Abstract
Glutathione-S-transferase (GST, EC2.5.1.18) multifunctional protease is important for detoxification, defense against biotic and abiotic stresses, and secondary metabolic material transport for plant growth and development. In this study, 71 members of the BpGST family were identified from the entire Betula platyphylla Suk. genome. Most of the members encode proteins with amino acid lengths ranging from 101 to 875 and were localized to the cytoplasm by a prediction. BpGSTs can be divided into seven subfamilies, with a majority of birch U and F subfamily members according to gene structure, conserved motifs and evolutionary analysis. GST family genes showed collinearity with 22 genes in Oryza sativa L., and three genes in Arabidopsis thaliana; promoter cis-acting elements predicted that the GST gene family is functional in growth, hormone regulation, and abiotic stress response. Most members of the F subfamily of GST (BpGSTFs) were expressed in roots, stems, leaves, and petioles, with the most expression observed in leaves. On the basis of the expression profiles of F subfamily genes (BpGSTF1 to BpGSTF13) during salt, mannitol and ABA stress, BpGSTF proteins seem to have multiple functions depending on the type of abiotic stress; for instance, BpGSTs may function at different times during abiotic stress. This study enhances understanding of the GST gene family and provides a basis for further exploration of their function in birch.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glutathione transferases, also known as glutathione S-transferases (GSTs), are ubiquitous in animals, plants, and microorganisms. They are a group of superfamily enzymes encoded by multiple genes, performing various functions, including promoting metabolism, and elimination of harmful xenobiotics or oxidative products through glutathione binding to these substances (Nutricati et al. 2006). For example, GSTs facilitate the covalent binding of reduced glutathione (GSH) with hydrophobic and electrophilic substrates, forming conjugates that are sequestered in vacuoles or transferred to the apoplast, thereby detoxifying both endogenous and exogenous harmful substances, such as herbicides, and aiding in tolerance to abiotic stresses (Öztetik 2008; Cicero et al. 2015). This detoxification process involves three stages: transformation, conjugation, and compartmentalization (Nianiou-Obeidat et al. 2017). Soluble GST is mainly distributed in the cytoplasm, with some present in chloroplasts and microsomes, and a minor amount in the nucleus and extracellular bodies (Dixon et al. 2002). In addition, a typical GST has two binding sites: the N-terminal GSH binding site (G-site; GST-N) and the adjacent electron substrate binding site (H-site; GST-C), primarily formed by the C-terminal. The GST-N is well- conserved, likely due to its role in binding GSH, while the GST-C is variable, possibly because of its ability to combine with multiple substances (Edwards and Dixon 2005; Sylvestre-Gonon et al. 2019).
Based on the homology of plant proteins and gene structure characteristics, the GST family is divided into eight subfamilies: F (Phi), U (Tau), T (Theta), Z (Zeta), L (Lambda), DHAR, EF1Bγ, and TCHQD (Jain et al. 2010). The F and U subfamilies, unique to plants, have the largest number of members and the greatest abundance compared to other subfamilies. They have a broader substrate spectrum, allowing them to bind to various harmful xenobiotics, such as pesticides and herbicides, to protect plants from toxicity (Munyampundu et al. 2016). Theta-type GSTs possess glutathione peroxidase (GPOX) activity, reducing H2O2 produced by plants under oxidative stress. Zeta-type GSTs with maleoylacetone isomerase activity are involved in the degradation of tyrosine in organisms, indicating common metabolism of compounds in living cells (Board et al. 1997). DHAR-type and Lambda-type GSTs are grouped as mercaptotransferases, mainly functioning as peroxidases. DHAR type members catalyze ascorbate synthesis under stress conditions such as excess light and drought, enhancing plant antioxidant capacity (Lallement et al. 2014). The TCHQD protein may have a serine residue at its active site (Mohsenzadeh et al. 2011). However, the nature of the catalytic residue in the EF1Bγ subfamily remains unclear. It has been reported that members of the EF1Bγ subfamily play a role in the oxidative stress response pathway (Olarewaju et al. 2004).
GST affects growth and development by participating in plant physiology and metabolism, increases resistance to adversity, and is involved in cellular signaling and other pathways. In flax (Linum usitatissimum L.), GST is important for detoxification of reactive oxygen species (ROS) and cell wall modification (Dmitriev et al 2016). In walnut species (Juglans regia L.), JrGSTTau1 plays a positive role in osmotic tolerance and is modulated by upstream regulators (Yang et al. 2019). The tobacco GST gene expression was significantly up-regulated by various stresses, and transgenic tobacco plants expressing Tau-class SbGST genes exhibited enhanced tolerance to abiotic stress (Jha et al. 2011). MaGSTs play a pivotal role in both development and abiotic stress responses in banana (Wang et al. 2013). GST has been reported as a key component in the metabolism of anthocyanins, flavonols, proanthocyanidin, cinnamic acid (Wei et al. 2019; Liu et al. 2019a, b), methyl jasmonate (Zhang et al. 2022a, b), salicylic acid (Gullner et al. 2018), auxin (Wu et al. 2023), and ethylene (Zhang et al. 2022a, b). Previous studies showed that GST-mediated transport is key in anthocyanin accumulation in various plant species, including sweet cherry (Qi et al. 2022), maize (Marrs 1996), petunia (Mueller et al. 2000), pear (Li et al. 2022), Arabidopsis (Yamazaki et al. 2008; Sun et al. 2012), grape (Conn et al. 2008), litchi (Hu et al. 2016), apple (Jiang et al. 2019), kiwifruit (Liu et al. 2019a, b), and peach (Zhao et al. 2020).
Betula platyphylla Suk., a forest species with significant economic importance, is widespread in northern China (Lyu et al. 2020). It is a vital timber resource for the construction and furniture industries and also plays an essential role in ecosystem restoration as a pioneering species in forests recovering from fires. The species prefers sunlight, demonstrates cold resistance, and thrives in acidic and moist soils, highlighting its adaptability and ecological value (Jing et al. 2020; Geng et al. 2022).
Understanding the function of the GST gene family is instrumental in the study of molecular mechanisms of development and stress tolerance of plants. However, the genome-wide GST gene family has not been comprehensively evaluated in birch. In this study, 71 BpGST genes were identified, and their basic biological information analyzed, which included gene structure, evolutionary relationship, conserved motif, gene collinearity and promoter cis-acting elements. Combined with the analysis of the expression pattern of Phi BpGSTFs in different organs and under salt, mannitol, and ABA abiotic stresses, this study provides reference and the basis for exploring the function of GST family genes.
