Abstract
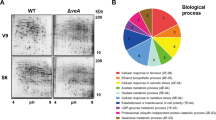
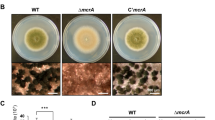
The COP9 signalosome (CSN) is a highly conserved multiprotein complex in all eukaryotes and involved in regulation of organism development. In filamentous fungi, several lines of evidence indicate that fungal development and secondary metabolism (SM) are mediated by the fifth subunit of CSN, called CsnE. Here we uncover a connection with CsnE and conidial formation as well as SM regulation in the plant endophytic fungus Pestalotiopsis fici. A homology search of the P. fici genome with CsnE, involved in sexual development and SM in Aspergillus nidulans, identified PfCsnE. Deletion of PfcsnE resulted in a mutant that stopped conidial production, but the conidia are recovered in a PfcsnE complemented strain. This indicates that PfCsnE is required for the formation of conidia. Secondary metabolite analysis demonstrated that the ΔPfcsnE strain produced more chloroisosulochrin, less ficiolide A production in comparison to wild type (WT). Transcriptome analysis of WT and ΔPfcsnE strains indicated that PfcsnE impacts the expression levels of 8.37% of 14,797 annotated genes. Specifically, nine biosynthetic gene clusters (BGCs) were up-regulated and three BGCs were down-regulated by PfCsnE. Our results suggest that PfCsnE plays major roles in SM regulation and conidial development in P. fici.
Similar content being viewed by others
References
Bayram, O., Feussner, K., Dumkow, M., Herrfurth, C., Feussner, I., and Braus, G.H. (2016). Changes of global gene expression and secondary metabolite accumulation during light-dependent Aspergillus nidulans development. Fungal Genet Biol 87, 30–53.
Beckmann, E.A., Kohler, A.M., Meister, C., Christmann, M., Draht, O.W., Rakebrandt, N., Valerius, O., and Braus, G.H. (2015). Integration of the catalytic subunit activates deneddylase activity in vivo as final step in fungal COP9 signalosome assembly. Mol Microbiol 97, 110–124.
Brakhage, A.A. (2013). Regulation of fungal secondary metabolism. Nat Rev Microbiol 11, 21–32.
Braus, G.H., Irniger, S., and Bayram, O. (2010). Fungal development and the COP9 signalosome. Curr Opin Microbiol 13, 672–676.
Busch, S., Eckert, S.E., Krappmann, S., and Braus, G.H. (2003). The COP9 signalosome is an essential regulator of development in the filamentous fungus Aspergillus nidulans. Mol Microbiol 49, 717–730.
Busch, S., Schwier, E.U., Nahlik, K., Bayram, O., Helmstaedt, K., Draht, O.W., Krappmann, S., Valerius, O., Lipscomb, W.N., and Braus, G.H. (2007). An eight-subunit COP9 signalosome with an intact JAMM motif is required for fungal fruit body formation. Proc Natl Acad Sci USA 104, 8089–8094.
Castrillo, M., and Avalos, J. (2014). Light-mediated participation of the VIVID-like protein of Fusarium fujikuroi VvdA in pigmentation and development. Fungal Genet Biol 71, 9–20.
Christmann, M., Schmaler, T., Gordon, C., Huang, X., Bayram, O., Schinke, J., Stumpf, S., Dubiel, W., and Braus, G.H. (2013). Control of multicellular development by the physically interacting deneddylases DEN1/DenA and COP9 signalosome. PLoS Genet 9, e1003275.
Colot, H.V., Park, G., Turner, G.E., Ringelberg, C., Crew, C.M., Litvinkova, L., Weiss, R.L., Borkovich, K.A., and Dunlap, J.C. (2006). A high-throughput gene knockout procedure for Neurospora reveals functions for multiple transcription factors. Proc Natl Acad Sci USA 103, 10352–10357.
Cope, G.A., Suh, G.S., Aravind, L., Schwarz, S.E., Zipursky, S.L., Koonin, E.V., and Deshaies, R.J. (2002). Role of predicted metalloprotease motif of Jab1/Csn5 in cleavage of Nedd8 from Cul1. Science 298, 608–611.
