Abstract
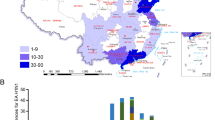
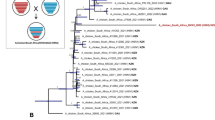
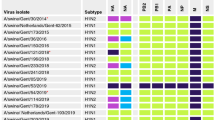
Animal influenza viruses continue to pose a threat to human public health. The Eurasian avian-like H1N1 (EA H1N1) viruses are widespread in pigs throughout Europe and China and have caused human infections in several countries, indicating their pandemic potential. To carefully monitor the evolution of the EA H1N1 viruses in nature, we collected nasal swabs from 103,110 pigs in 22 provinces in China between October 2013 and December 2019, and isolated 855 EA H1N1 viruses. Genomic analysis of 319 representative viruses revealed that these EA H1N1 viruses formed eight different genotypes through reassortment with viruses of other lineages circulating in humans and pigs, and two of these genotypes (G4 and G5) were widely distributed in pigs. Animal studies indicated that some strains have become highly pathogenic in mice and highly transmissible in ferrets via respiratory droplets. Moreover, two-thirds of the EA H1N1 viruses reacted poorly with ferret serum antibodies induced by the currently used H1N1 human influenza vaccine, suggesting that existing immunity may not prevent the transmission of the EA H1N1 viruses in humans. Our study reveals the evolution and pandemic potential of EA H1N1 viruses and provides important insights for future pandemic preparedness.
Similar content being viewed by others
References
Abolnik, C., Pieterse, R., Peyrot, B.M., Choma, P., Phiri, T.P., Ebersohn, K., Heerden, C.J., Vorster, A.A., Zel, G., Geertsma, P.J., et al. (2019). The incursion and spread of highly pathogenic avian influenza H5N8 clade 2.3.4.4 within South Africa. Avian Dis 63, 149–156.
Anderson, T.K., Macken, C.A., Lewis, N.S., Scheuermann, R.H., Van Reeth, K., Brown, I.H., Swenson, S.L., Simon, G., Saito, T., Berhane, Y., et al. (2016). A phylogeny-based global nomenclature system and automated annotation tool for H1 hemagglutinin genes from swine influenza A viruses. mSphere 1, e00275.
Baek, Y.G., Lee, Y.N., Lee, D.H., Shin, J.I., Lee, J.H., Chung, D.H., Lee, E. K., Heo, G.B., Sagong, M., Kye, S.J., et al. (2021). Multiple reassortants of H5N8 clade 2.3.4.4b highly pathogenic avian influenza viruses detected in South Korea during the winter of 2020–2021. Viruses 13, 490.
Bouckaert, R., Heled, J., Kühnert, D., Vaughan, T., Wu, C.H., Xie, D., Suchard, M.A., Rambaut, A., and Drummond, A.J. (2014). BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput Biol 10, e1003537.
Chen, H., Li, Y., Li, Z., Shi, J., Shinya, K., Deng, G., Qi, Q., Tian, G., Fan, S., Zhao, H., et al. (2006). Properties and dissemination of H5N1 viruses isolated during an influenza outbreak in migratory waterfowl in western China. J Virol 80, 5976–5983.
Cui, P., Shi, J., Wang, C., Zhang, Y., Xing, X., Kong, H., Yan, C., Zeng, X., Liu, L., Tian, G., et al. (2022a). Global dissemination of H5N1 influenza viruses bearing the clade 2.3.4.4b HA gene and biologic analysis of the ones detected in China. Emerging Microbes Infects 11, 1693–1704.
Cui, P., Zeng, X., Li, X., Li, Y., Shi, J., Zhao, C., Qu, Z., Wang, Y., Guo, J., Gu, W., et al. (2022b). Genetic and biological characteristics of the globally circulating H5N8 avian influenza viruses and the protective efficacy offered by the poultry vaccine currently used in China. Sci China Life Sci 65, 795–808.