Materials and methods
Identification and sequence analysis of the GST gene birch family
Protein sequences of the Arabidopsis thaliana GST gene family were obtained (http://www.arabidopsis.org/). A local blast database was constructed using white birch proteome sequences (Chen et al. 2021), with A. thaliana GST protein sequence as the seed for Blastp search with the E value cutoff of 1e − 10. The hidden Markov model of GST was downloaded from the Pfam database (http://pfam.xfam.org/) for PF02798, PF00043, PF13410, and PF13417. Searches were conducted using HMMER 3.0 software (http://hmmer.janelia.org) based on E value ≤ 1e − 5. Repetitive sequences from both methods were removed. Using the CDD (https://www.ncbi.nlm.nih.gov/cdd) for domain validation, sequences without GST were deleted, resulting in the final identification of candidate genes.
Analysis of physical and chemical properties
The ExPASy online software ProtParam (https://web.expasy.org/protparam/) was used to predict the physical and chemical properties of the protein primary structure, including the isoelectric point (pI), amino acid number, and relative molecular mass (https://wolfpsort.hgc.jp/).
Phylogenetic tree construction
The protein sequences BpGSTs and AtGSTs were analyzed using MEGA 7 software with ClustalX multiple sequence alignment. An evolutionary tree was constructed with the NJ method (execution parameters: Poisson correction, pairwise deletion, and bootstrap 1000 duplications). The birch GST gene family members were renamed according to the evolutionary tree subfamily.
Gene structure analysis
Gene structure maps were generated using the GFF file annotated by B. platyphylla genome in TBtools software (Chen et al. 2020).
Conservative motif analysis
Motifs with high similarity in the family genes were detected by MEME, setting the width of conserved sites to ≥ 5 and ≤ 50, and the maximum number of conserved sequences to 20. The conservative motif map of the gene family was drawn by TBtools.
Chromosomal localization analysis
Using genome annotation information, the gene location information and structure were visualized using TBtools software.
Analysis of cis-acting elements of promoter
Based on genome annotation information, 2000 bp upstream of the gene was analyzed using PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html) for promoter cis-acting element analysis. The results were visualized by TBtools.
Plant materials
Birch in vitro seedlings were cultured in a woody plant medium containing 20 g L−1 sucrose and 7.5–8 g L−1 agar. The climate chamber was set at 24 ± 2 °C light for 16 h using a 46 µmol m−2 s−1 cold white, fluorescent lamp, 8 h of darkness, and a relative humidity of 65% to 75%. Four-week-old seedlings, approximately 2.5 cm high, were transferred to a medium containing 150 mM NaCl, 200 mM mannitol and 100 µM ABA for abiotic stress. Samples were collected on 1, 2, 4, and 7 days after treatment, and the untreated samples are to be named ‘day 0’, serving as the control group. Each group was replicated three times, frozen in liquid nitrogen and stored at − 80 °C.
RNA extraction and cDNA synthesis
Total RNA was extracted from the samples by the CTAB method (Gambino et al. 2008) and RNA integrity was confirmed by electrophoresis on a 1% agarose gel. cDNA was synthesized with the ReverTra Ace® qPCR RT Master Mix with gDNA Remover kit (TOYOBO, Shanghai) according to the manufacturer’s instructions.
Real-time fluorescence quantitative PCR analysis
The synthetic cDNA was diluted tenfold as a template for relative quantitative PCR using a Roche LightCycler 480 II fluorescence quantitative PCR instrument. Reaction conditions included a 95 °C reaction for 30 s; 95 °C for 10 s, 60 °C for 10 s, 72 °C for 30 s for 45 cycles, followed by 95 °C for 10 s, 60 °C for 1 min, and 97 °C for 15 s to confirm PCR product specificity with melting curves. All samples were replicated three times and three technical replicates. Birch 18S rRNA (18S) was used as an internal control. Primer design was based on the online program Primer-Blast (Primer designing tool, https://www.ncbi.nlm.nih.gov/tools/primer-blast/) with all analyzed gene primers listed in Table S2.
For RT-qPCR, TOYOBO THUNDERBIRD SYBR qPCR Mix Without Rox (TOYOBO, Shanghai) was used, totaling 20 µL of 10 µL 2 × SYBR qMix, forward and reverse primers 0.6 µL (10 µM), 2 µL cDNA, and 4.8 µL ddH2O. The relative expression level of the target gene was determined by the 2−ΔΔCt method (Yaish et al. 2010).
Results
Identification and analysis of birch GST gene family
Based on BLASTp and HMM search, 71 GST family members were identified in the birch genome after removing the redundant transcripts. Based on protein sequence phylogenetic subfamily classification and chromosome location information, they were named BpGSTU1-BpGSTU41, BpGSTF1-BpGSTF13, BpDHAR1-BPDHAR3, BpGSTL1-BpGSTL3, BpGSTT1, BpGSTZ1-BpGSTZ5, and BpTCHQD1-BpTCHQD5 (Table S1).
The 71 GST gene family members encoded proteins varied in size from 101 to 875 amino acids, with molecular weights ranging from 94.86 to 116.38 kDa. The BpDHAR3 had the highest molecular weight and the highest number of amino acids (875 amino acid residues). Conversely, the shortest BpGSTU16 protein contained only 101 amino acid residues. The theoretical isoelectric point (pI) was between 4.43 and 9.82. Fifty-two BpGST members had a pI < 7, indicating these proteins may possess acidic properties; the remaining nineteen BpGST members had a pI > 7, suggesting these proteins could have alkaline characteristics. The instability index ranged between 21.78 and 67.27, with fourty-six BpGST proteins with an instability index < 40. In contrast, the other twenty-five BpGST proteins had a high instability index, indicating that they could be unstable proteins. The aliphatic index of these proteins ranged between 73.87 and 112.86. The GRAVY (Grand Average of Hydropathy) index, which estimates a protein’s hydrophobic or hydrophilic character, indicated that most BpGST proteins, except for BpGSTU23, BpGSTU25, BpGSTF11, and BpGSTZ2, were hydrophilic with negative values. Subcellular prediction results indicated that most BpGSTs were localized in the cytoplasm, followed by the nucleus, mitochondria, peroxisome, and the cytoskeleton (Table S1).