Gerke, J., Bayram, O., Feussner, K., Landesfeind, M., Shelest, E., Feussner, I., and Braus, G.H. (2012). Breaking the silence: protein stabilization uncovers silenced biosynthetic gene clusters in the fungus Aspergillus nidulans. Appl Environ Microbiol 78, 8234–8244.
Gerke, J., and Braus, G.H. (2014). Manipulation of fungal development as source of novel secondary metabolites for biotechnology. Appl Microbiol Biotechnol 98, 8443–8455.
Hoffmeister, D., and Keller, N.P. (2007). Natural products of filamentous fungi: enzymes, genes, and their regulation. Nat Prod Rep 24, 393–416.
Keller, N.P. (2015). Translating biosynthetic gene clusters into fungal armor and weaponry. Nat Chem Biol 11, 671–677.
Kim, T., Hofmann, K., Von Arnim, A.G., and Chamovitz, D.A. (2001). PCI complexes: pretty complex interactions in diverse signaling pathways. Trends Plant Sci 6, 379–386.
Lima, J.F., Malavazi, I., Von Zeska Kress Fagundes, M.R., Savoldi, M., Goldman, M.H., Schwier, E., Braus, G.H., and Goldman, G.H. (2005). The csnD/csnE signalosome genes are involved in the Aspergillus nidulans DNA damage response. Genetics 171, 1003–1015.
Liu, L. (2011). Bioactive metabolites from the plant endophyte Pestalotiopsis fici. Mycology 2, 37–45.
Liu, L., Liu, S.C., Jiang, L.H., Chen, X.L., Guo, L.D., and Che, Y.S. (2008). Chloropupukeananin, the first chlorinated pupukeanane derivative, and its precursors from Pestalotiopsis fici. Org Lett 10, 1397–1400.
Menon, S., Chi, H., Zhang, H., Deng, X.W., Flavell, R.A., and Wei, N. (2007). COP9 signalosome subunit 8 is essential for peripheral T cell homeostasis and antigen receptor-induced entry into the cell cycle from quiescence. Nat Immunol 8, 1236–1245.
Mundt, K.E., Liu, C., and Carr, A.M. (2002). Deletion mutants in COP9/signalosome subunits in fission yeast Schizosaccharomyces pombe display distinct phenotypes. Mol Biol Cell 13, 493–502.
Nahlik, K., Dumkow, M., Bayram, O., Helmstaedt, K., Busch, S., Valerius, O., Gerke, J., Hoppert, M., Schwier, E., Opitz, L., Westermann, M., Grond, S., Feussner, K., Goebel, C., Kaever, A., Meinicke, P., Feussner, I., and Braus, G.H. (2010). The COP9 signalosome mediates transcriptional and metabolic response to hormones, oxidative stress protection and cell wall rearrangement during fungal development. Mol Microbiol 78, 964–979.
Netzker, T., Fischer, J., Weber, J., Mattern, D.J., Konig, C.C., Valiante, V., Schroeckh, V., and Brakhage, A.A. (2015). Microbial communication leading to the activation of silent fungal secondary metabolite gene clusters. Front Microbiol 6, 299.
Pruss, S., Fetzner, R., Seither, K., Herr, A., Pfeiffer, E., Metzler, M., Lawrence, C.B., and Fischer, R. (2014). Role of the Alternaria alternata blue-light receptor LreA (white-collar 1) in spore formation and secondary metabolism. Appl Environ Microbiol 80, 2582–2591.
Rohlfs, M., and Churchill, A.C. (2011). Fungal secondary metabolites as modulators of interactions with insects and other arthropods. Fungal Genet Biol 48, 23–34.
Rutledge, P.J., and Challis, G.L. (2015). Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat Rev Microbiol 13, 509–523.
Schwechheimer, C., and Isono, E. (2010). The COP9 signalosome and its role in plant development. Eur J Cell Biol 89, 157–162.
Wang, P.M., Choera, T., Wiemann, P., Pisithkul, T., Amador-Noguez, D., and Keller, N.P. (2015a). TrpE feedback mutants reveal roadblocks and conduits toward increasing secondary metabolism in Aspergillus fumigatus. Fungal Genet Biol 89, 102–113.