Czudai-Matwich, V., Otte, A., Matrosovich, M., Gabriel, G., and Klenk, H. D. (2014). PB2 mutations D701N and S714R promote adaptation of an influenza H5N1 virus to a mammalian host. J Virol 88, 8735–8742.
Deng, G., Shi, J., Wang, J., Kong, H., Cui, P., Zhang, F., Tan, D., Suzuki, Y., Liu, L., Jiang, Y., et al. (2015). Genetics, receptor binding, and virulence in mice of H10N8 influenza viruses isolated from ducks and chickens in live poultry markets in China. J Virol 89, 6506–6510.
de Jong, J.C., Paccaud, M.F., Ronde-Verloop, F.M., Huffels, N.H., Verwei, C., Weijers, T.F., Bangma, P.J., Kregten, E., Kerckhaert, J.A.M., Wicki, F., et al. (1988). Isolation of swine-like influenza A(H1N1) viruses from man in Switzerland and The Netherlands. Annales de lInstitut Pasteur Virologie 139, 429–437.
Dürrwald, R., Wedde, M., Biere, B., Oh, D.Y., Heßler-Klee, M., Geidel, C., Volmer, R., Hauri, A.M., Gerst, K., Thürmer, A., et al. (2020). Zoonotic infection with swine A/H1avN1 influenza virus in a child, Germany, June 2020. Eurosurveillance 25, 2001638.
El-Shesheny, R., Kandeil, A., Mostafa, A., Ali, M.A., and Webby, R.J. (2021). H5 influenza viruses in Egypt. Cold Spring Harb Perspect Med 11, a038745.
Fraaij, P.L.A., Wildschut, E.D., Houmes, R.J., Swaan, C.M., Hoebe, C.J., de Jonge, H.C.C., Tolsma, P., de Kleer, I., Pas, S.D., Oude Munnink, B. B., et al. (2016). Severe acute respiratory infection caused by swine influenza virus in a child necessitating extracorporeal membrane oxygenation (ECMO), the Netherlands, October 2016. Eurosurveillance 21, 30416.
Gao, S., Anderson, T.K., Walia, R.R., Dorman, K.S., Janas-Martindale, A., and Vincent, A.L. (2017). The genomic evolution of H1 influenza A viruses from swine detected in the United States between 2009 and 2016. J Gen Virol 98, 2001–2010.
Gao, Y., Zhang, Y., Shinya, K., Deng, G., Jiang, Y., Li, Z., Guan, Y., Tian, G., Li, Y., Shi, J., et al. (2009). Identification of amino acids in HA and PB2 critical for the transmission of H5N1 avian influenza viruses in a mammalian host. PLoS Pathog 5, e1000709.
Garten, R.J., Davis, C.T., Russell, C.A., Shu, B., Lindstrom, S., Balish, A., Sessions, W.M., Xu, X., Skepner, E., Deyde, V., et al. (2009). Antigenic and genetic characteristics of swine-origin 2009 A(H1N1) influenza viruses circulating in humans. Science 325, 197–201.
Gregory, V., Bennett, M., Thomas, Y., Kaiser, L., Wunderli, W., Matter, H., Hay, A., and Lin, Y.P. (2003). Human infection by a swine influenza A (H1N1) virus in Switzerland. Arch Virol 148, 793–802.
Gu, W., Shi, J., Cui, P., Yan, C., Zhang, Y., Wang, C., Zhang, Y., Xing, X., Zeng, X., Liu, L., et al. (2022). Novel H5N6 reassortants bearing the clade 2.3.4.4b HA gene of H5N8 virus have been detected in poultry and caused multiple human infections in China. Emerging Microbes Infects 11, 1174–1185.
Guan, L., Shi, J., Kong, X., Ma, S., Zhang, Y., Yin, X., He, X., Liu, L., Suzuki, Y., Li, C., et al. (2019). H3N2 avian influenza viruses detected in live poultry markets in China bind to human-type receptors and transmit in guinea pigs and ferrets. Emerging Microbes Infects 8, 1280–1290.
