Abstract
Purpose
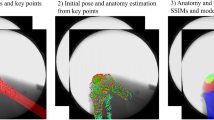
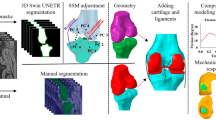
Patient-specific biomechanical models of the knee joint can effectively aid in understanding the reasons for pathologies and improve diagnostic methods and treatment procedures. For deeper research of knee diseases, the development of biomechanical models with appropriate configurations is essential. In this study, we mainly focus on the development of a personalized biomechanical model for the investigation of knee joint pathologies related to patellar motion using automated methods.
Methods
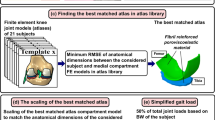
This study presents a biomechanical model created for patellar motion pathologies research and some techniques for automating the generation of the biomechanical model. To generate geometric models of bones, the U-Net neural network was adapted for 3D input datasets. The method uses the same neural network for segmentation of femur, tibia, patella and fibula. The total size of the train/validation (75/25%) dataset is 18,183 3D volumes of size \(512\times 512\times 4\) voxels. The configuration of the biomechanical knee model proposed in the paper includes six degrees of freedom for the tibiofemoral and patellofemoral joints, lateral and medial contact surfaces for femur and tibia, and ligaments, representing, among other things, the medial and lateral stabilizers of the knee cap. The development of the personalized biomechanical model was carried out using the OpenSim software system. The automated model generation was implemented using OpenSim Python scripting commands.
Results
The neural network for bones segmentation achieves mean DICE 0.9838. A biomechanical model for realistic simulation of patellar movement within the trochlear groove was proposed. Generation of personalized biomechanical models was automated.
Conclusions
In this paper, we have implemented a neural network for the segmentation of 3D CT scans of the knee joint to produce a biomechanical model for the study of knee cap motion pathologies. Most stages of the generation process have been automated and can be used to generate patient-specific models.













