://doi.org/10.1146/annurev.ecolsys.110308.120232
Walter, G. M., Aguirre, J. D., Blows, M. W., & Ortiz-Barrientos, D. (2018). Evolution of genetic variance during adaptive radiation. The American Naturalist, 191(4), E108–E128. https://doi.org/10.1086/696123
Whitlock, M. C., Phillips, P. C., & Fowler, K. (2002). Persistence of changes in the genetic covariance matrix after a bottleneck. Evolution, 56(10), 1968–1975. https://doi.org/10.1111/j.0014-3820.2002.tb00122.x
Willis, J. H., Coyne, J. A., & Kirkpatrick, M. (1991). Can one predict the evolution of quantitative characters without genetics. Evolution, 45(2), 441–444. https://doi.org/10.2307/2409678
Wyman, M. J., & Rowe, L. (2014). Male bias in distributions of additive genetic, residual, and phenotypic variances of shared traits. The American Naturalist, 184(3), 326–337. https://doi.org/10.1086/677310
Acknowledgements
We thank two anonymous reviewers for insightful comments, Mikkel Brydegaard for providing Matlab code for quantifying wing melanization and Jesús Ortega, Jodie Lilley, Emma Kärrnäs, and Mathilde Schnuriger for help in the field and in the lab, and Geir H. Bolstad for advice on evolvability analyses. This study was funded by a Wenner-Gren Fellowship, a Crafoord grant and a Swedish Research Council grant to A. R. Ø.H.O. acknowledge support from the Swedish Research Council.
Funding
Open access funding provided by Lund University. Vetenskapsrådet, 2021-04777, 2018-04537, Crafoordska Stiftelsen, 20190787.
Author information
Authors and Affiliations
Contributions
A. R. conceived of and designed the study. K. J. N. and A. R. performed field work, K. J. N. eclosed the flies and quantified traits, K.J.N. performed the statistical analysis with support from M.T. and Ø.H.O., K. J. N. wrote the first draft of the manuscript, and all authors contributed equally to the writing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nilsson, K.J., Tsuboi, M., Opedal, Ø.H. et al. Colonization of a Novel Host Plant Reduces Phenotypic Variation. Evol Biol 51, 269–282 (2024). https://doi.org/10.1007/s11692-024-09634-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11692-024-09634-7