Hatta, M., Gao, P., Halfmann, P., and Kawaoka, Y. (2001). Molecular basis for high virulence of Hong Kong H5N1 influenza A viruses. Science 293, 1840–1842.
He, J., Wu, Q., Yu, J.L., He, L., Sun, Y., Shi, Y.L., Chen, Q.Q., Ge, Y.L., Zhang, Z.H., Li, W.W., et al. (2020). Sporadic occurrence of H9N2 avian influenza infections in human in Anhui province, eastern China: A notable problem. Microb Pathog 140, 103940.
Jallow, M.M., Fall, A., Barry, M.A., Diop, B., Sy, S., Goudiaby, D., Fall, M., Enouf, V., Niang, M.N., and Dia, N. (2020). Genetic characterization of the first detected human case of low pathogenic avian influenza A/H9N2 in sub-Saharan Africa, Senegal. Emerging Microbes Infects 9, 1092–1095.
Kilbourne, E.D. (2006). Influenza pandemics of the 20th Century. Emerg Infect Dis 12, 9–14.
Kong, H., Ma, S., Wang, J., Gu, C., Wang, Z., Shi, J., Deng, G., Guan, Y., and Chen, H. (2019). Identification of key amino acids in the PB2 and M1 proteins of H7N9 influenza virus that affect its transmission in guinea pigs. J Virol 94, e01180.
Lakdawala, S.S., Jayaraman, A., Halpin, R.A., Lamirande, E.W., Shih, A. R., Stockwell, T.B., Lin, X., Simenauer, A., Hanson, C.T., Vogel, L., et al. (2015). The soft palate is an important site of adaptation for transmissible influenza viruses. Nature 526, 122–125.
Lewis, N.S., Banyard, A.C., Whittard, E., Karibayev, T., Al Kafagi, T., Chvala, I., Byrne, A., Meruyert (Akberovna), S., King, J., Harder, T., et al. (2021). Emergence and spread of novel H5N8, H5N5 and H5N1 clade 2.3.4.4 highly pathogenic avian influenza in 2020. Emerging Microbes Infects 10, 148–151.
Li, X., Guo, L., Liu, C., Cheng, Y., Kong, M., Yang, L., Zhuang, Z., Liu, J., Zou, M., Dong, X., et al. (2019). Human infection with a novel reassortant Eurasian-avian lineage swine H1N1 virus in northern China. Emerging Microbes Infects 8, 1535–1545.
Li, X., Shi, J., Guo, J., Deng, G., Zhang, Q., Wang, J., He, X., Wang, K., Chen, J., Li, Y., et al. (2014). Genetics, receptor binding property, and transmissibility in mammals of naturally isolated H9N2 Avian Influenza viruses. PLoS Pathog 10, e1004508.
Li, Z., Chen, H., Jiao, P., Deng, G., Tian, G., Li, Y., Hoffmann, E., Webster, R.G., Matsuoka, Y., and Yu, K. (2005). Molecular basis of replication of duck H5N1 influenza viruses in a mammalian mouse model. J Virol 79, 12058–12064.
Li, Z., Zhao, X., Huang, W., Yang, L., Cheng, Y., Tan, M., Li, X., Liu, J., Chen, Z., Wei, H., et al. (2022). Etiological characteristics of the first human infection with the G4 genotype Eurasian avian-like H1N1 swine influenza virus in Yunnan province, China. Chin J Virol 38, 290–297.
Liu, K., Ding, P., Pei, Y., Gao, R., Han, W., Zheng, H., Ji, Z., Cai, M., Gu, J., Li, X., et al. (2022). Emergence of a novel reassortant avian influenza virus (H10N3) in Eastern China with high pathogenicity and respiratory droplet transmissibility to mammals. Sci China Life Sci 65, 1024–1035.
Lycett, S.J., Bodewes, R., Pohlmann, A., Banks, J., Bányai, K., Boni, M.F., Bouwstra, R., Breed, A.C., Brown, I.H., Chen, H., et al. (2016). Role for migratory wild birds in the global spread of avian influenza H5N8. Science 354, 213–217.