Phylogenetic analysis of GST gene family
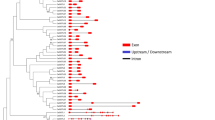
The evolutionary classification of 53 AtGSTs and 71 BpGSTs based on their amino acid sequences showed that the 71 BpGST protein members in B. platyphylla could be divided into seven subfamilies, including Tau, Zeta, Lambda, Theta, DHAR, TCHQD, and Phi subfamily. Similar to the classification in Arabidopsis, more than half of BpGST members belonged to the U and F subfamilies. Among the 71 birch family members, the U subfamily had the most members with 41 (Fig. 1); F subfamily followed with 13 members. The subfamilies, DHAR and L, consisted of 3 members each. Subfamily T only had one member, BpGSTZ1, and subfamily Z and TCHQD both had 5 respectively (Fig. 1). Notably, the Phi, Theta, and Theta subfamilies were clustered on the same evolutionary clade, indicating a closer relationship among these subfamilies.
Analysis of the structure and motif of GST family genes in birch
For the 71 GST gene structures in birch (Fig. 2), the number of introns did not exceed 12. Twenty-six BpGSTUs, including BpGSTU2, BpGSTU3, BpGSTU7, and others, contained a single intron. Additionally, BpGSTU16, BpGSTF8 and BpGSTF11 lacked introns entirely. The U subfamily members generally shared a consistent gene structure, with most containing 1–3 introns and 2–4 exons. An exception within this subfamily was BpGTU6, which displayed a unique structure of four introns and five exons. Within the F subfamily, BpGSTF3, BpGSTF6, BpGSTF9, BpGSTF12, and BpGSTF13 were distinguished by possessing either 5' or 3' untranslated regions (UTRs), a feature absent in the other eight members. The gene structures within the L, Z, TCHQD, and T subfamilies exhibited considerable variation, highlighting the diversity and complexity of the GST gene structures in birch. Among these, members of the L subfamily were notable for having more than 8 introns and exons, indicating a significant degree of conservation. BpGSTT1, the unique member of the T subfamily, featured 6 introns and 7 exons. Additionally, BpDHAR3 of the DHAR subfamily possessed the longest CDS with 2628 bp, while BpGSTU14 was remarkable for its long introns within the entire GST family. Notably, there existed 12 introns and 13 exons in BpGSTF10, further emphasizing the intricate genetic framework present within these subfamilies.
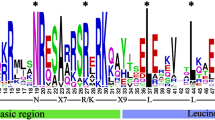
Twenty conserved motifs in the BpGST gene family were identified using the MEME online tool (Fig. 3). The distributions of various motifs in GST proteins are shown in Fig S1. Motif 3 was present in 61 GST proteins, indicating that it can be used as a marker to recognize BpGSTs. Additionally, motif 7 was found in 21 GST proteins. Within the T subfamily, with the exception of BpGSTU16 and BpGSTU39, all members harbored the conserved motif 1, suggesting that this motif may play a significant role in the subfamily. Moreover, the T subfamily was noted for exhibiting the greatest diversity of motif types, highlighting its complex genetic architecture. Interestingly, each member of the L subfamily contained motifs 3, 7, and 15, suggesting these shared motifs might provide insights into the evolutionary divergence of the L subfamily from other gene groups. The arrangement of conserved motifs in BpGSTU5-BpGSTU15, BpGSTU17-BpGSTU30, and BpGSTU33-BpGSTU38 showed remarkable similarity, particularly with motifs 6, 3, and 1 being in comparable positions. This consistency in motif arrangement suggests that these genes may perform similar biological functions, attributable to their shared motifs.
Chromosomal localization and gene collinearity analysis of BpGSTs genes in birch
To understand the distribution on chromosomes, the chromosomal localization of BpGSTs genes was studied based on birch genomic information. The results suggest that 66 BpGSTs genes were localized on 14 chromosomes (Fig. 4). Chromosome 5 harbored the largest number of BpGSTs genes (15), followed by chromosome 6 with 10 BpGSTs genes, while chromosomes 2, 7, and 10 had only one BpGST gene each. Together, these findings indicated that the BpGSTs distribution was not relevant for chromosome size.
According to tandem replication criteria (Qiao et al. 2015), it usually results in gene clusters but segmental duplication might cause family members to become separated. Notably, multiple genes formed dense gene clusters such as BpGSTF2 and BpGSTF3 as well as BpGSTU9 and BpGSTU10 on chromosome 5, and BpGSTU16, BpGSTU17 and BpGSTU18 on chromosome 6. In total, seventeen pairs of tandem duplicated genes were detected in the BpGST gene family, most of which were in the Tau, Phi, Zeta, and TCHQD subfamilies (Fig. 4). Furthermore, five pairs of segmental duplicated genes were found and connected by red curves (Fig. 5). For example, BpGSTF2 and BpGSTF5 on chromosome 5 were highly collinear with genes BpGSTF8 and BpGST13 on chromosomes 8 and 13, respectively. BpGSTU5 on chromosome 4 was collinear with BpGSTU30 on chromosome 10. BpGSTL2 on chromosome 6 and BpGSTL3 on chromosome 14 were collinear, as were BpGSTF26 on chromosome 8 and BpGSTF29 on chromosome 9. Additionally, interspecific collinear relationships between birch GST family members and model plants Arabidopsis and monocotyledon Oryza sativa were analyzed. It was found that the BpGST gene in birch was more closely related to Arabidopsis thaliana than to monocotyledon rice (Fig. 6).
Analysis of 2 kb cis-acting elements upstream of BpGST family genes
The upstream 2 kb base sequences of 71 BpGST family members were extracted and cis-acting elements predicted using PlantCARE online software. In addition to core promoter elements such as TATA-box and CAAT-box, the majority of regulatory elements are involved in light response, as well as abscisic acid acting elements (ABRE), stress response elements (STRE), anaerobic inducible response elements (ARE), and MYB binding sites. MYB binding sites also participate in various processes such as light response, drought induction, and flavonoid biosynthesis (Fig. 7). The presence of diverse regulatory elements related to abiotic stress response suggests that BpGSTs may play a role in plant stress resistance.