Similar content being viewed by others
References
Loudon J (2016) Biomechanics and pathomechanics of the patellofemoral joint. Int J Sports Phys Ther 11:820–830
Tarabarko IN, Lychagin AV, Bobrov DS (2018) Status Quo: diagnostics and surgical treatment of excessive lateral pressure syndrome. Orthop Surg. https://doi.org/10.17238/ISSN2226-2016.2018.1.46-51
Coburn SL, Barton CJ, Filbay SR, Hart HF, Rathleff MS, Crossley KM (2018) Quality of life in individuals with patellofemoral pain: a systematic review including meta-analysis. Phys Ther Sport 33:96–108
Boling M, Padua D, Marshall S, Guskiewicz K, Pyne S, Beutler A (2010) Gender differences in the incidence and prevalence of patellofemoral pain syndrome. Scand J Med Sci Sports 20(5):725–730
Robinson RL, Nee RJ (2007) Analysis of hip strength in females seeking physical therapy treatment for unilateral patellofemoral pain syndrome. J Orthop Sports Phys Ther 37:232–238
Fulkerson JP, Arendt EA (2000) Anterior knee pain in females. Clin Orthop Relat Res 431:69–73
Petersen W, Ellermann A, Gösele-Koppenburg A, Best R, Rembitzki IV, Brüggemann GP, Liebau C (2014) Patellofemoral pain syndrome. Knee Surg Sports Traumatol Arthrosc 22(10):2264–74. https://doi.org/10.1007/s00167-013-2759-6
Al-Hakim W, Kumar Jaiswal P, Khan W, Johnstone D (2012) The non-operative treatment of anterior knee pain. Open Orthop J 6(Suppl 2: M10):320–326
Blond L, Hansen L (1998) Patellofemoral pain syndrome in athletes: a 5.7-year retrospective follow-up study of 250 athletes. Acta Orthop Belg 64:393–400
Myer GD, Ford KR, Barber Foss KD, Goodman A, Ceasar A, Rauh MJ, Divine JG, Hewett TE (2010) The incidence and potential pathomechanics of patellofemoral pain in female athletes. Clin Biomech (Bristol, Avon) 25(7):700–707
Thomas MJ, Wood L, Selfe J, Peat G (2010) Anterior knee pain in younger adults as a precursor to subsequent patellofemoral osteoarthritis: a systematic review. BMC Musculoskelet Disord 11:201–208
Utting MR, Davies G, Newman JH (2005) Is anterior knee pain a predisposing factor to patellofemoral osteoarthritis? Knee 12:362–365
Barton CJ, Menz HB, Levinger P, Webster KE, Crossley KM (2011) Greater peak rearfoot eversion predicts foot orthoses efficacy in individuals with patellofemoral pain syndrome. Br J Sports Med 45(9):697–701
Draper CE, Besier TF, Santos JM, Jennings F, Fredericson M, Gold GE, Beaupre GS, Delp SL (2009) Using real-time MRI to quantify altered joint kinematics in subjects with patellofemoral pain and to evaluate the effects of a patellar brace or sleeve on joint motion. J Orthop Res 27(5):571–577
Witvrouw E, Lysens R, Bellemans J, Cambier D, Vanderstraeten G (2000) Intrinsic risk factors for the development of anterior knee pain in an athletic population. A two-year prospective study. Am J Sports Med 28:480–489
Wilson NA, Press JM, Koh JL, Hendrix RW, Zhang LQ (2009) In vivo noninvasive evaluation of abnormal patellar tracking during squatting in patients with patellofemoral pain. J Bone Joint Surg Am 91(3):558–566
Schmitz A, Piovesan D (2016) Development of an open-source, discrete element knee model. IEEE Trans Biomed Eng 63(10):2056–67. https://doi.org/10.1109/TBME.2016.2585926
Xu H, Bloswick D, Merryweather A (2014) An improved OpenSim gait model with multiple degrees of freedom knee joint and knee ligaments. Comput Methods Biomech Biomed Eng. https://doi.org/10.1080/10255842.2014.889689
Bedo BLS, Catelli DS, Lamontagne M, Santiago PRP (2020) A custom musculoskeletal model for estimation of medial and lateral tibiofemoral contact forces during tasks with high knee and hip flexions. Comput Methods Biomech Biomed Engin 23(10):658–663. https://doi.org/10.1080/10255842.2020.1757662
Delp SL, Anderson FC, Arnold AS, Loan P, Habib A, John CT, Guendelman E, Thelen DG (2007) OpenSim: open-source software to create and analyze dynamic simulations of movement. IEEE Trans Biomed Eng 54(11):1940–50. https://doi.org/10.1109/TBME.2007.901024
Delp SL, Loan JP, Hoy MG, Zajac FE, Topp EL, Rosen JM (1990) An interactive graphics-based model of the lower extremity to study orthopaedic surgical procedures. IEEE Trans Biomed Eng 37:757–767
Yamaguchi GT, Zajac FE (1989) A planar model of the knee joint to characterize the knee extensor mechanism. J Biomech 21:1–10
Anderson FC, Pandy MG (1999) A dynamic optimization solution for vertical jumping in three dimensions. Comput Methods Biomech Biomed Eng 2:201–231
Stredney DL (1982) The representation of anatomical structures through computer animation for scientific, educational and artistic applications. Master Thesis, The Ohio State University
Ridhma K, Manvjeet K, Sanjeev S, Devendra Ch (2021) Review of automated segmentation approaches for knee images. IET Image Process. https://doi.org/10.1049/ipr2.12045
Powers CM (2000) Patellar kinematics, part II: the influence of the depth of the trochlear groove in subjects with and without patellofemoral pain. Phys Ther 80(10):965–78
Dong C, Zhao C, Kong L, Piao K, Hao K, Wang F (2022) Medialization of trochlear groove was correlated with extended lateral trochlear in trochlear dysplasia: a transverse CT analysis. J Orthop Surg Res 17(1):276. https://doi.org/10.1186/s13018-022-03166-6
Yurova AS et al An advanced patellofemoral joint biomechanical model for knee pathologies analysis. Sechenov Medical J 15 (to appear)
Patellofemoral joint model for knee pathologies analysis. https://simtk.org/projects/kneepatfemjoint
Ciliberti FK, Guerrini L, Gunnarsson AE, Recenti M, Jacob D, Cangiano V, Tesfahunegn YA, Islind AS, Tortorella F, Tsirilaki M, Jónsson H Jr, Gargiulo P, Aubonnet R (2022) CT- and MRI-Based 3D Reconstruction of Knee Joint to Assess Cartilage and Bone. Diagnostics 12:279. https://doi.org/10.3390/diagnostics12020279
Salamatova VY, Yurova AS, Vassilevski YV, Wang L (2019) Automatic segmentation algorithms and personalized geometric modelling for a human knee. Russ J Numer Anal Math Model 34(6):361–367. https://doi.org/10.1515/rnam-2019-0031
Ronneberger O, Fischer Ph, Brox Th (2015) U-net: convolutional networks for biomedical image segmentation, medical image computing and computer-assisted intervention (MICCAI), LNCS, vol 9351. Springer, Berlin, pp 234–241
Isensee F, Jaeger PF, Kohl SAA, Petersen J, Maier-Hein KH (2021) nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods 18(2):203–211. https://doi.org/10.1038/s41592-020-01008-z
Ioffe S, Szegedy C (2015) Batch normalization: accelerating deep network training by reducing internal covariate shift. In: Proceedings of the 32nd international conference on international conference on machine learning—Volume 37 (ICML’15). JMLR.org, pp 448–456
Srivastava N, Hinton G, Krizhevsky A, Sutskever I, Salakhutdinov R (2014) Dropout: a simple way to prevent neural networks from overfitting. J Mach Learn Res 15:1929–1958
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–302
An open-source multiple-platform application for interactive, scientific visualization. https://www.paraview.org/
GMSH Open Source Mesh Generator. https://gmsh.info/doc/texinfo/gmsh.html
Muntoni A, Cignoni P (2021) PyMeshLab, Zenodo. https://doi.org/10.5281/zenodo.4438750
Amis AA, Senavongse W, Bull AM (2006) Patellofemoral kinematics during knee flexion-extension: an in vitro study. J Orthop Res 24(12):2201–11. https://doi.org/10.1002/jor.20268
Yurova A, Salamatova V, Lychagin A, Vassilevski Y (2022) Automatic detection of attachment sites for knee ligaments and tendons on CT images. Int J Comput Assist Radiol Surg 17(2):393–402. https://doi.org/10.1007/s11548-021-02527-6
Acknowledgements
The authors would like to acknowledge Fedor Loginov for the assistance in processing personalized data.
Funding
The research was funded by Russian Science Foundation Grant 21-71-30023.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
The data were obtained retrospectively from anonymized databases and not generated intentionally for the study. For this type of study, formal consent is not required.
Informed consent
There is no informed consent required for the work reported in this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix A Pseudocode for automated model generation functions
Appendix A Pseudocode for automated model generation functions
-
1.

-
2.

-
3.

-
4.

-
5.

-
6.

-
7.

Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yurova, A., Lychagin, A., Kalinsky, E. et al. Automated personalization of biomechanical knee model. Int J CARS (2024). https://doi.org/10.1007/s11548-024-03075-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11548-024-03075-5