Meng, F., Yang, H., Qu, Z., Chen, Y., Zhang, Y., Zhang, Y., Liu, L., Zeng, X., Li, C., Kawaoka, Y., et al. (2022). A Eurasian avian-like H1N1 swine influenza reassortant virus became pathogenic and highly transmissible due to mutations in its PA gene. Proc Natl Acad Sci USA 119, e2203919119.
Nguyen, L.T., Schmidt, H.A., von Haeseler, A., and Minh, B.Q. (2015). IQTREE: a fast and effective stochastic algorithm for estimating Maximum-Likelihood phylogenies. Mol Biol Evol 32, 268–274.
Pan, Y., Cui, S., Sun, Y., Zhang, X., Ma, C., Shi, W., Peng, X., Lu, G., Zhang, D., Liu, Y., et al. (2018). Human infection with H9N2 avian influenza in northern China. Clin Microbiol Infect 24, 321–323.
Parys, A., Vandoorn, E., King, J., Graaf, A., Pohlmann, A., Beer, M., Harder, T., and Van Reeth, K. (2021). Human infection with Eurasian avian-like swine influenza A(H1N1) virus, the Netherlands, September 2019. Emerg Infect Dis 27, 939–943.
Pensaert, M., Ottis, K., Vandeputte, J., Kaplan, M.M., and Bachmann, P.A. (1981). Evidence for the natural transmission of influenza A virus from wild ducks to swine and its potential importance for man. Bull World Health Organ 59, 75–78.
Qi, X., Cui, L., Jiao, Y., Pan, Y., Li, X., Zu, R., Huo, X., Wu, B., Tang, F., Song, Y., et al. (2013). Antigenic and genetic characterization of a European avian-like H1N1 swine influenza virus from a boy in China in 2011. Arch Virol 158, 39–53.
Rambaut, A., Lam, T.T., Max Carvalho, L., and Pybus, O.G. (2016). Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol 2, vew007.
Rimmelzwaan, G.F., de Jong, J.C., Bestebroer, T.M., van Loon, A.M., Claas, E.C.J., Fouchier, R.A.M., and Osterhaus, A.D.M.E. (2001). Antigenic and genetic characterization of swine influenza A (H1N1) viruses isolated from pneumonia patients in the Netherlands. Virology 282, 301–306.
Rovida, F., Piralla, A., Marzani, F.C., Moreno, A., Campanini, G., Mojoli, F., Pozzi, M., Girello, A., Chiapponi, C., Vezzoli, F., et al. (2017). Swine influenza A (H1N1) virus (SIV) infection requiring extracorporeal life support in an immunocompetent adult patient with indirect exposure to pigs, Italy, October 2016. Eurosurveillance 22, 30456.
Sancho, B.A., Teres, M.O., Cuenca, S.M., Val, P.R., Villanueva, P.S., Casas, I., Pozo, F., and Brena, P.P. (2009). Human case of swine influenza A (H1N1), Aragon, Spain, November 2008. Euro Surveill 14, 19120.
Shi, J., Deng, G., Kong, H., Gu, C., Ma, S., Yin, X., Zeng, X., Cui, P., Chen, Y., Yang, H., et al. (2017). H7N9 virulent mutants detected in chickens in China pose an increased threat to humans. Cell Res 27, 1409–1421.
Shi, J., Deng, G., Ma, S., Zeng, X., Yin, X., Li, M., Zhang, B., Cui, P., Chen, Y., Yang, H., et al. (2018). Rapid evolution of H7N9 highly pathogenic viruses that emerged in China in 2017. Cell Host Microbe 24, 558–568.e7.
Shope, R.E. (1931). Swine influenza. J Exp Med 54, 349–359.
Song, W., Wang, P., Mok, B.W.Y., Lau, S.Y., Huang, X., Wu, W.L., Zheng, M., Wen, X., Yang, S., Chen, Y., et al. (2014). The K526R substitution in viral protein PB2 enhances the effects of E627K on influenza virus replication. Nat Commun 5, 5509.