Gene expression patterns of BpGSTFs in different tissues
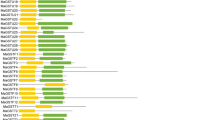
To clarify the role of Phi BpGST genes in birch growth and development, the expression patterns of 13 BpGSTF genes in different birch tissues were analyzed. This study revealed that BpGSTFs exhibit varied expression patterns in different tissues (Fig. 8). Most BpGSTFs genes were expressed in all 4 tissues examined, with 9 BpGSTF genes (BpGSTF1, BpGSTF2, BpGSTF3, BpGSTF5, BpGSTF6, BpGSTF10, BpGSTF11, BpGSTF12 and BpGSTF13) showing high relative expression in leaves. 9 genes (BpGSTF9, BpGSTF11, BpGSTF12 and BpGSTF13) displayed specific expression patterns in one or some tissues; for example, all genes have low expression in stems, while BpGSTF11 exhibited similar expression in leaves and petioles, and BpGSTF5 and BpGSTF11 had similar expression patterns. BpGSTF1, BpGSTF2, and BpGSTF3 shared similar high expression patterns in leaves. BpGSTF4’s expression in the petiole was about 70 times higher than in roots and leaves, suggesting a possible role in petiole development. The expression patterns of BpGSTF7 and BpGSTF8 were similar, with higher expression in roots and petioles. BpGSTF10 showed similar expression in stems and petioles, approximately four times higher in leaves. The expression of BpGSTF6 varied significantly in roots, stems, leaves, and petioles, indicating diverse functions of this gene in birch.
Analysis of BpGSTFs gene expression patterns in four tissues (root, stem, leaf and petiole). The level in the root is expressed as 1. Error bars represent the standard deviation of the mean using three technical replicates. Data were analyzed using one-way variance (ANOVA). Using the LSD method for multiple comparisons, different letters represent significant differences (P < 0.05)
Expression pattern analysis of BpGSTF genes in response to salt and osmotic stress
Considering the importance of F subfamily genes to abiotic stresses, the expression patterns of BpGSTFs under salt and osmotic stress were analyzed. The results reveal that each gene had significant differences in response and the time of response varied as well. The expression patterns of BpGSTF gene family members under NaCl stress indicated that BpGSTF1, BpGSTF5, and BpGSTF11 showed a trend of down-regulation (Fig. 9). The expression of BpGSTF2, BpGSTF3, and BpGSTF9 genes peaked on the second day after salt treatment, followed by a gradual decrease. BpGSTF6, BpGSTF10, and BpGSTF13 exhibited similar expression patterns, with peak expression on the first day. Notably, BpGSTF13 was up-regulated most significantly on the second day after salt treatment, about 15 times higher than controls, suggesting that the gene was strong inducted by NaCl. The expression of BpGSTF4, BpGSTF7, and BpGSTF8 initially increased, then dropped to their lowest value on the fourth day, which was 7.3, 1.7, and 7.8 times lower, respectively, than their highest values. On the seventh day of salt treatment, the expression of these three genes slightly increased. BpGSTF1 displayed a unique expression pattern, being significantly induced after one day of salt stress, reaching its highest expression, then decreasing the following day and increasing again on the fourth day, but with little variation from the highest value, followed by a subsequent decrease.
To examine the expression of BpGSTFs under osmotic stress, stem segments with terminal buds of tissue culture seedlings were cut and cultured in rooting medium containing 200 mM mannitol WPM for 0, 1, 2, 4, and 7 d. Total RNAs were extracted for quantitative analysis (Fig. 10). The results show that, after mannitol treatment, the expression of BpGSTF1, BpGSTF5, BpGSTF6, BpGSTF7, BpGSTF8, and BpGSTF10 initially decreased and then increased. The expression patterns of BpGSTF5, BpGSTF6, BpGSTF7, and BpGSTF8 were similar, with relative expression levels two days after treatment being significantly low, only 2.3%, 7.1%, 5.0%, and 3.2% of the control (day zero), respectively, and then gradually decreased. The expression patterns of other family members BpGSTF2, BpGSTF3, BpGSTF4, BpGSTF9, BpGSTF11, and BpGSTF13 initially increased and then decreased. BpGSTF4, BpGSTF9, and BpGSTF11 were significantly induced and reached their highest value on the first day, increasing by 1.2, 7.5, and 1.6 times, respectively. They showed a down-regulation trend on the second day, up-regulated again on the fourth day, and then nearly dropped to zero on the seventh day. The expression of BpGSTF13 peaked 2 days after the treatment, about 20 times that of the control, and then decreased.
Expression pattern analysis of BpGSTF genes in response to ABA
The RT-qPCR results demonstrate that the expression of BpGSTFs were induced by ABA. Post-ABA treatment, the expression patterns were primarily categorized into three types (Fig. 11). Firstly, the expression of BpGSTF1, BpGSTF10, BpGSTF6, BpGSTF7, and BpGSTF9 were significantly up-regulated in the initial 2 days of ABA treatment, decreased on the 4th day, and then showed a temporary up-regulation in the last time period. Secondly, the expression levels of BpGSTF5, BpGSTF8, and BpGSTF13 peaked on the 1st day post-treatment, registering 5.6, 7.1, and 4.2 times their levels on day 0, respectively, followed by a brief increase and then a decrease on the 4th day. It is hypothesized that these genes may share similar functions in response to ABA treatment. Thirdly, BpGSTF3 and BpGSTF12 exhibited an initial up-regulation followed by a decline; in contrast, BpGSTF11 consistently trended downward. The expression patterns of BpGSTF2 and BpGSTF4 were distinct. For BpGSTF2, it increased after 1 day of stress, decreased after 2 days, then rose again between 2 and 4 days, before gradually declining. BpGSTF4 showed a continuous downward trend in the initial 2 days of ABA treatment, was significantly induced on the fourth day, and then declined.
Discussion
The GST gene family is extensively found across various plant species, playing crucial roles in numerous growth and development processes and exerting regulatory functions (Shehu et al. 2018). To date, significant numbers of GST gene family members have been identified in a variety of plants. For example, there are 55, 49, 90, 23, 42, 79, 61, 54, and 97 GST members in Arabidopsis thaliana (Sappl et al. 2009), Gossypium arboreu (Dong et al. 2016), Solanum lycopersicum (Islam et al. 2017), Malus domestica (Jiang et al. 2019), Zea mays L. (McGonigle et al. 2000), O. sativa L. (Jain et al. 2010), Citrus sinensis (Licciardello et al. 2014), Prunus persica (Zhao et al. 2017) and Actinidia Lindl (Liu et al. 2019a, b). However, there have been no related reports on the identification and functional study of GST gene family members in B. platyphylla. Therefore, we identified and characterized the GST genes in birch. This research broadens our insight into the diversity of the GST gene family and also contributes significantly to clarifying their role in growth and stress tolerance in birch.