Wang, X., Wu, F., Liu, L., Liu, X., Che, Y., Keller, N.P., Guo, L., and Yin, W.B. (2015b). The bZIP transcription factor PfZipA regulates secondary metabolism and oxidative stress response in the plant endophytic fungus Pestalotiopsis fici. Fungal Genet Biol 81, 221–228.
Wei, N., Chamovitz, D.A., and Deng, X.W. (1994a). Arabidopsis COP9 is a component of a novel signaling complex mediating light control of development. Cell 78, 117–124.
Wei, N., and Deng, X.W. (2003). The COP9 signalosome. Annu Rev Cell Dev Biol 19, 261–286.
Wei, N., Kwok, S.F., Von Arnim, A.G., Lee, A., Mcnellis, T.W., Piekos, B., and Deng, X.W. (1994b). Arabidopsis COP8, COP10, and COP11 genes are involved in repression of photomorphogenic development in darkness. Plant Cell 6, 629–643.
Wei, N., Serino, G., and Deng, X.W. (2008). The COP9 signalosome: more than a protease. Trends Biochem Sci 33, 592–600.
Wu, G., Zhou, H., Zhang, P., Wang, X., Li, W., Zhang, W., Liu, X., Liu, H.W., Keller, N.P., An, Z., and Yin, W.B. (2016). Polyketide production of pestaloficiols and macrodiolide ficiolides revealed by manipulations of epigenetic regulators in an endophytic fungus. Org Lett 18, 1832–1835.
Xu, X., Liu, L., Zhang, F., Wang, W., Li, J., Guo, L., Che, Y., and Liu, G. (2014). Identification of the first diphenyl ether gene cluster for pestheic acid biosynthesis in plant endophyte Pestalotiopsis fici. Chembiochem 15, 284–292.
Yaegashi, J., Oakley, B.R., and Wang, C.C. (2014). Recent advances in genome mining of secondary metabolite biosynthetic gene clusters and the development of heterologous expression systems in Aspergillus nidulans. J Ind Microbiol Biotechnol 41, 433–442.
Yin, W.B., Reinke, A.W., Szilagyi, M., Emri, T., Chiang, Y.M., Keating, A.E., Pocsi, I., Wang, C.C., and Keller, N.P. (2013). bZIP transcription factors affecting secondary metabolism, sexual development and stress responses in Aspergillus nidulans. Microbiology 159, 77–88.
Yu, D., Xu, F., Zi, J., Wang, S., Gage, D., Zeng, J., and Zhan, J. (2013). Engineered production of fungal anticancer cyclooligomer depsipeptides in Saccharomyces cerevisiae. Metab Eng 18, 60–68.
Yun, C.S., Motoyama, T., and Osada, H. (2015). Biosynthesis of the mycotoxin tenuazonic acid by a fungal NRPS-PKS hybrid enzyme. Nat Commun 6, 8758.
Zhang, A., Lu, P., Dahl-Roshak, A.M., Paress, P.S., Kennedy, S., Tkacz, J.S., and An, Z. (2003). Efficient disruption of a polyketide synthase gene (pks1) required for melanin synthesis through Agrobacterium-mediated transformation of Glarea lozoyensis. Mol Genet Genomics 268, 645–655.
Zhou, C., Seibert, V., Geyer, R., Rhee, E., Lyapina, S., Cope, G., Deshaies, R.J., and Wolf, D.A. (2001). The fission yeast COP9/signalosome is involved in cullin modification by ubiquitin-related Ned8p. BMC Biochem 2, 7.
Zhou, Z., Wang, Y., Cai, G., and He, Q. (2012). Neurospora COP9 signalosome integrity plays major roles for hyphal growth, conidial development, and circadian function. PLoS Genet 8, e1002712.
Acknowledgements
Wenbing Yin is a scholar of “the 100 Talents Project” of Chinese Academy of Sciences. This work was supported by the National Key Research and Development Program (2016YFD0400105), National Natural Science Foundation of China (31670402 and 31400334), and Sate Key Laboratory of Mycology Open Project (SKLMKF 2015-1).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zheng, Y., Wang, X., Zhang, X. et al. COP9 signalosome subunit PfCsnE regulates secondary metabolism and conidial formation in Pestalotiopsis fici . Sci. China Life Sci. 60, 656–664 (2017). https://doi.org/10.1007/s11427-017-9068-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-017-9068-4