Sun, H., Xiao, Y., Liu, J., Wang, D., Li, F., Wang, C., Li, C., Zhu, J., Song, J., Sun, H., et al. (2020). Prevalent Eurasian avian-like H1N1 swine influenza virus with 2009 pandemic viral genes facilitating human infection. Proc Natl Acad Sci USA 117, 17204–17210.
Sun, W., Cheng, S.S.M., Lam, K.N.T., Kwan, T.C., Wong, R.W.K., Lau, L. H.K., Liu, G.Y.Z., Luk, L.L.H., Li, J.K.C., Gu, H., et al. (2022). Natural reassortment of Eurasian avian-like swine H1N1 and avian H9N2 influenza viruses in pigs, China. Emerg Infect Dis 28, 1509–1512.
Tumpey, T.M., Maines, T.R., Van Hoeven, N., Glaser, L., Solorzano, A., Pappas, C., Cox, N.J., Swayne, D.E., Palese, P., Katz, J.M., et al. (2007). A two-amino acid change in the hemagglutinin of the 1918 influenza virus abolishes transmission. Science 315, 655–659.
Um, S., Siegers, J.Y., Sar, B., Chin, S., Patel, S., Bunnary, S., Hak, M., Sor, S., Sokhen, O., Heng, S., et al. (2019). Human Infection with Avian Influenza A(H9N2) virus, Cambodia, February 2021. Emerg Infect Dis 27, 2742–2745.
Wang, D.Y., Qi, S.X., Li, X.Y., Guo, J.F., Tan, M.J., Han, G.Y., Liu, Y.F., Lan, Y., Yang, L., Huang, W.J., et al. (2013). Human infection with Eurasian avian-like influenza A(H1N1) virus, China. Emerg Infect Dis 19, 1709–1711.
Wang, S.Y., Wen, F., Yu, L.X., Wang, J., Wang, M.Z., Yan, J.C., Zhou, Y.J., Tong, W., Shan, T.L., Li, G.X., et al. (2022a). Potential threats to human health from Eurasian Avian-Like Swine Influenza A(H1N1) virus and its reassortants. Emerg Infect Dis 28, 1489–1493.
Wang, Y., Wang, M., Zhang, H., Zhao, C., Zhang, Y., He, G., Deng, G., Cui, P., Li, Y., Liu, W., et al. (2022b). Emergence, evolution, and biological characteristics of H10N4 and H10N8 avian Influenza viruses in migratory wild birds detected in eastern China in 2020. Microbiol Spectr 10, e0080722.
Wang, Z., Yang, H., Chen, Y., Tao, S., Liu, L., Kong, H., Ma, S., Meng, F., Suzuki, Y., Qiao, C., et al. (2017). A single-amino-acid substitution at position 225 in hemagglutinin alters the transmissibility of Eurasian avian-like H1N1 swine influenza virus in guinea pigs. J Virol 91, e00800.
Wasilenko, J.L., Lee, C.W., Sarmento, L., Spackman, E., Kapczynski, D.R., Suarez, D.L., and Pantin-Jackwood, M.J. (2008). NP, PB1, and PB2 viral genes contribute to altered replication of H5N1 avian influenza viruses in chickens. J Virol 82, 4544–4553.
WHO. (2022). World Organisation for Animal Health, Avian Influenza Weekly Update. Available from URL: https://www.who.int/docs/de-fault-source/wpro—documents/emergency/surveillance/avian-influen-za/ai-20220204.pdf?sfvrsn=223ca73f_183.
Xie, J.F., Zhang, Y.H., Zhao, L., Xiu, W.Q., Chen, H.B., Lin, Q., Weng, Y. W., and Zheng, K.C. (2018). Emergence of Eurasian Avian-Like swine influenza A (H1N1) virus from an adult case in Fujian province, China. Virol Sin 33, 282–286.