Based on BLAST and HMM search, 71 members of the GST gene family were identified in the B. platyphylla genome (Table S1). Physicochemical analysis revealed that 25 were classified as unstable proteins. Apart from BpGSTU23, BpGSTU25, BpGSTF11, and BpGSTZ2, which exhibited hydrophobic properties, the remaining were hydrophilic. This distinction could influence their functionality within the cell. Phylogenetic analysis divided these BpGST genes into U, F, DHAR, L, T, Z, and TCHQD subfamilies. Notably, compared to Arabidopsis, Triticeae (Hao et al. 2021) and Ficus carica L. (Liu et al. 2023), the EF1Bγ subfamily was absent in B. platyphylla, a difference possibly resulting from gene loss over the course of evolution. The loss of the EF1Bγ subfamily may be related to gene functional substitution and adaptation to environmental pressures during evolution (Jia et al. 2023), which should be further studied. In addition, the L and U families clustered together, while Z, T, and F families, and DHAR and TCHQD families formed separate branches.
Exon–intron structure is crucial in gene evolution (Xu et al. 2012). In birch, BpGST genes within the same subfamily exhibited similar exon–intron structures, particularly in the T and L subfamilies. A previous study highlighted that, compact genetic structures with fewer introns, enable timely stress responses (Jeffares et al. 2008), suggesting that T and DHAR members with fewer introns might rapidly respond to stress. In addition, most BpGST members of the same subfamily had conserved motifs. These results reveal that BpGST members in same subfamily are evolutionarily conserved across distinct phylogenetic groups, indicating they might perform similar biological functions.
Collinearity analysis involving birch, Arabidopsis, and rice revealed 22 relationships between the GST family and Arabidopsis, and three with rice, suggesting higher homology of the GST gene family with the Arabidopsis GST gene family. Chromosomal analysis indicated that the widespread distribution of BpGST in birch contributes to the diversity and complexity of this gene family, potentially key to their roles in catalysis and detoxification (Abdul Kayum et al. 2018).
Cis elements are known to control or regulate gene expression, thereby influencing the plant’s response to stress and developmental changes (Narusaka et al 2003). In this study’s analysis of upstream cis-acting elements, 43 different cis-acting elements were detected across the 71 members of the GST gene family. These elements can be categorized into groups associated with light response, hormone response, development, abiotic stress response, and other functions. This diversity of cis elements in the BpGST gene promoters suggests that they might work synergistically to endow BpGSTs with the potential to respond to various stimuli. Significantly, multiple GST gene family members contained MYB binding sites, indicating MYB is likely one of the major factors involved in the regulation of BpGSTs transcriptional expression.
Analysis of gene expression patterns can provide crucial insights into their physiological functions. The expression analysis of the BpGSTFs gene family in birch revealed that 13 exhibited specific expression in different tissues under normal growth conditions, suggesting a significant regulatory role in growth and development. The expression patterns of 13 genes in the Phi BpGSTs gene family in response to high salt, drought, and ABA treatment showed that they were influenced by stress treatment and ABA. However, the expression patterns varied, indicating that each member might have distinct functions in response to stress. In other species, GST gene family members have also shown significant responses to stress stimuli (Simarani et al. 2016). For instance, overexpression of GsGST from soybeans enhanced drought and salt tolerance in transgenic tobacco (Ji et al. 2010). Expression of GmGSTL1 from transgenic soybeans could mitigate salt stress symptoms (Chan and Lam 2014). Sharma et al. (2014) reported that OsGSTU4 in rice is induced by ABA and participates in ABA-dependent processes, providing enhanced tolerance in transgenic plants. In Brassica oleracea, Vijayakumar et al. (2016) identified 65 glutathione transferases (BoGST), with most being highly expressed after 1 and 6 h in cold-sensitive and cold-tolerant lines, respectively, identifying three BoGST (BoGSTU10/19/24) genes as candidates for stress resistance. Wang et al. (2019) reported that 14 of 330 TaGST genes in wheat could respond to different abiotic stresses and hormones, especially salt stress and abscisic acid. In rice, 79 GST genes were identified, with many regulated under various abiotic stresses (20), arsenate stress (32), and biotic stress (48) (Jain et al. 2010).
Conclusion
In summary, the BpGST gene family in birch were wide investigated at the genome level. 71 BpGSTs were identified, and were categorized into seven subfamilies which, in the same branch of the evolutionary tree, had similar exon/intron structures and motif constitutions. These BpGSTs were unevenly located on 14 chromosomes. In addition, eighteen pairs of tandem duplication and five pairs of segmental duplicated genes and were found, suggesting that gene duplication is crucial in the evolution and expansion of the birch GST gene family. Tissue-specific expression of BpGSTFs indicated that these genes may have divergent functions in growth and development. The analysis of cis-acting elements and gene expression profiles under multiple stresses and ABA treatment showed that BpGSTFs play a critical regulatory role in resistance to abiotic stress, and the time of stress response was different as well. These results provide a basis for studying functional characterization of BpGSTs genes in birch and other woody species.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files. The GST protein sequences of Arabidopsis thaliana were downloaded from TAIR database (https://www.arabidopsis.org/). The GST protein sequences of O. sativa and B. platyphylla Suk. were downloaded from Phytozome genome databases (https://phytozome.jgi.doe.gov/pz/portal.html). The accession IDs or numbers of all sequences used in the present study are listed in Table S1, and all databases are available to the public.