Xu, C., Xu, B., Wu, Y., Yang, S., Jia, Y., Liang, W., Yang, D., He, L., Zhu, W., Chen, Y., et al. (2020). A single amino acid at position 431 of the PB2 protein determines the virulence of H1N1 swine influenza viruses in mice. J Virol 94, e01930–e01919.
Xu, G., Zhang, X., Gao, W., Wang, C., Wang, J., Sun, H., Sun, Y., Guo, L., Zhang, R., Chang, K.C., et al. (2016). Prevailing PA mutation K356R in avian influenza H9N2 virus increases mammalian replication and pathogenicity. J Virol 90, 8105–8114.
Yang, H., Chen, Y., Qiao, C., He, X., Zhou, H., Sun, Y., Yin, H., Meng, S., Liu, L., Zhang, Q., et al. (2016). Prevalence, genetics, and transmissibility in ferrets of Eurasian avian-like H1N1 swine influenza viruses. Proc Natl Acad Sci USA 113, 392–397.
Yu, G., Smith, D.K., Zhu, H., Guan, Y., Lam, T.T.Y., and McInerny, G. (2016). ggtree: an r package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol Evol 8, 28–36.
Zhang, Q., Shi, J., Deng, G., Guo, J., Zeng, X., He, X., Kong, H., Gu, C., Li, X., Liu, J., et al. (2013a). H7N9 influenza viruses are transmissible in ferrets by respiratory droplet. Science 341, 410–414.
Zhang, Y., Zhang, Q., Gao, Y., He, X., Kong, H., Jiang, Y., Guan, Y., Xia, X., Shu, Y., Kawaoka, Y., et al. (2012). Key molecular factors in hemagglutinin and PB2 contribute to efficient transmission of the 2009 H1N1 pandemic influenza virus. J Virol 86, 9666–9674.
Zhang, Y., Zhang, Q., Kong, H., Jiang, Y., Gao, Y., Deng, G., Shi, J., Tian, G., Liu, L., Liu, J., et al. (2013b). H5N1 hybrid viruses bearing 2009/H1N1 virus genes transmit in guinea pigs by respiratory droplet. Science 340, 1459–1463.
Zhang, Y., Zhao, C., Hou, Y., Chen, Y., Meng, F., Zhuang, Y., Liu, L., Suzuki, Y., Shi, J., Deng, G., et al. (2021). Pandemic threat posed by H3N2 avian influenza virus. Sci China Life Sci 64, 1984–1987.
Zhong, G., Le, M.Q., Lopes, T.J.S., Halfmann, P., Hatta, M., Fan, S., Neumann, G., and Kawaoka, Y. (2018). Mutations in the PA protein of avian H5N1 influenza viruses affect polymerase activity and mouse virulence. J Virol 92, e01557.
Zhu, W., Zhang, H., Xiang, X., Zhong, L., Yang, L., Guo, J., Xie, Y., Li, F., Deng, Z., Feng, H., et al. (2016). Reassortant Eurasian avian-Like influenza A(H1N1) virus from a severely Ill child, Hunan province, China, 2015. Emerg Infect Dis 22, 1930–1936.
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2021YFD1800200, 2021YFC2301700).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Compliance and ethics The author(s) declare that they have no conflict of interest.
Electronic supplementary material
11427_2022_2208_MOESM3_ESM.pdf
Table S1. The reference strains of H1N1 used in the phylogenetic analysis, with the isolation date, subtype, and HA gene clade.
11427_2022_2208_MOESM4_ESM.pdf
Table S2. Mutations detected in the HA, PB2, and PA of EA H1N1 viruses that contribute to increased binding to human-type receptors, pathogenicity, and transmission in mammals.
Rights and permissions
About this article
Cite this article
Meng, F., Chen, Y., Song, Z. et al. Continued evolution of the Eurasian avian-like H1N1 swine influenza viruses in China. Sci. China Life Sci. 66, 269–282 (2023). https://doi.org/10.1007/s11427-022-2208-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-022-2208-0