References
Abdul Kayum M, Nath UK, Park JI, Biswas MK, Choi EK, Song JY, Kim HT, Nou IS (2018) Genome-wide identification, characterization, and expression profiling of glutathione S-transferase (GST) family in pumpkin reveals likely role in cold-stress tolerance. Genes 9(2):84. https://doi.org/10.3390/genes9020084
Board PG, Baker RT, Chelvanayagam G, Jermiin LS (1997) Zeta, a novel class of glutathione transferases in a range of species from plants to humans. Biochem J 328(3):929–935. https://doi.org/10.1042/bj3280929
Chan C, Lam HM (2014) A putative lambda class glutathione S-transferase enhances plant survival under salinity stress. Plant Cell Phyiol 55(3):570–579. https://doi.org/10.1093/pcp/pct201
Chen CJ, Chen H, Zhang Y, Thomas HR, Frank MH, He YH, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13(8):1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Chen S, Wang YC, Yu LL, Zheng T, Wang S, Yue Z, Jiang J, Kumari S, Zheng CF, Tang HB, Li J, Li YQ, Chen JJ, Zhang WB, Kuang HH, Robertson JS, Zhao PX, Li HY, Shu SQ, Yordanov YS, Huang HJ, Goodstein DM, Gai Y, Qi Q, Min JM, Xu CY, Wang SB, Qu GZ, Paterson AH, Sankoff D, Wei HR, Liu GF, Yang CP (2021) Genome sequence and evolution of Betula platyphylla. Hortic Res 8(1):37. https://doi.org/10.1038/s41438-021-00481-7
Cicero LL, Madesis P, Tsaftaris A, Lo Piero AR (2015) Tobacco plants over-expressing the sweet orange tau glutathione transferases (CsGSTUs) acquire tolerance to the diphenyl ether herbicide fluorodifen and to salt and drought stresses. Phytochemistry 116:69–77. https://doi.org/10.1016/j.phytochem.2015.03.004
Conn S, Curtin C, Bézier A, Franco C, Zhang W (2008) Purification, molecular cloning, and characterization of glutathione S-transferases (GSTs) from pigmented Vitis vinifera L. cell suspension cultures as putative anthocyanin transport proteins. J Exp Bot 59(13):3621–3634. https://doi.org/10.1093/jxb/ern217
Dixon DP, Davis BG, Edwards R (2002) Functional divergence in the glutathione transferase superfamily in plants: identification of two classes with putative functions in redox homeostasis in Arabidopsis thaliana. J Biol Chem 277(34):30859–30869. https://doi.org/10.1074/jbc.M202919200
Dmitriev AA, Krasnov GS, Rozhmina TA, Kishlyan NV, Zyablitsin AV, Sadritdinova AF, Snezhkina AV, Fedorova MS, Yurkevich OY, Muravenko OV, Bolsheva NL, Kudryavtseva AV, Melnikova NV (2016) Glutathione S-transferases and UDP-glycosyltransferases are involved in response to aluminum stress in flax. Front Plant Sci 7:1920. https://doi.org/10.3389/fpls.2016.01920
Dong YT, Li C, Zhang Y, He QL, Daud MK, Chen JH, Zhu SJ (2016) Glutathione S-Transferase gene family in Gossypium raimondii and G arboreum: comparative genomic study and their expression under salt stress. Front Plant Sci 7:139. https://doi.org/10.3389/fpls.2016.00139
Gambino G, Perrone I, Gribaudo I (2008) A rapid and effective method for RNA extraction from different tissues of grapevine and other woody plants. Phytochem Anal 19(6):520–525. https://doi.org/10.1002/pca.1078
Geng WL, Li YY, Sun DQ, Li B, Zhang PY, Chang H, Rong TQ, Liu Y, Shao JW, Liu ZY, Zhu HR, Lou YY, Wang QQ, Zhang JB (2022) Prediction of the potential geographical distribution of Betula platyphylla Suk. in China under climate change scenarios. PLoS One 17(3):e0262540. https://doi.org/10.1371/journal.pone.0262540
Gullner G, Komives T, Király L, Schröder P (2018) Glutathione S-transferase enzymes in plant-pathogen interactions. Front Plant Sci 9:1836. https://doi.org/10.3389/fpls.2018.01836
Hao YC, Xu SS, Lyu ZF, Wang HW, Kong LR, Sun SL (2021) Comparative analysis of the glutathione S-transferase gene family of four Triticeae species and transcriptome analysis of GST genes in common wheat responding to salt stress. Int J Genomics 2021:6289174. https://doi.org/10.1155/2021/6289174
Hu B, Zhao JT, Lai B, Qin YH, Wang HC, Hu GB (2016) LcGST4 is an anthocyanin-related glutathione S-transferase gene in Litchi chinensis Sonn. Plant Cell Rep 35(4):831–843. https://doi.org/10.1007/s00299-015-1924-4
Islam S, Rahman IA, Islam T, Ghosh A (2017) Genome-wide identification and expression analysis of glutathione S-transferase gene family in tomato: gaining an insight to their physiological and stress-specific roles. PLoS ONE 12(11):e0187504. https://doi.org/10.1371/journal.pone.0187504
Jain M, Ghanashyam C, Bhattacharjee A (2010) Comprehensive expression analysis suggests overlapping and specific roles of rice glutathione S-transferase genes during development and stress responses. BMC Genomics 11:73. https://doi.org/10.1186/1471-2164-11-73
Jeffares DC, Penkett CJ, Bähler J (2008) Rapidly regulated genes are intron poor. Trends Genet 24(8):375–378. https://doi.org/10.1016/j.tig.2008.05.006
Jha B, Sharma A, Mishra A (2011) Expression of SbGSTU (tau class glutathione S-transferase) gene isolated from Salicornia brachiata in tobacco for salt tolerance. Mol Biol Rep 38(7):4823–4832. https://doi.org/10.1007/s11033-010-0625-x
Ji W, Zhu YM, Li Y, Yang L, Zhao XW, Cai H, Bai X (2010) Over-expression of a glutathione S-transferase gene, GsGST, from wild soybean (Glycine soja) enhances drought and salt tolerance in transgenic tobacco. Biotechnol Lett 32(8):1173–1179. https://doi.org/10.1007/s10529-010-0269-x
Jia JS, Ge N, Wang QY, Zhao LT, Chen C, Chen JW (2023) Genome-wide identification and characterization of members of the LEA gene family in Panax notoginseng and their transcriptional responses to dehydration of recalcitrant seeds. BMC Genomics 24(1):126. https://doi.org/10.1186/s12864-023-09229-0
Jiang SH, Chen M, He NB, Chen XL, Wang N, Sun QG, Zhang TL, Xu HF, Fang HC, Wang YC, Zhang ZY, Wu SJ, Chen XS (2019) MdGSTF6, activated by MdMYB1, plays an essential role in anthocyanin accumulation in apple. Hortic Res 6:40. https://doi.org/10.1038/s41438-019-0118-6
Jing GH, Hu TM, Liu J, Cheng JM, Li W (2020) Biomass estimation, nutrient accumulation, and stoichiometric characteristics of dominant tree species in the semi-arid region on the Loess Plateau of China. Sustainability 12(1):339. https://doi.org/10.3390/su12010339
Lallement PA, Brouwer B, Keech O, Hecker A, Rouhier N (2014) The still mysterious roles of cysteine-containing glutathione transferases in plants. Front Pharmacol 5:192. https://doi.org/10.3389/fphar.2014.00192
Li B, Zhang XZ, Duan RW, Han CH, Yang J, Wang L, Wang SK, Su YL, Wang L, Dong YF, Xue HB (2022) Genomic analysis of the glutathione S-transferase family in pear (Pyrus communis) and functional identification of PcGST57 in anthocyanin accumulation. Int J Mol Sci 23(2):746. https://doi.org/10.3390/ijms23020746
Licciardello C, D’Agostino N, Traini A, Recupero GR, Frusciante L, Chiusano ML (2014) Characterization of the glutathione S-transferase gene family through ESTs and expression analyses within common and pigmented cultivars of Citrus sinensis (L.) Osbeck. BMC Plant Biol 14:39. https://doi.org/10.1186/1471-2229-14-39
Liu YF, Qi YW, Zhang AL, Wu HX, Liu ZD, Ren XL (2019a) Molecular cloning and functional characterization of AcGST1, an anthocyanin-related glutathione S-transferase gene in kiwifruit (Actinidia chinensis). Plant Mol Biol 100(4–5):451–465. https://doi.org/10.1007/s11103-019-00870-6
Liu YJ, Jiang H, Zhao Y, Li X, Dai XL, Zhuang JH, Zhu MQ, Jiang XL, Wang PQ, Gao LP, Xia T (2019b) Three Camellia sinensis glutathione S-transferases are involved in the storage of anthocyanins, flavonols, and proanthocyanidins. Planta 250(4):1163–1175. https://doi.org/10.1007/s00425-019-03206-2
Liu LB, Zheng SX, Yang DK, Zheng J (2023) Genome-wide in silico identification of glutathione S-transferase (GST) gene family members in fig (Ficus carica L.) and expression characteristics during fruit color development. PeerJ 11:e14406. https://doi.org/10.7717/peerj.14406
Lyu ZY, Yun RX, Wu T, Ma YJ, Chen ZJ, Jin YT, Li JX (2020) Altitudinal differentiation in the radial growth of Betula platyphylla and its response to climate in cold temperate forest a case of Oakley mountain Northeast China. Ying Yong Sheng Tai Xue Bao 31(6):1889–1897. https://doi.org/10.13287/j.1001-9332.202006.011
Marrs KA (1996) The functions and regulation of glutathione S-transferases in plants. Annu Rev Plant Physiol Plant Mol Biol 47:127–158. https://doi.org/10.1146/annurev.arplant.47.1.127
McGonigle B, Keeler SJ, Lau SM, Koeppe MK, O’Keefe DP (2000) A genomics approach to the comprehensive analysis of the glutathione S-transferase gene family in soybean and maize. Plant Physiol 124(3):1105–1120. https://doi.org/10.1104/pp.124.3.1105
Mohsenzadeh S, Esmaeili M, Moosavi F, Shahrtash M, Saffari B, Mohabatkar H (2011) Plant glutathione S-transferase classification, structure and evolution. Afr J Biotechnol 10:8160–8165. https://doi.org/10.5897/AJB11.1024
Mueller LA, Goodman CD, Silady RA, Walbot V (2000) AN9, a petunia glutathione S-transferase required for anthocyanin sequestration, is a flavonoid-binding protein. Plant Physiol 123(4):1561–1570. https://doi.org/10.1104/pp.123.4.1561
Munyampundu JP, Xu YP, Cai XZ (2016) Phi class of glutathione S-transferase gene superfamily widely exists in nonplant taxonomic groups. Evol Bioinform Online 12:59–71. https://doi.org/10.4137/EBO.S35909
Narusaka Y, Nakashima K, Shinwari ZK, Sakuma Y, Furihata T, Abe H, Narusaka M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Interaction between two cis-acting elements, ABRE and DRE, in ABA-dependent expression of Arabidopsis rd29A gene in response to dehydration and high-salinity stresses. Plant J 34(2):137–148. https://doi.org/10.1046/j.1365-313X.2003.01708.x
Nianiou-Obeidat I, Madesis P, Kissoudis C, Voulgari G, Chronopoulou E, Tsaftaris A, Labrou NE (2017) Plant glutathione transferase-mediated stress tolerance: functions and biotechnological applications. Plant Cell Rep 36(6):791–805. https://doi.org/10.1007/s00299-017-2139-7
Nutricati E, Miceli A, Blando F, De Bellis L (2006) Characterization of two Arabidopsis thaliana glutathione S-transferases. Plant Cell Rep 25(9):997–1005. https://doi.org/10.1007/s00299-006-0146-1
Olarewaju O, Ortiz P, Chowdhury W, Chatterjee I, Kinzy T (2004) The translation elongation factor eEF1B plays a role in the oxidative stress response pathway. RNA Biol 1:89–94. https://doi.org/10.4161/rna.1.2.1033
Öztetik E (2008) A tale of plant Glutathione S-transferases: since 1970. Bot Rev 74(3):419–437. https://doi.org/10.1007/s12229-008-9013-9
Qi XL, Liu CL, Song LL, Dong YX, Chen L, Li M (2022) A sweet cherry glutathione S-transferase gene, PavGST1, plays a central role in fruit skin coloration. Cells 11(7):1170. https://doi.org/10.3390/cells11071170
Qiao LY, Zhang XJ, Han X, Zhang L, Li X, Zhan HX, Ma J, Luo PG, Zhang WP, Cui L, Li XY, Chang ZJ (2015) A genome-wide analysis of the auxin/indole-3-acetic acid gene family in hexaploid bread wheat (Triticum aestivum L.). Front Plant Sci 6:770. https://doi.org/10.3389/fpls.2015.00770
Sappl PG, Carroll AJ, Clifton R, Lister R, Whelan J, Harvey Millar A, Singh KB (2009) The Arabidopsis glutathione transferase gene family displays complex stress regulation and co-silencing multiple genes results in altered metabolic sensitivity to oxidative stress. Plant J 58(1):53–68. https://doi.org/10.1111/j.1365-313X.2008.03761.x
Sharma R, Sahoo A, Devendran R, Jain M (2014) Over-expression of a rice tau class glutathione s-transferase gene improves tolerance to salinity and oxidative stresses in Arabidopsis. PLoS ONE 9(3):e92900. https://doi.org/10.1371/journal.pone.0092900
Shehu D, Abdullahi N, Alias Z (2018) Cytosolic glutathione S-transferase in bacteria a review. Pol J Environ Stud 28(2):515–528. https://doi.org/10.15244/pjoes/85200
Simarani K, Yusoff WHAC, Alias Z (2016) Purification of glutathione Transferases (GSTs) from identified Rhizospheric bacteria. Sains Malays 45(7):1057–1062
Sun Y, Li H, Huang JR (2012) Arabidopsis TT19 functions as a carrier to transport anthocyanin from the cytosol to tonoplasts. Mol Plant 5(2):387–400. https://doi.org/10.1093/mp/ssr110
Sylvestre-Gonon E, Law SR, Schwartz M, Robe K, Keech O, Didierjean C, Dubos C, Rouhier N, Hecker A (2019) Functional, structural and biochemical features of plant serinyl-glutathione transferases. Front Plant Sci 10:608. https://doi.org/10.3389/fpls.2019.00608
Vijayakumar H, Thamilarasan SK, Shanmugam A, Natarajan S, Jung HJ, Park JI, Kim H, Chung MY, Nou IS (2016) Glutathione transferases superfamily: cold-inducible expression of distinct GST genes in Brassica oleracea. Int J Mol Sci 17(8):1211. https://doi.org/10.3390/ijms17081211
Wang Z, Huang SZ, Jia CH, Liu JH, Zhang JB, Xu BY, Jin ZQ (2013) Molecular cloning and expression of five glutathione S-transferase (GST) genes from banana (Musa acuminata L. AAA group, cv. Cavendish). Plant Cell Rep 32(9):1373–1380. https://doi.org/10.1007/s00299-013-1449-7
Wang RB, Ma JF, Zhang Q, Wu CL, Zhao HY, Wu YN, Yang GX, He GY (2019) Genome-wide identification and expression profiling of glutathione transferase gene family under multiple stresses and hormone treatments in wheat (Triticum aestivum L.). BMC Genomics 20(1):986. https://doi.org/10.1186/s12864-019-6374-x
Wei K, Wang LY, Zhang YZ, Ruan L, Li HL, Wu LY, Xu LY, Zhang CC, Zhou XG, Cheng H, Edwards R (2019) A coupled role for CsMYB75 and CsGSTF1 in anthocyanin hyperaccumulation in purple tea. Plant J 97(5):825–840. https://doi.org/10.1111/tpj.14161
Wu J, Lv S, Zhao L, Gao T, Yu C, Hu J, Ma F (2023) Advances in the study of the function and mechanism of the action of flavonoids in plants under environmental stresses. Planta 257(6):108. https://doi.org/10.1007/s00425-023-04136-w
Xu GX, Guo CC, Shan HY, Kong HZ (2012) Divergence of duplicate genes in exon-intron structure. Proc Natl Acad Sci USA 109(4):1187–1192. https://doi.org/10.1073/pnas.1109047109
Yaish MW, El-Kereamy A, Zhu T, Beatty PH, Good AG, Bi YM, Rothstein SJ (2010) The APETALA-2-like transcription factor OsAP2-39 controls key interactions between abscisic acid and gibberellin in rice. PLoS Genet 6(9):e1001098. https://doi.org/10.1371/journal.pgen.1001098
Yamazaki M, Shibata M, Nishiyama Y, Springob K, Kitayama M, Shimada N, Aoki T, Ayabe SI, Saito K (2008) Differential gene expression profiles of red and green forms of Perilla frutescens leading to comprehensive identification of anthocyanin biosynthetic genes. FEBS J 275(13):3494–3502. https://doi.org/10.1111/j.1742-4658.2008.06496.x
Yang GY, Chen SW, Li DP, Gao XQ, Su LY, Peng SB, Zhai MZ (2019) Multiple transcriptional regulation of walnut JrGSTTau1 gene in response to osmotic stress. Physiol Plant 166(3):748–761. https://doi.org/10.1111/ppl.12833
Zhang T, Liu L, Jia YL, Zhi JR, Yue WB, Li DY, Zeng G (2022a) Induced resistance combined with RNA interference attenuates the counteradaptation of the western flower Thrips. Int J Mol Sci 23(18):10886. https://doi.org/10.3390/ijms231810886
Zhang Y, Ming RH, Khan M, Wang Y, Dahro B, Xiao W, Li CL, Liu JH (2022) ERF9 of Poncirus trifoliata (L.) Raf undergoes feedback regulation by ethylene and modulates cold tolerance via regulating a glutathione S-transferase U17 gene. Plant Biotechnol J 20(1):183–200. https://doi.org/10.1111/pbi.13705
Zhao Y, Dong WQ, Wang K, Zhang B, Allan AC, Kui LW, Chen KS, Xu CJ (2017) Differential sensitivity of fruit pigmentation to ultraviolet light between two peach cultivars. Front Plant Sci 8:1552. https://doi.org/10.3389/fpls.2017.01552
Zhao Y, Dong WQ, Zhu YC, Allan AC, Kui LW, Xu CJ (2020) PpGST1, an anthocyanin-related glutathione S-transferase gene, is essential for fruit coloration in peach. Plant Biotechnol J 18(5):1284–1295. https://doi.org/10.1111/pbi.13291
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Project funding: This work was supported by the National Key Research and Development Program of China (No. 2021YFD2200304), Fundamental Research Funds for the Central Universities (2572022DQ08), and the National Natural Science Foundation of China (No32171738).
The online version is available at https://link.springer.com/.
Corresponding editor: Lei Yu.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hu, X., Zheng, T., Chen, W. et al. Genome-wide identification and expression analysis of the GST gene family of Betula platyphylla. J. For. Res. 35, 123 (2024). https://doi.org/10.1007/s11676-024-01767-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11676-024-01767-x